|
| 1 | +# gmr |
| 2 | + |
| 3 | +> Gaussian Mixture Models (GMMs) for clustering and regression in Python. |
| 4 | +
|
| 5 | +[](https://codecov.io/gh/AlexanderFabisch/gmr) |
| 6 | +[](https://doi.org/10.21105/joss.03054) |
| 7 | +[](https://zenodo.org/badge/latestdoi/17119390) |
| 8 | + |
| 9 | +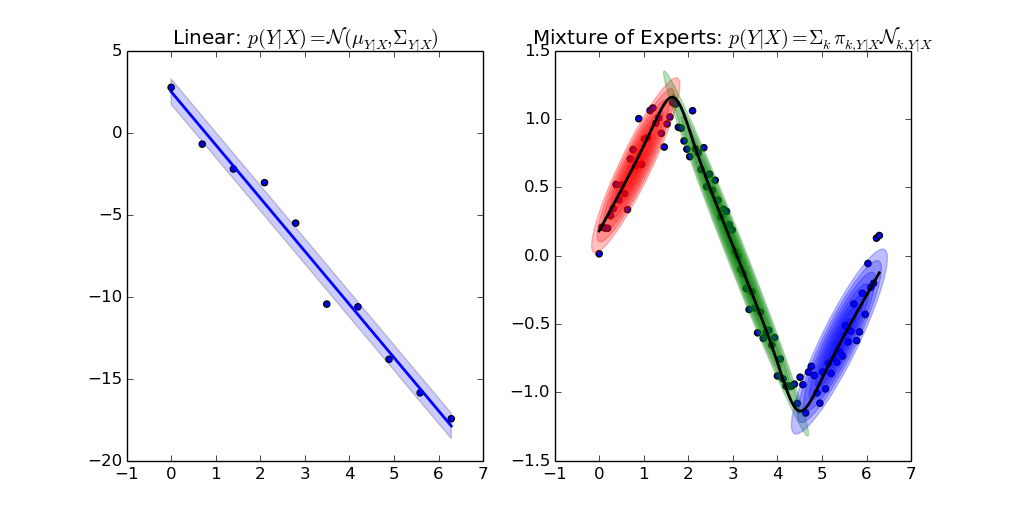 |
| 10 | + |
| 11 | +[(Source code of example)](https://github.com/AlexanderFabisch/gmr/blob/master/examples/plot_regression.py) |
| 12 | + |
| 13 | +* Source code repository: https://github.com/AlexanderFabisch/gmr |
| 14 | +* License: [New BSD / BSD 3-clause](https://github.com/AlexanderFabisch/gmr/blob/master/LICENSE) |
| 15 | +* Releases: https://github.com/AlexanderFabisch/gmr/releases |
| 16 | +* [API documentation](https://alexanderfabisch.github.io/gmr/) |
| 17 | + |
| 18 | +## Documentation |
| 19 | + |
| 20 | +### Installation |
| 21 | + |
| 22 | +Install from [PyPI](https://pypi.python.org/pypi): |
| 23 | + |
| 24 | +```bash |
| 25 | +pip install gmr |
| 26 | +``` |
| 27 | + |
| 28 | +If you want to be able to run all examples, pip can install all necessary |
| 29 | +examples with |
| 30 | + |
| 31 | +```bash |
| 32 | +pip install gmr[all] |
| 33 | +``` |
| 34 | + |
| 35 | +You can also install `gmr` from source: |
| 36 | + |
| 37 | +```bash |
| 38 | +python setup.py install |
| 39 | +# alternatively: pip install -e . |
| 40 | +``` |
| 41 | + |
| 42 | +### Example |
| 43 | + |
| 44 | +Estimate GMM from samples, sample from GMM, and make predictions: |
| 45 | + |
| 46 | +```python |
| 47 | +import numpy as np |
| 48 | +from gmr import GMM |
| 49 | + |
| 50 | +# Your dataset as a NumPy array of shape (n_samples, n_features): |
| 51 | +X = np.random.randn(100, 2) |
| 52 | + |
| 53 | +gmm = GMM(n_components=3, random_state=0) |
| 54 | +gmm.from_samples(X) |
| 55 | + |
| 56 | +# Estimate GMM with expectation maximization: |
| 57 | +X_sampled = gmm.sample(100) |
| 58 | + |
| 59 | +# Make predictions with known values for the first feature: |
| 60 | +x1 = np.random.randn(20, 1) |
| 61 | +x1_index = [0] |
| 62 | +x2_predicted_mean = gmm.predict(x1_index, x1) |
| 63 | +``` |
| 64 | + |
| 65 | +For more details, see: |
| 66 | + |
| 67 | +```python |
| 68 | +help(gmr) |
| 69 | +``` |
| 70 | + |
| 71 | +or have a look at the |
| 72 | +[API documentation](https://alexanderfabisch.github.io/gmr/) |
| 73 | + |
| 74 | +You can see the results of all the examples [here](https://github.com/AlexanderFabisch/gmr/tree/master/examples/examples-with-gmr.ipynb>). |
| 75 | + |
| 76 | +You can find worked examples in [this Google Colab notebook](https://colab.research.google.com/drive/1fJK7z8Jhn04O6NxuPZMdLCsXT5HjvnyD?usp=sharing). |
| 77 | + |
| 78 | +### How Does It Compare to scikit-learn? |
| 79 | + |
| 80 | +There is an implementation of Gaussian Mixture Models for clustering in |
| 81 | +[scikit-learn](https://scikit-learn.org/stable/modules/classes.html#module-sklearn.mixture>) |
| 82 | +as well. Regression could not be easily integrated in the interface of |
| 83 | +sklearn. That is the reason why I put the code in a separate repository. |
| 84 | +It is possible to initialize GMR from sklearn though: |
| 85 | + |
| 86 | +```python |
| 87 | +from sklearn.mixture import GaussianMixture |
| 88 | +from gmr import GMM |
| 89 | +gmm_sklearn = GaussianMixture(n_components=3, covariance_type="diag") |
| 90 | +gmm_sklearn.fit(X) |
| 91 | +gmm = GMM( |
| 92 | + n_components=3, priors=gmm_sklearn.weights_, means=gmm_sklearn.means_, |
| 93 | + covariances=np.array([np.diag(c) for c in gmm_sklearn.covariances_])) |
| 94 | +``` |
| 95 | + |
| 96 | +For model selection with sklearn we furthermore provide an optional |
| 97 | +regressor interface. |
| 98 | + |
| 99 | + |
| 100 | +### Gallery |
| 101 | + |
| 102 | +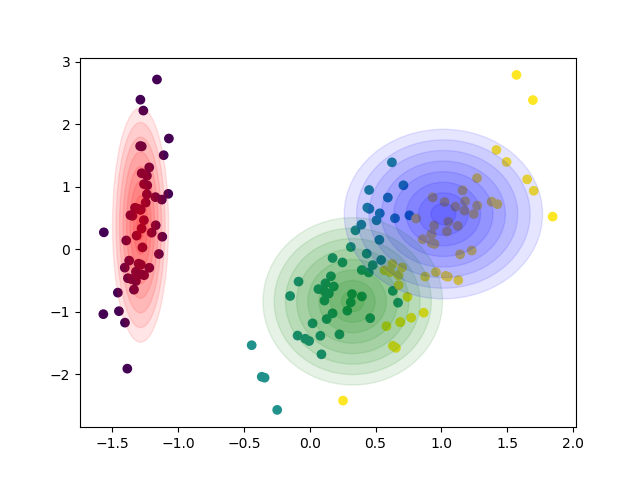 |
| 103 | + |
| 104 | +[Diagonal covariances](https://github.com/AlexanderFabisch/gmr/blob/master/examples/plot_iris_from_sklearn.py) |
| 105 | + |
| 106 | +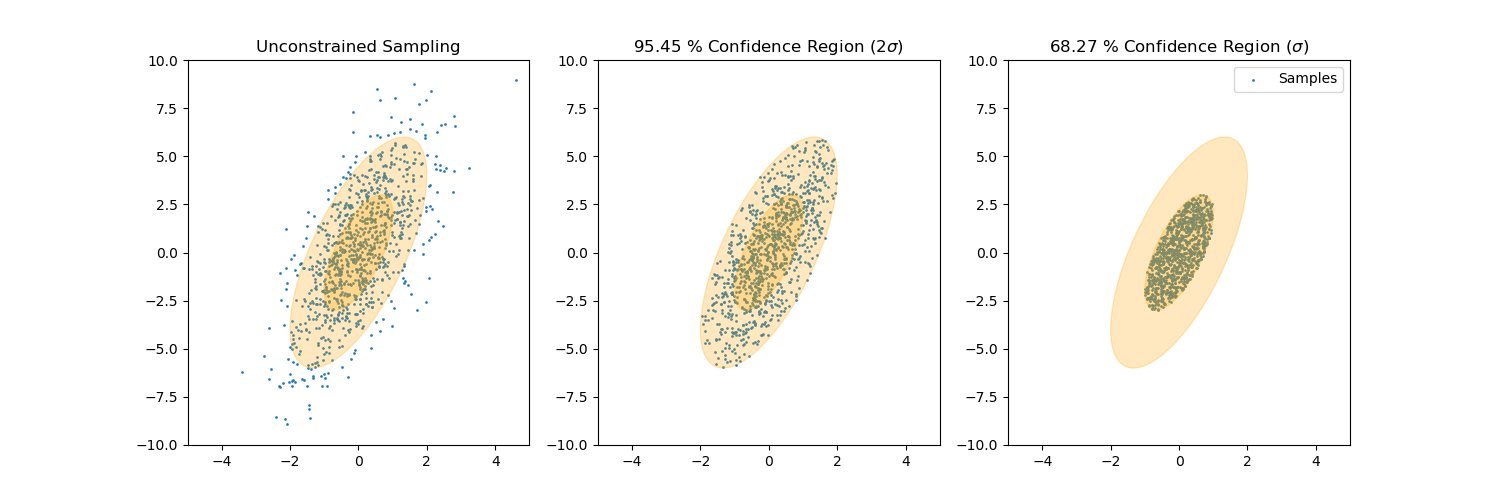 |
| 107 | + |
| 108 | +[Sample from confidence interval](https://github.com/AlexanderFabisch/gmr/blob/master/examples/plot_sample_mvn_confidence_interval.py) |
| 109 | + |
| 110 | +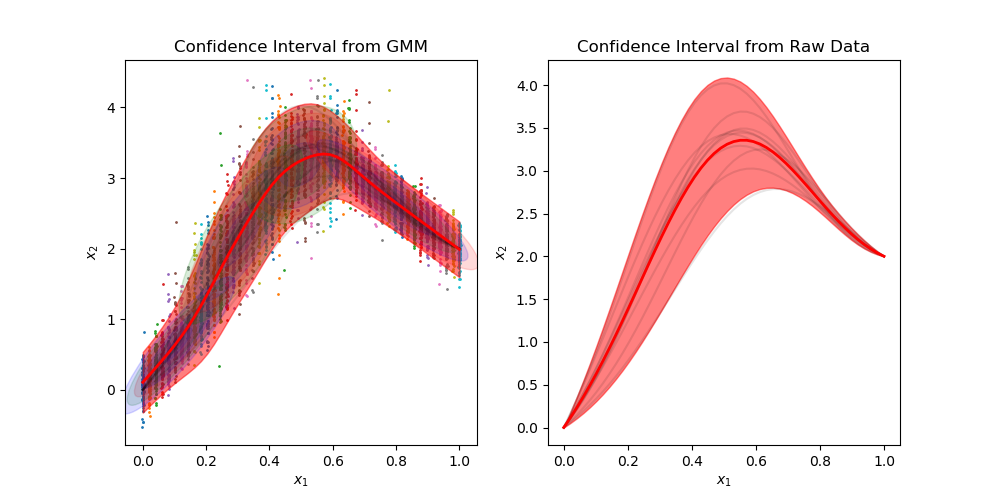 |
| 111 | + |
| 112 | +[Generate trajectories](https://github.com/AlexanderFabisch/gmr/blob/master/examples/plot_trajectories.py) |
| 113 | + |
| 114 | +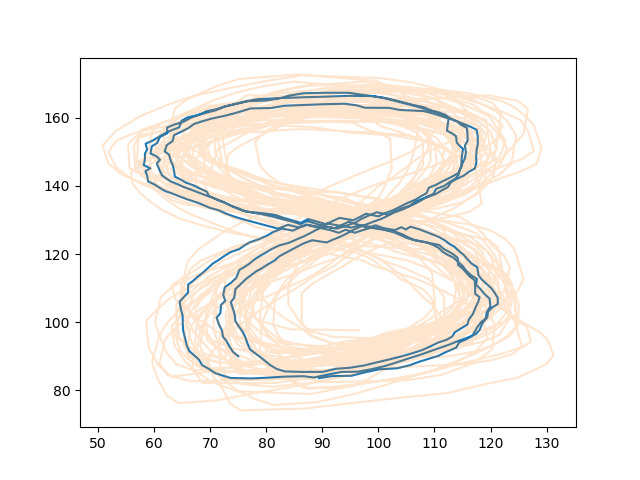 |
| 115 | + |
| 116 | +[Sample time-invariant trajectories](https://github.com/AlexanderFabisch/gmr/blob/master/examples/plot_time_invariant_trajectories.py) |
| 117 | + |
| 118 | +You can find [all examples here](https://github.com/AlexanderFabisch/gmr/tree/master/examples). |
| 119 | + |
| 120 | + |
| 121 | +### Saving a Model |
| 122 | + |
| 123 | +This library does not directly offer a function to store fitted models. Since |
| 124 | +the implementation is pure Python, it is possible, however, to use standard |
| 125 | +Python tools to store Python objects. For example, you can use pickle to |
| 126 | +temporarily store a GMM: |
| 127 | + |
| 128 | +```python |
| 129 | +import numpy as np |
| 130 | +import pickle |
| 131 | +import gmr |
| 132 | +gmm = gmr.GMM(n_components=2) |
| 133 | +gmm.from_samples(X=np.random.randn(1000, 3)) |
| 134 | + |
| 135 | +# Save object gmm to file 'file' |
| 136 | +pickle.dump(gmm, open("file", "wb")) |
| 137 | +# Load object from file 'file' |
| 138 | +gmm2 = pickle.load(open("file", "rb")) |
| 139 | +``` |
| 140 | + |
| 141 | +It might be required to store models more permanently than in a pickle file, |
| 142 | +which might break with a change of the library or with the Python version. |
| 143 | +In this case you can choose a storage format that you like and store the |
| 144 | +attributes `gmm.priors`, `gmm.means`, and `gmm.covariances`. These can be |
| 145 | +used in the constructor of the GMM class to recreate the object and they can |
| 146 | +also be used in other libraries that provide a GMM implementation. The |
| 147 | +MVN class only needs the attributes `mean` and `covariance` to define the |
| 148 | +model. |
| 149 | + |
| 150 | + |
| 151 | +### API Documentation |
| 152 | + |
| 153 | +API documentation is available [here](https://alexanderfabisch.github.io/gmr/). |
| 154 | + |
| 155 | + |
| 156 | +### Citation |
| 157 | + |
| 158 | +If you use the library gmr in a scientific publication, I would appreciate |
| 159 | +citation of the following paper: |
| 160 | + |
| 161 | +> Fabisch, A., (2021). gmr: Gaussian Mixture Regression. Journal of Open Source |
| 162 | +> Software, 6(62), 3054, https://doi.org/10.21105/joss.03054 |
| 163 | +
|
| 164 | +Bibtex entry: |
| 165 | + |
| 166 | +```bibtex |
| 167 | +@article{Fabisch2021, |
| 168 | +doi = {10.21105/joss.03054}, |
| 169 | +url = {https://doi.org/10.21105/joss.03054}, |
| 170 | +year = {2021}, |
| 171 | +publisher = {The Open Journal}, |
| 172 | +volume = {6}, |
| 173 | +number = {62}, |
| 174 | +pages = {3054}, |
| 175 | +author = {Alexander Fabisch}, |
| 176 | +title = {gmr: Gaussian Mixture Regression}, |
| 177 | +journal = {Journal of Open Source Software} |
| 178 | +} |
| 179 | +``` |
| 180 | + |
| 181 | + |
| 182 | +## Contributing |
| 183 | + |
| 184 | +### How can I contribute? |
| 185 | + |
| 186 | +If you discover bugs, have feature requests, or want to improve the |
| 187 | +documentation, you can open an issue at the |
| 188 | +[issue tracker](https://github.com/AlexanderFabisch/gmr/issues) |
| 189 | +of the project. |
| 190 | + |
| 191 | +If you want to contribute code, please open a pull request via |
| 192 | +GitHub by forking the project, committing changes to your fork, |
| 193 | +and then opening a |
| 194 | +[pull request](https://github.com/AlexanderFabisch/gmr/pulls) |
| 195 | +from your forked branch to the main branch of `gmr`. |
| 196 | + |
| 197 | + |
| 198 | +### Development Environment |
| 199 | + |
| 200 | +I would recommend to install `gmr` from source in editable mode with `pip` and |
| 201 | +install all dependencies: |
| 202 | + |
| 203 | +```bash |
| 204 | +pip install -e .[all,test,doc] |
| 205 | +``` |
| 206 | + |
| 207 | +You can now run tests with |
| 208 | + |
| 209 | +```bash |
| 210 | +pytest |
| 211 | +``` |
| 212 | + |
| 213 | +This will also generate a coverage report and output an HTML overview to |
| 214 | +the folder `htmlcov/`. |
| 215 | + |
| 216 | +### Generate Documentation |
| 217 | + |
| 218 | +The API documentation is generated with |
| 219 | +[pdoc3](https://pdoc3.github.io/pdoc/). If you want to regenerate it, |
| 220 | +you can run |
| 221 | + |
| 222 | +```bash |
| 223 | +pdoc gmr --html --skip-errors |
| 224 | +``` |
| 225 | + |
| 226 | + |
| 227 | +## Related Publications |
| 228 | + |
| 229 | +The first publication that presents the GMR algorithm is |
| 230 | + |
| 231 | +> [1] Z. Ghahramani, M. I. Jordan, "Supervised learning from incomplete data via an EM approach," Advances in Neural Information Processing Systems 6, 1994, pp. 120-127, http://papers.nips.cc/paper/767-supervised-learning-from-incomplete-data-via-an-em-approach |
| 232 | +
|
| 233 | +but it does not use the term Gaussian Mixture Regression, which to my knowledge occurs first in |
| 234 | + |
| 235 | +> [2] S. Calinon, F. Guenter and A. Billard, "On Learning, Representing, and Generalizing a Task in a Humanoid Robot," in IEEE Transactions on Systems, Man, and Cybernetics, Part B (Cybernetics), vol. 37, no. 2, 2007, pp. 286-298, doi: [10.1109/TSMCB.2006.886952](https://doi.org/10.1109/TSMCB.2006.886952). |
| 236 | +
|
| 237 | +A recent survey on various regression models including GMR is the following: |
| 238 | + |
| 239 | +> [3] F. Stulp, O. Sigaud, "Many regression algorithms, one unified model: A review," in Neural Networks, vol. 69, 2015, pp. 60-79, doi: [10.1016/j.neunet.2015.05.005](https://doi.org/10.1016/j.neunet.2015.05.005). |
| 240 | +
|
| 241 | +Sylvain Calinon has a good introduction in his [slides on nonlinear regression](https://calinon.ch/misc/EE613/EE613-nonlinearRegression.pdf) for his [machine learning course](http://calinon.ch/teaching_EPFL.htm). |
0 commit comments