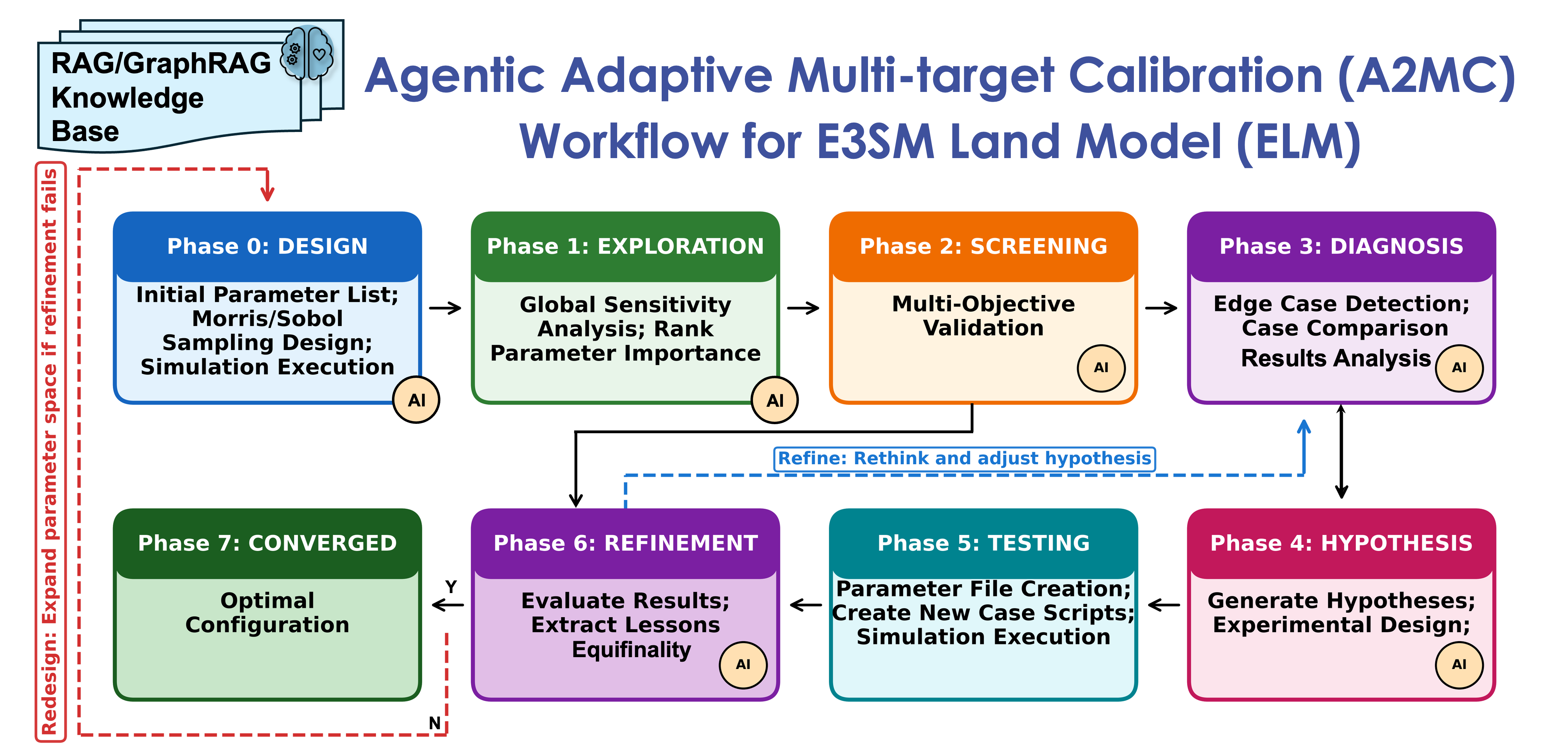
Predicting ecosystem responses to extreme events and changes in environmental conditions with land surface models such as DOE's E3SM Land Model (ELM) remains challenging. These models, such as ELM-FATES (Functionally Assembled Terrestrial Ecosystem Simulator) and ELM-ReSOM (Reaction-network-based model of soil organic matter and Microbes), contain hundreds to thousands of parameters governing vegetation dynamics, biogeochemistry cycling, nutrient uptake, allocation, and life strategy. These parameters must be carefully calibrated against observations at each study site or region. Traditional calibration is a months-to-years-long manual process requiring iterative ensemble runs on HPC, manual inspection of sensitivity analyses, and expert decisions about which parameters to adjust and why. A2MC (Agentic Adaptive Multi-target Calibration), developed at Lawrence Berkeley National Laboratory, reduces this process from months/years to days through an autonomous 7-phase calibration workflow that seamlessly combines LLM reasoning, model-specific RAG/GraphRAG knowledge retrieval, interpretable diagnostics, hypothesis-driven refinement, and persistent adaptive memory, supporting DOE's NGEE-Arctic and Belowground Biogeochemistry SFA projects and accelerating the ModEx (Model-Experiment) loop.
Traditional black-box optimization methods (genetic algorithms, gradient descent) for model calibration identify parameter values that minimize a cost function but provide no insight into why those values work. When calibration fails, scientists have no actionable explanation. A2MC transforms this through interpretable, hypothesis-driven agentic capabilities:
- AI-powered root cause diagnosis. When calibration targets are not met, an LLM reasons over simulation outputs augmented with model-specific RAG/GraphRAG context (e.g., 2,707 documentation chunks, 1,299-node knowledge graph with 2,200 edges for ELM-FATES) to identify mechanistic causes, e.g., diagnosing that arctic graminoid fine-root biomass is under-predicted due to P-limitation in the nutrient competition mechanism.
- Hypothesis generation and experimental design. Based on diagnosis, the agent generates specific, testable parameter modification hypotheses with predicted outcomes, then designs and submits targeted HPC experiments to validate them.
- Multi-target constraint satisfaction. The agent simultaneously optimizes parameters across multiple observational targets (biomass, fluxes, phenology), reducing equifinality by ensuring solutions are mechanistically defensible rather than just numerically coincidental.
- Adaptive memory across sessions. A two-tier persistent memory system records discoveries, failed approaches, and parameter insights in JSON stores, allowing the agent to avoid repeating failures across calibration campaigns at different sites.
- Flexible iteration paths. Rather than following a fixed loop, the agent selects among four iteration paths (re-diagnose, redesign, skip testing, converge) to minimize HPC resource consumption.
A2MC is implemented as a custom Python orchestrator with a 7-phase state machine (Exploration → Screening → Diagnosis → Hypothesis → Testing → Refinement → Converged) running on NERSC Perlmutter login nodes. The orchestrator manages three nested iteration loops, including the full phase 0-7 calibration rounds (expanding the parameter space), experiment cycles (full HPC runs), and skip-testing cycles (lightweight hypothesis testing against existing ensemble data). Workflow state is persisted to JSON after every phase transition, enabling checkpoint/resume across HPC sessions.
Before each AI reasoning call, the orchestrator assembles a rich context by querying three knowledge sources: (1) a hybrid RAG system combining ChromaDB vector search over model documentation chunks with a NetworkX knowledge graph capturing parameter–mechanism–output relationships; (2) a two-tier adaptive memory system that stores mechanistic discoveries, experiment outcomes, and failed approaches in JSON stores at both the framework level (generic model knowledge) and the site level (site-specific calibration lessons); and (3) structured data from previous phases (sensitivity rankings, screening results, diagnostic figures). This context is passed alongside simulation outputs to the LLM, which returns structured JSON responses, including diagnoses with root causes, hypotheses with specific parameter modifications, and/or experiment designs, that are validated before execution.
For HPC integration, the orchestrator directly submits SLURM jobs, polls job status, and extracts results from NetCDF output files. Each experiment uses a session-tagged naming scheme to support concurrent calibration sessions on the same cluster.
Key technologies and frameworks:
- Orchestration: Custom Python state machine with JSON-based checkpoint/resume and session-tagged concurrency
- LLM: Multi-provider support, including Anthropic Claude, OpenAI GPT, and other models via DOE's CBorg proxy (configurable via
A2MC_AI_PROVIDER) - Knowledge retrieval: Hybrid RAG (ChromaDB) + GraphRAG (NetworkX) built from model documentation and codebase wikis (e.g., 2,707 chunks / 1,299 nodes / 2,200 edges for ELM-FATES)
- Sensitivity analysis: SALib (Morris, Sobol, LHS methods)
- HPC execution: NERSC Perlmutter via SLURM (ensemble size and resource usage scale with parameter set and site configuration)
- Memory: Two-tier JSON knowledge stores (generic model knowledge + site-specific calibration lessons)
- Language: Python 3.10+; key deps: anthropic, openai, SALib, netCDF4, chromadb, sentence-transformers
- Repository: https://github.com/jingtao-lbl/A2MC-elm
- Documentation: README and inline phase documentation in repository
- License: BSD-3-Clause
- Contact: Jing Tao — jingtao@lbl.gov (Earth and Environmental Sciences Area, Lawrence Berkeley National Laboratory)
Last Updated: 03/15/2026 Contributed by: Jing Tao (Lawrence Berkeley National Laboratory)