You signed in with another tab or window. Reload to refresh your session.You signed out in another tab or window. Reload to refresh your session.You switched accounts on another tab or window. Reload to refresh your session.Dismiss alert
Copy file name to clipboardExpand all lines: content/100.figure-table-legends.md
+14-14Lines changed: 14 additions & 14 deletions
Display the source diff
Display the rich diff
Original file line number
Diff line number
Diff line change
@@ -1,7 +1,7 @@
1
1
## Figure Titles and Legends {.page_break_before}
2
2
3
3
<!-- Figure 1 -->
4
-
{#fig:fig1 tag="1" width="7in"}
4
+
{#fig:fig1 tag="1" width="7in"}
5
5
6
6
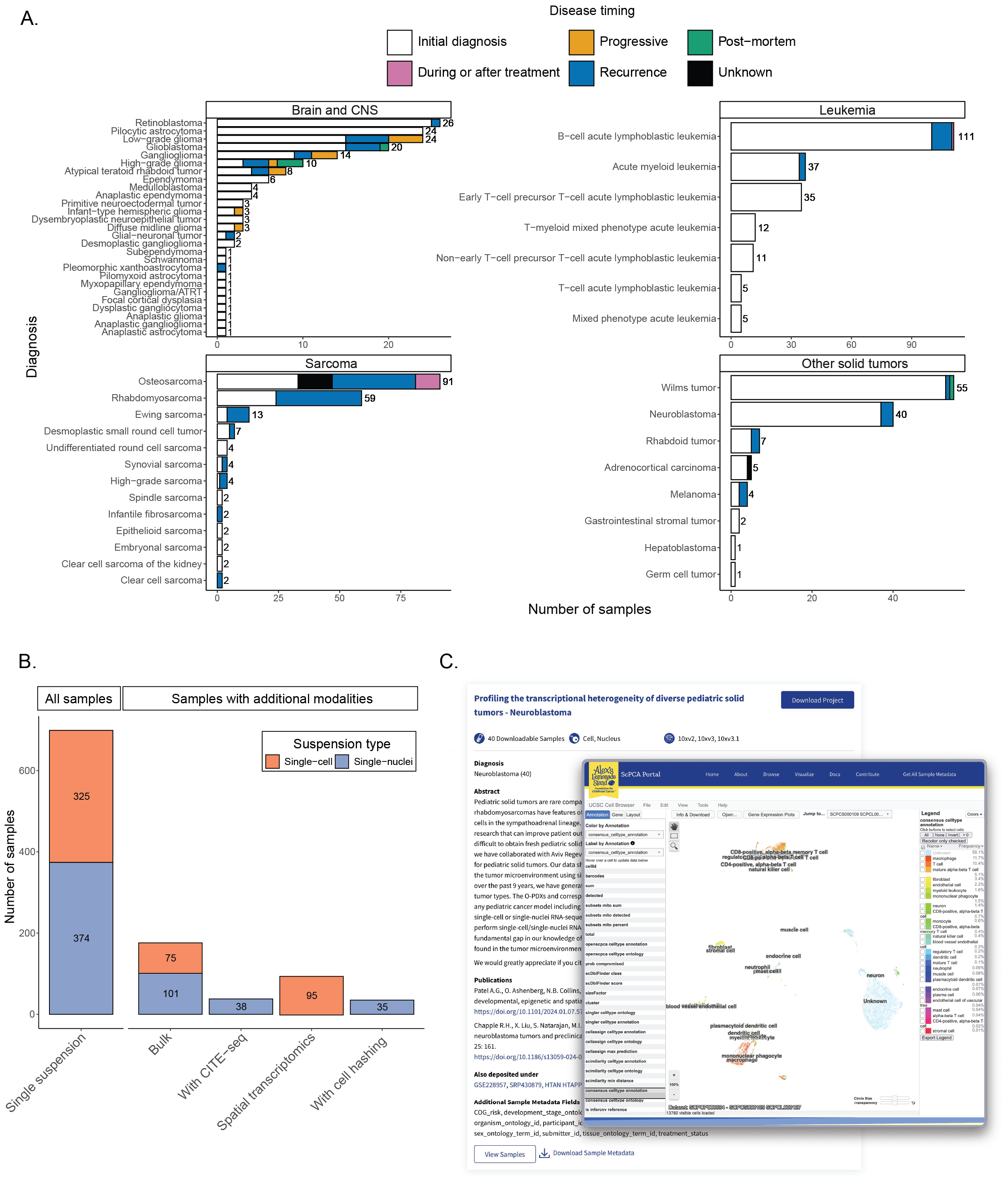
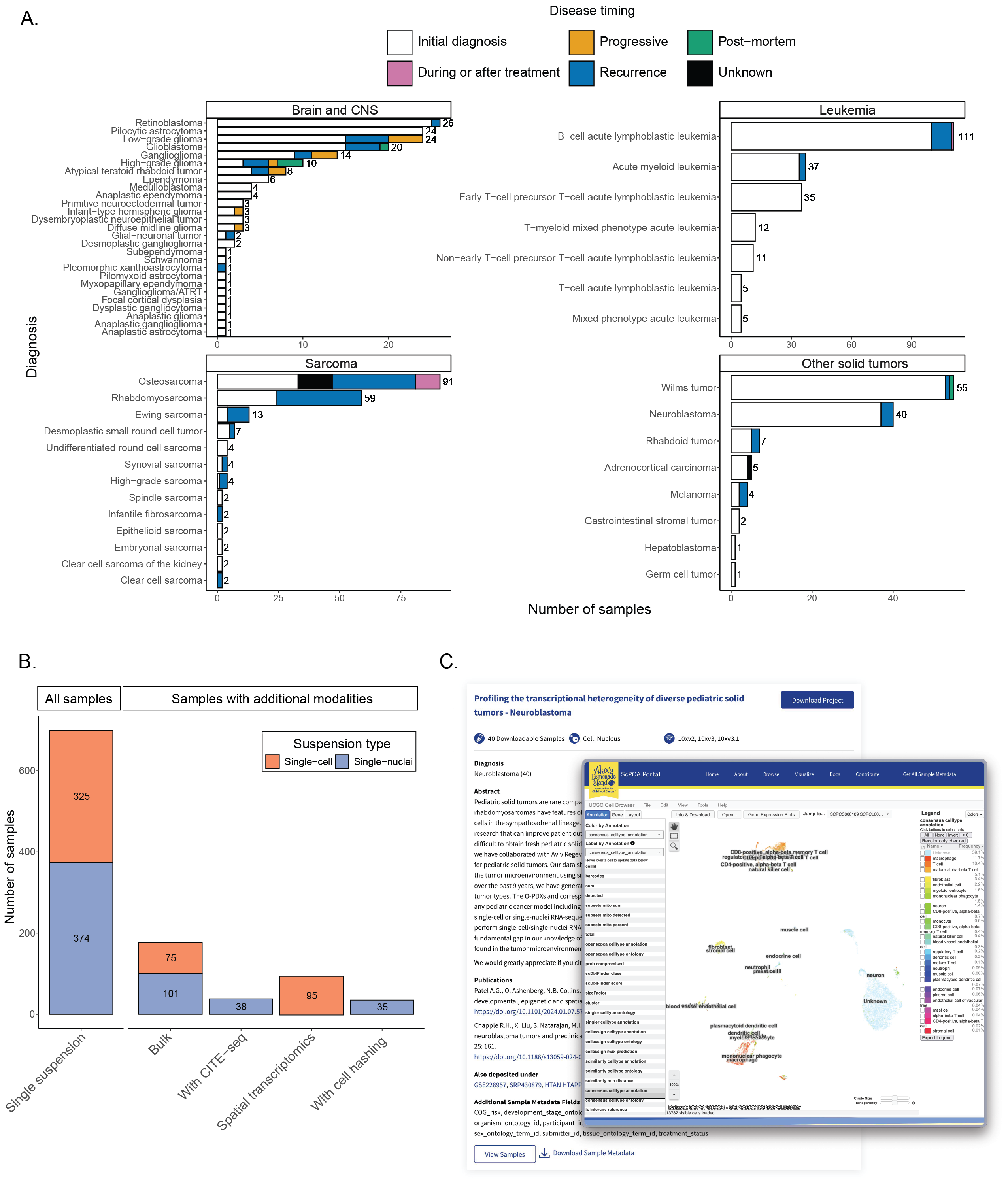
A. Barplots showing sample counts across four main cancer groupings in the ScPCA Portal with the total number of samples for each cancer type displayed to the right of each bar.
7
7
Each bar is colored based on the number of samples with the indicated disease timing.
@@ -23,7 +23,7 @@ The visualization employs the UCSC Cell Browser [@doi:10.1093/bioinformatics/bta
23
23
24
24
25
25
<!-- Figure 2 -->
26
-
{#fig:fig2 tag="2" width="7in"}
26
+
{#fig:fig2 tag="2" width="7in"}
27
27
28
28
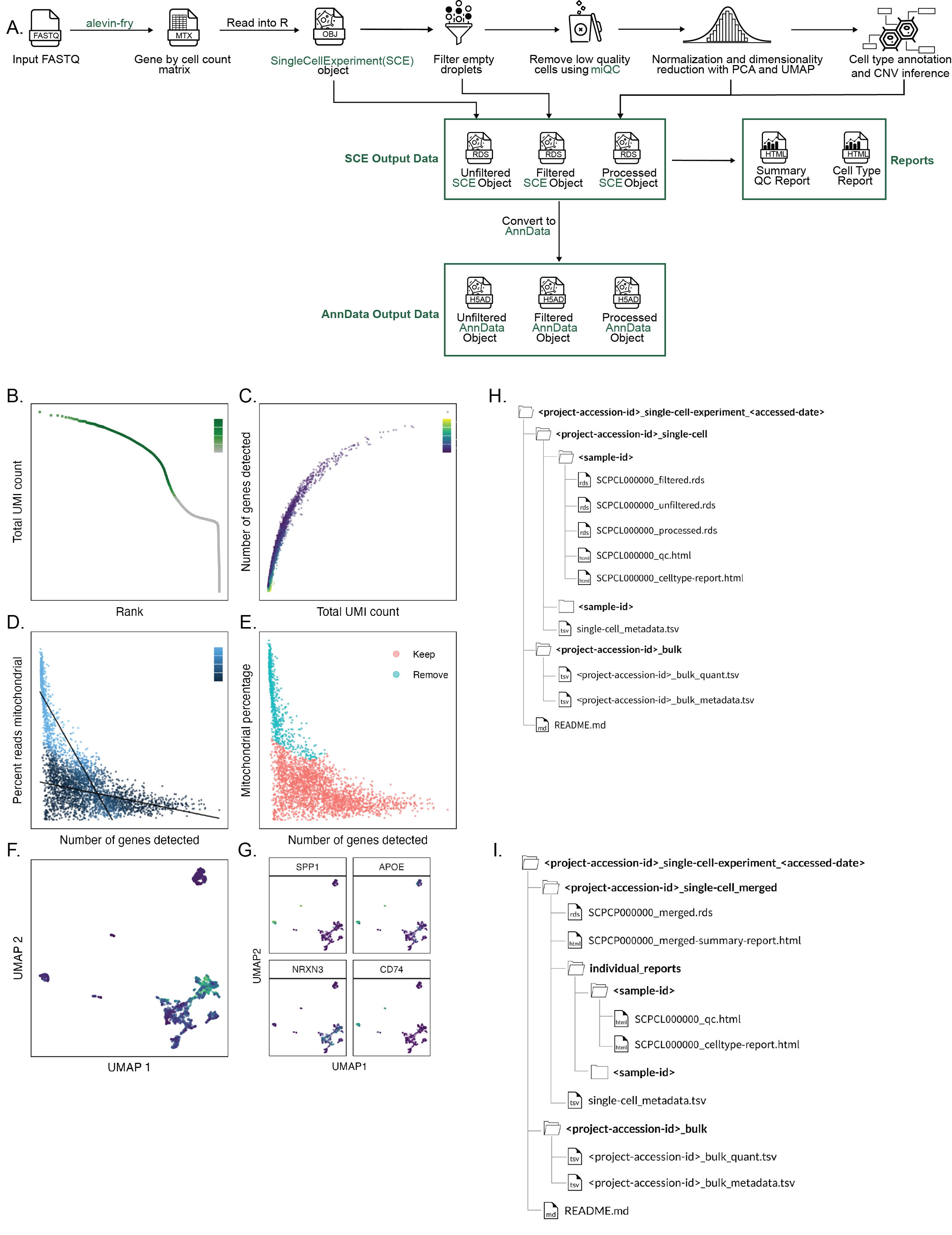
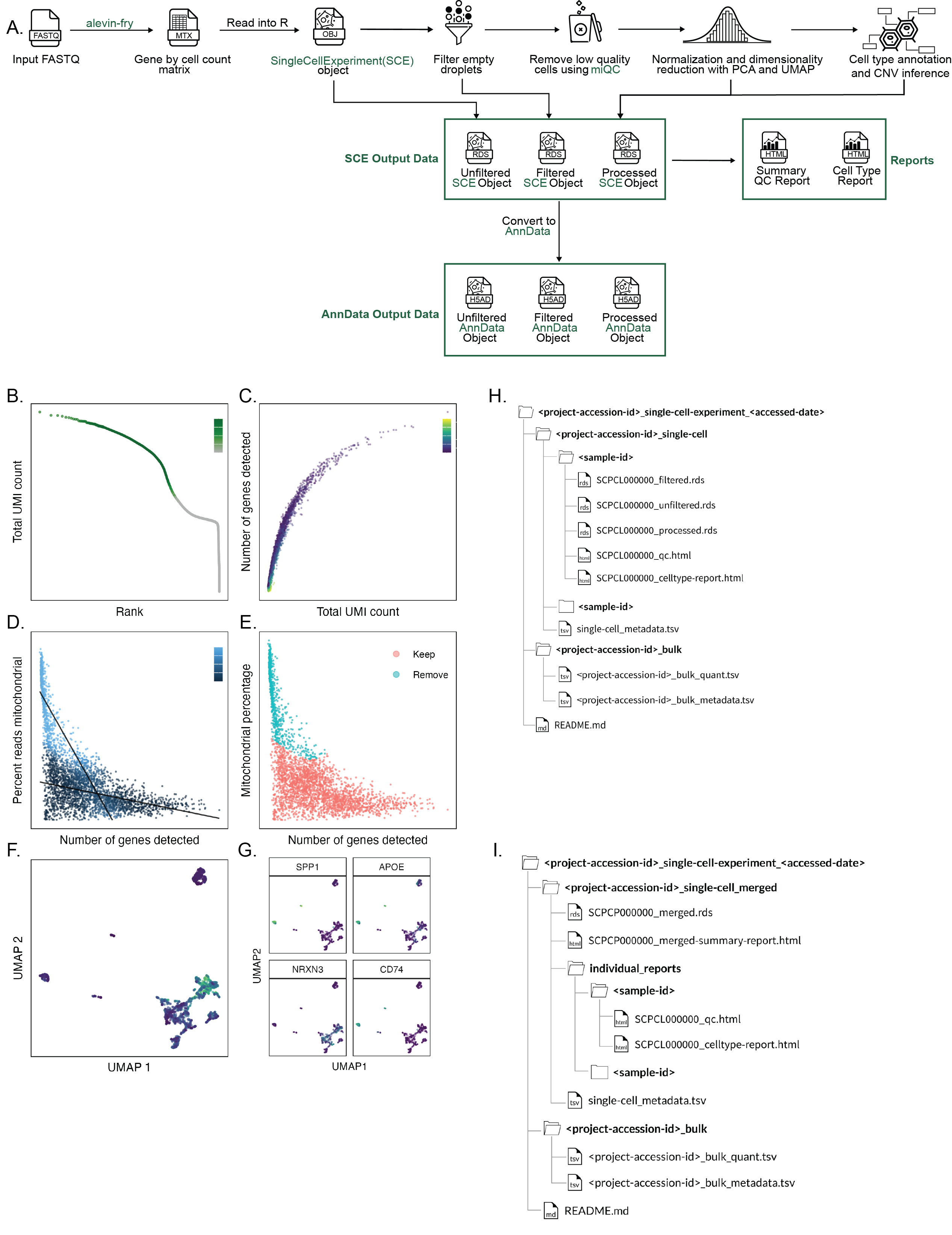
A. Overview of `scpca-nf`, the primary workflow for processing single-cell and single-nuclei RNA-seq data for the ScPCA Portal.
29
29
Mapping is first performed with `alevin-fry` to generate a gene-by-cell count matrix, which is read into `R` and converted into a `SingleCellExperiment` (`SCE`) object.
@@ -72,7 +72,7 @@ As in panel (H), additional files `single-cell_metadata.tsv`, `_bulk_quant.tsv`,
72
72
73
73
74
74
<!-- Figure 3 -->
75
-
{#fig:fig3 tag="3" width="7in"}
75
+
{#fig:fig3 tag="3" width="7in"}
76
76
77
77
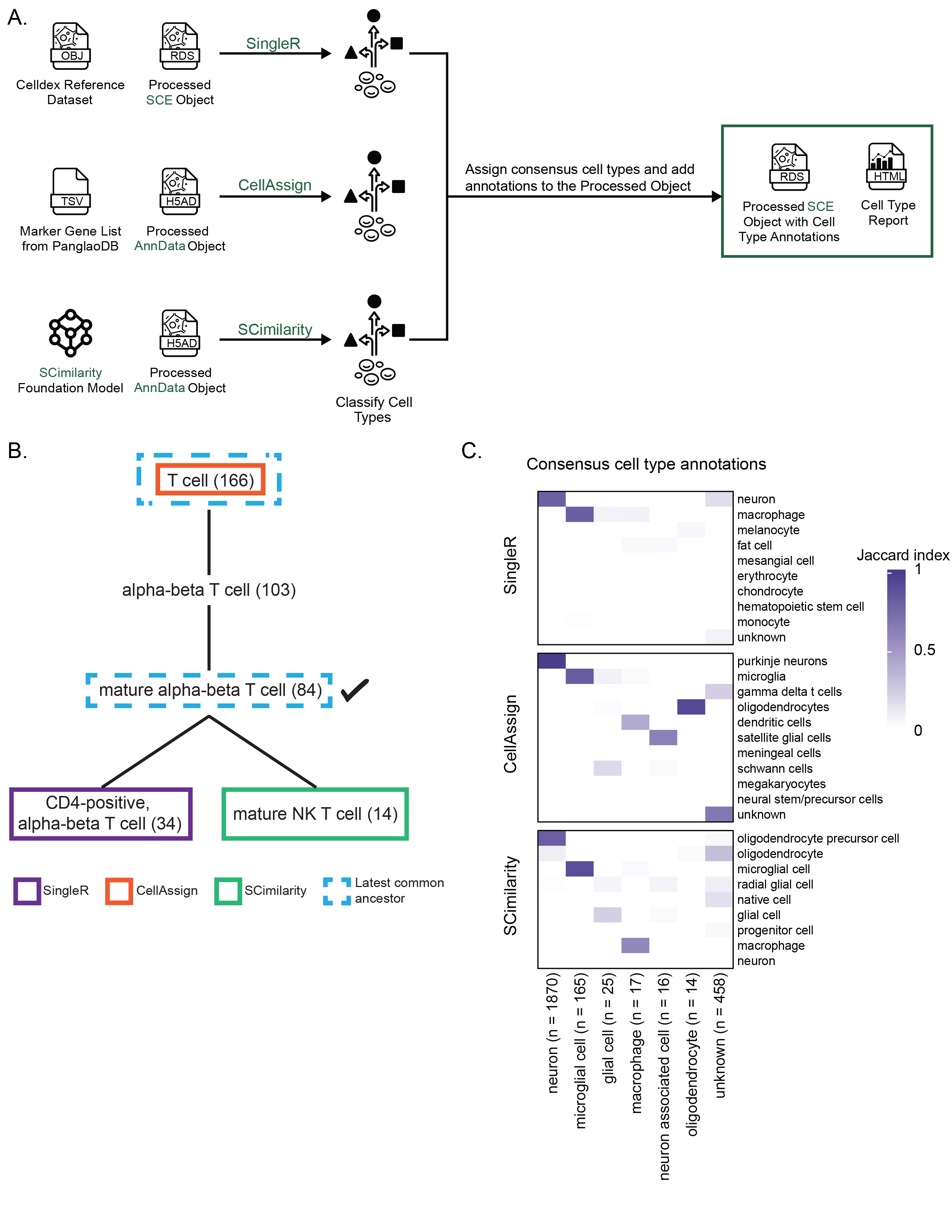
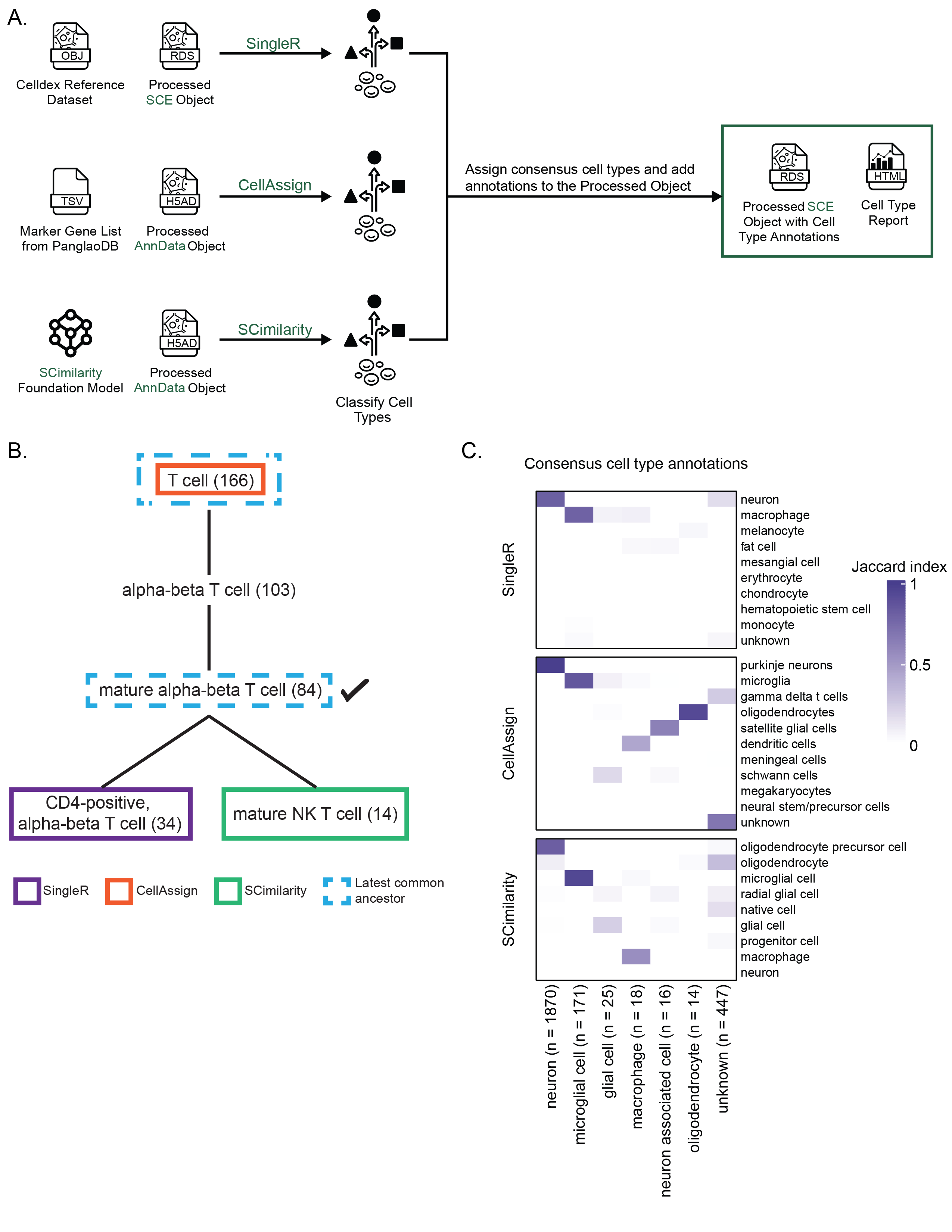
A. Expanded view of the process for adding cell type annotations within `scpca-nf`, as introduced in Figure {@fig:fig2}A.
78
78
Cell type annotation is performed on the `Processed SCE Object`.
@@ -96,7 +96,7 @@ For this figure specifically, the heatmap shows only the top seven consensus cel
96
96
97
97
98
98
<!-- Figure 4 -->
99
-
{#fig:fig4 tag="4" width="7in"}
99
+
{#fig:fig4 tag="4" width="7in"}
100
100
101
101
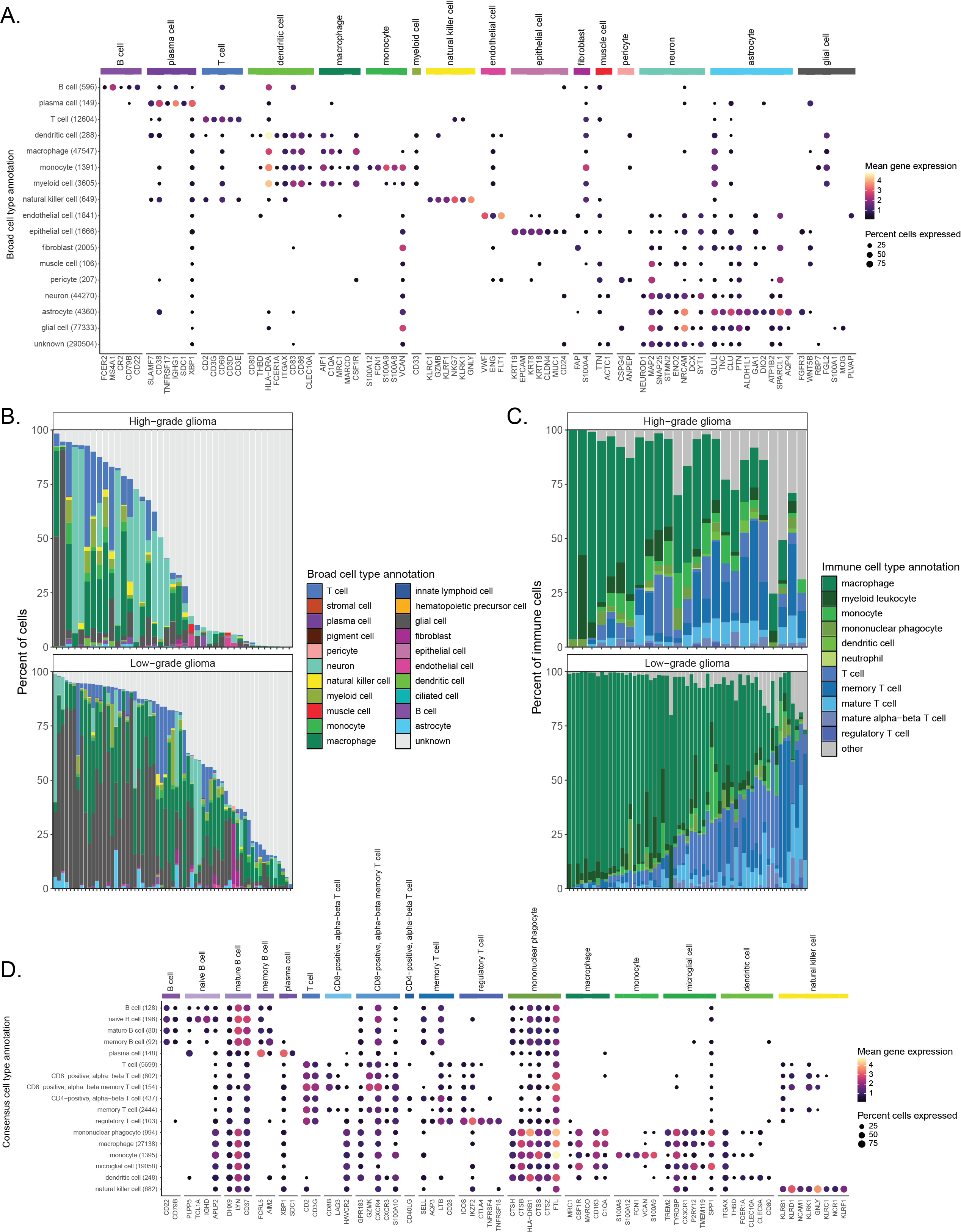
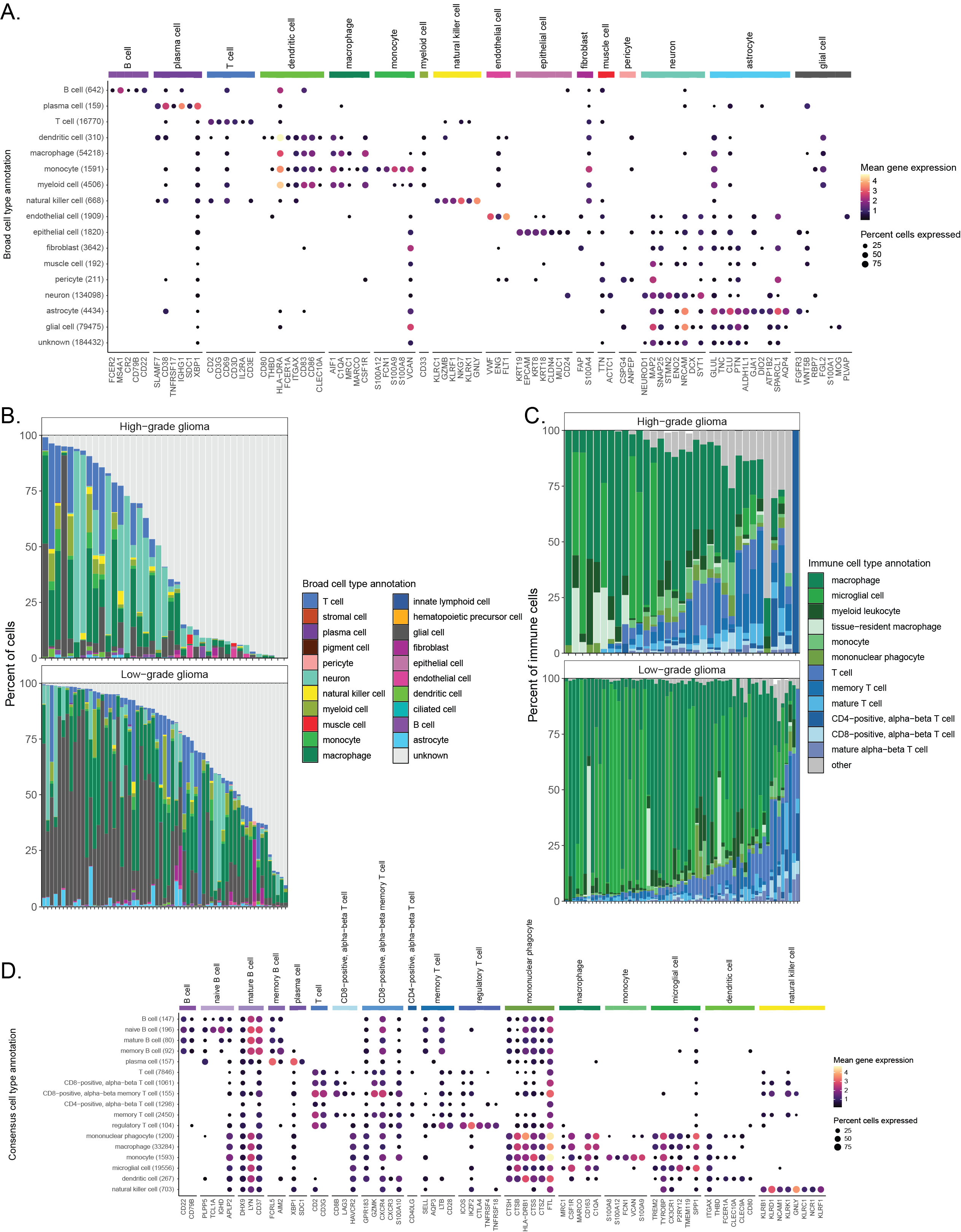
A. Dot plot showing expression of cell-type-specific marker genes across all libraries from brain and central nervous system (CNS) tumors, excluding multiplexed libraries.
102
102
Expression is shown for each broad cell type annotation, where each broad cell type annotation is a collection of similar consensus cell type annotations.
@@ -112,7 +112,7 @@ C. Barplot showing all consensus cell types classified as immune cells across li
112
112
The percentage shown corresponds to the percentage of immune cells classified as the indicated consensus cell type.
113
113
Only libraries comprised of at least 1\% immune cells, based on consensus cell type annotations, are shown.
114
114
Specific consensus cell types for myeloid and lymphocyte immune cells are shown, with all other consensus immune cell types included in "other."
115
-
Any myeloid or lymphocyte immune cell types with fewer than 1000 cells across all libraries are also include in "other".
115
+
Any myeloid or lymphocyte immune cell types with fewer than 1000 cells across all libraries are also included in "other".
116
116
117
117
D. Dot plot showing expression of cell-type-specific marker genes across all non-multiplexed libraries from brain and CNS tumors, considering only immune cells.
118
118
Expression is shown for each consensus cell type annotation.
@@ -125,7 +125,7 @@ Cell types without associated marker genes in `CellMarker 2.0` are not shown, in
125
125
<br><br>
126
126
127
127
<!-- Figure 5 -->
128
-
{#fig:fig5 tag="5" width="7in"}
128
+
{#fig:fig5 tag="5" width="7in"}
129
129
130
130
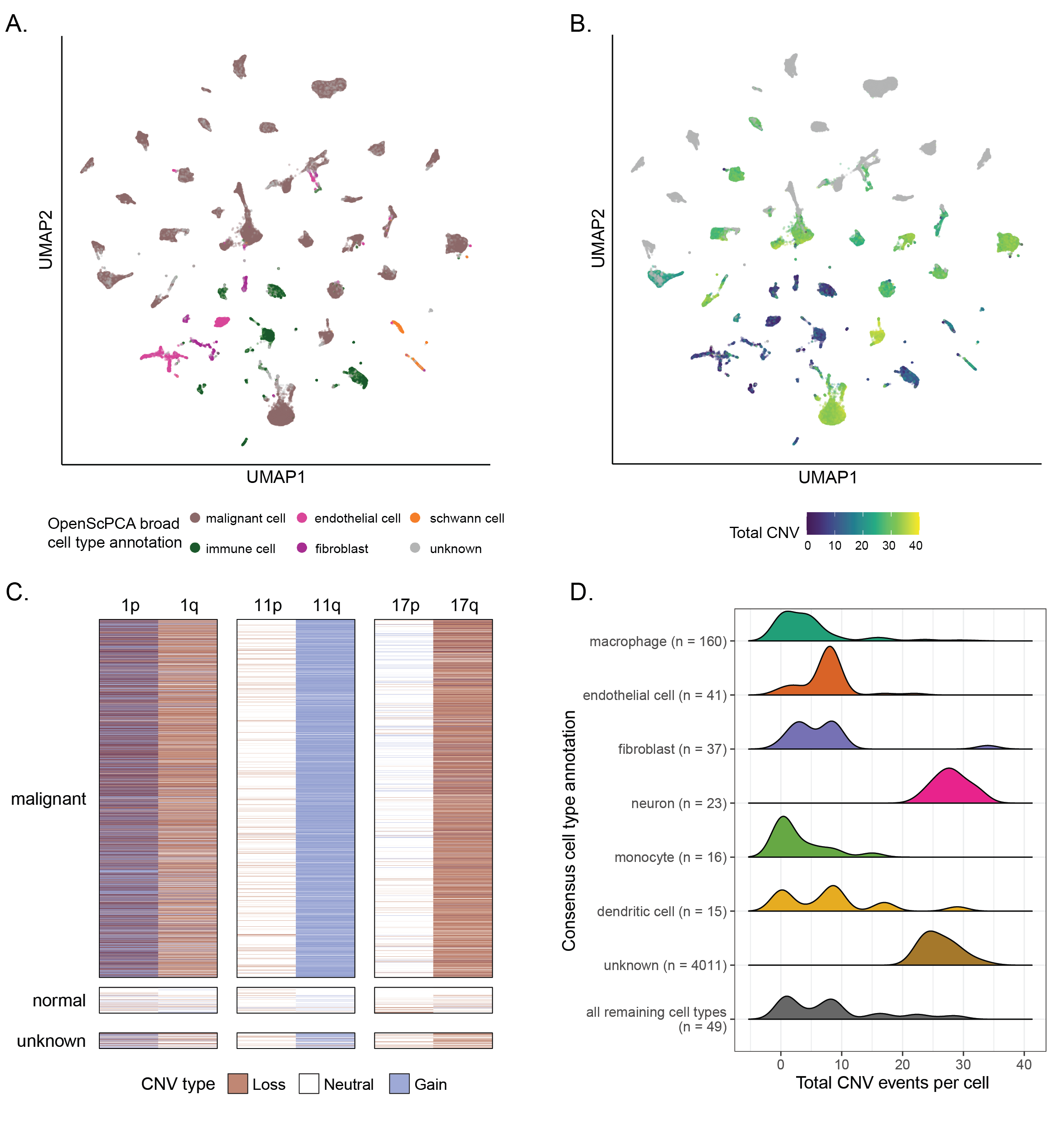
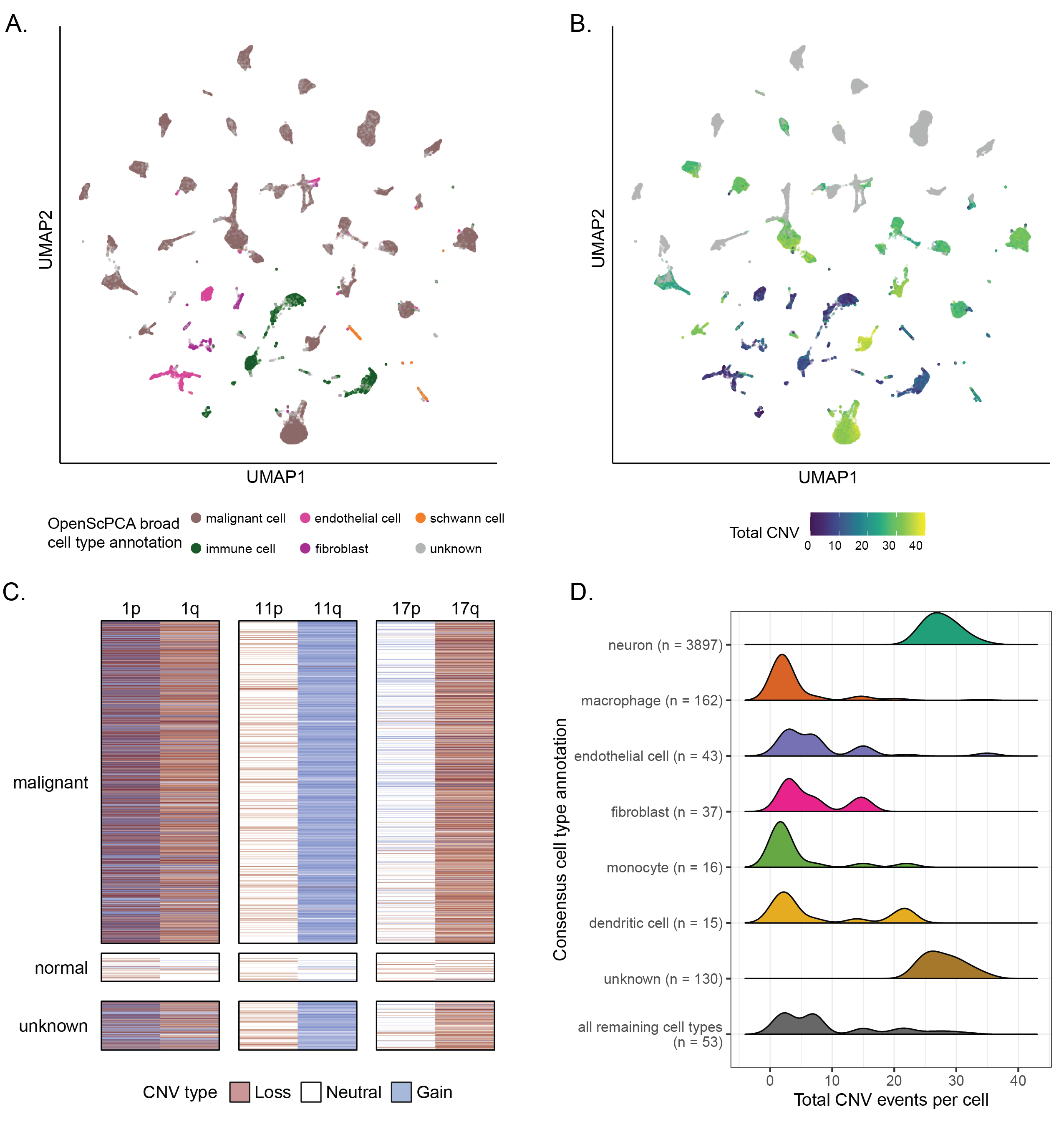
A. UMAP highlighting cell type annotations made with the OpenScPCA Project, collapsed into broad annotation groups, for all libraries in the neuroblastoma-only ScPCA Project `SCPCP000004` (N = 42).
131
131
The UMAP was constructed from the merged `SCPCP000004` object such that all libraries contribute an equal weight, but no batch correction was performed.
@@ -146,7 +146,7 @@ The ridge plot shown is for library `SCPCL000130`.
146
146
147
147
148
148
<!-- Figure 6 -->
149
-
{#fig:fig6 tag="6" width="7in"}
149
+
{#fig:fig6 tag="6" width="7in"}
150
150
151
151
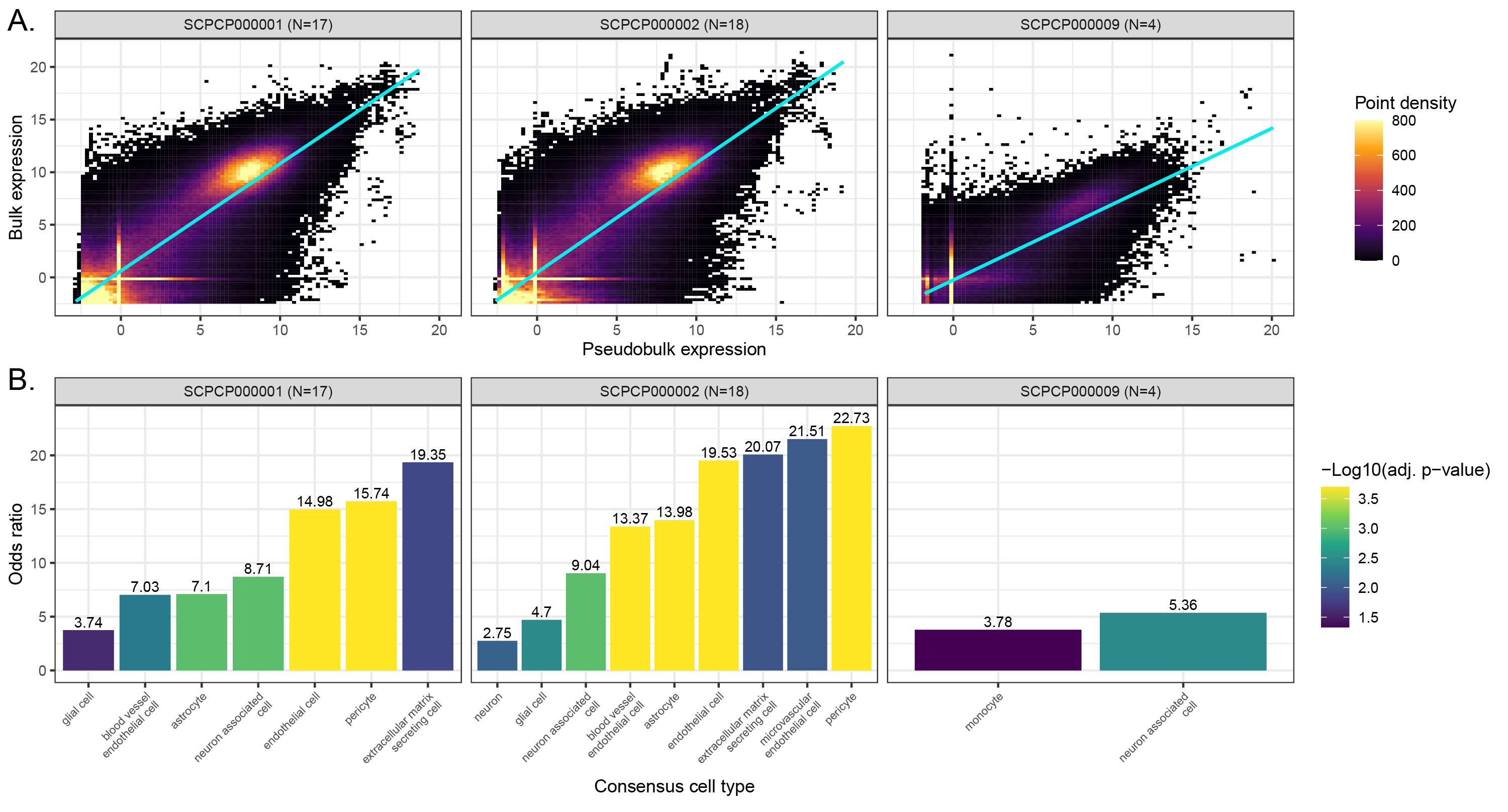
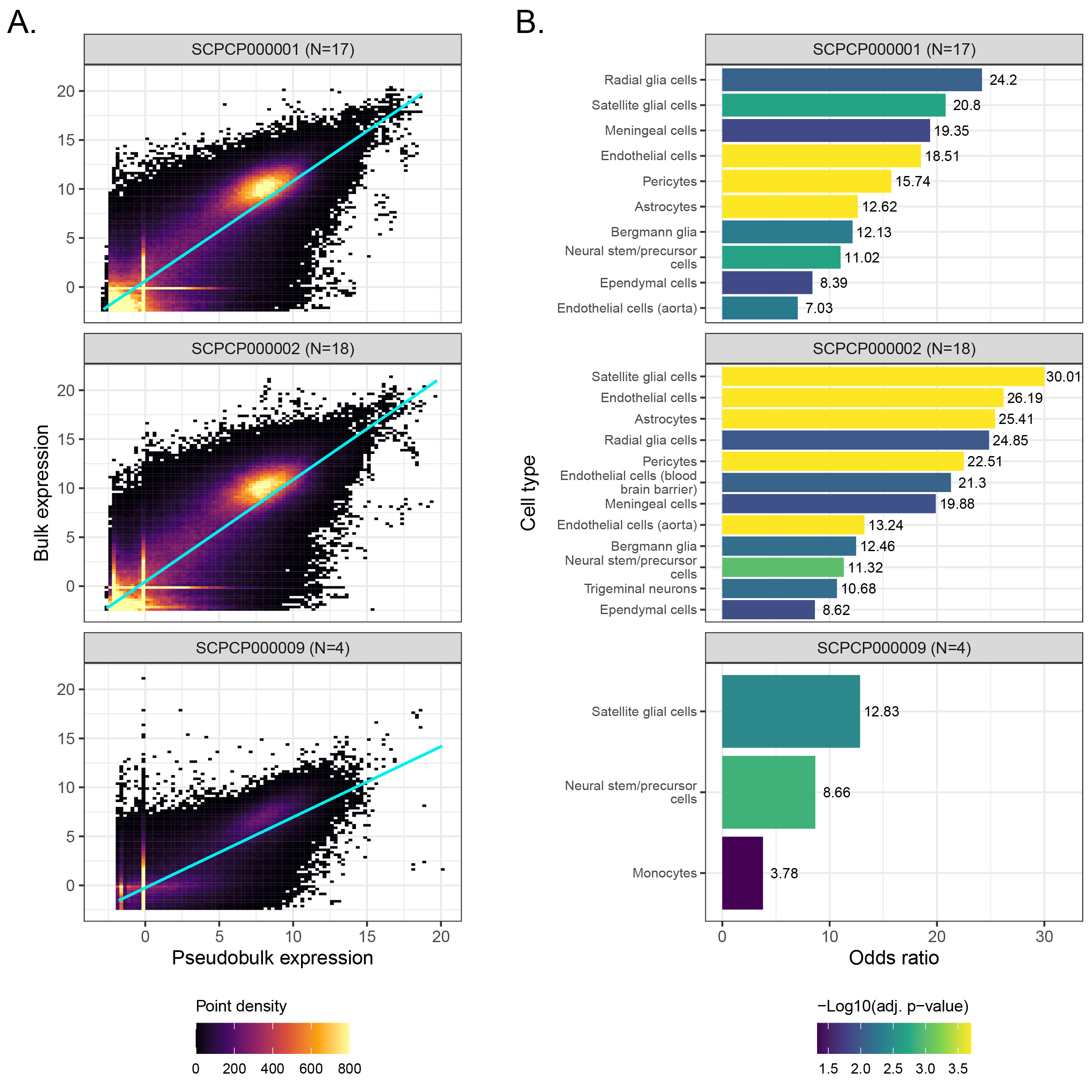
A. Scatter plots colored by point density of `DESeq2`-transformed and normalized bulk RNA-seq expression compared to pseudobulk expression from single-cell/nuclei RNA-seq.
152
152
Samples with RNA-seq for both bulk and single-cell/nuclei modalities, excluding multiplexed samples, from ScPCA projects comprising brain and central nervous system tumors are shown, with the number of samples considered per project shown in parentheses.
@@ -186,7 +186,7 @@ The reference includes marker genes for all cell types present in each organ.
186
186
<br><br>
187
187
188
188
<!-- Figure S1 -->
189
-
{#fig:figS1 tag="S1" width="7in"}
189
+
{#fig:figS1 tag="S1" width="7in"}
190
190
191
191
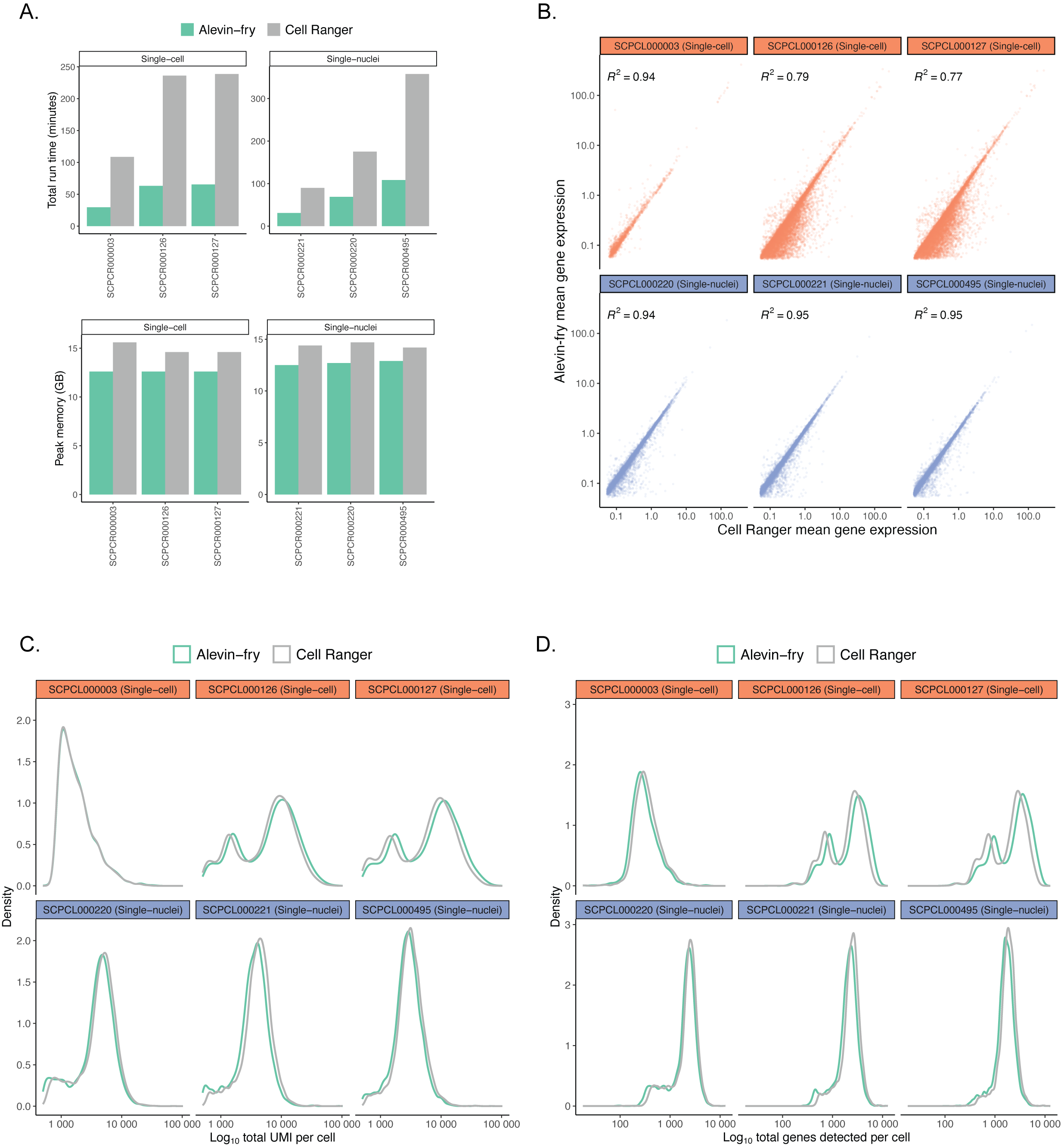
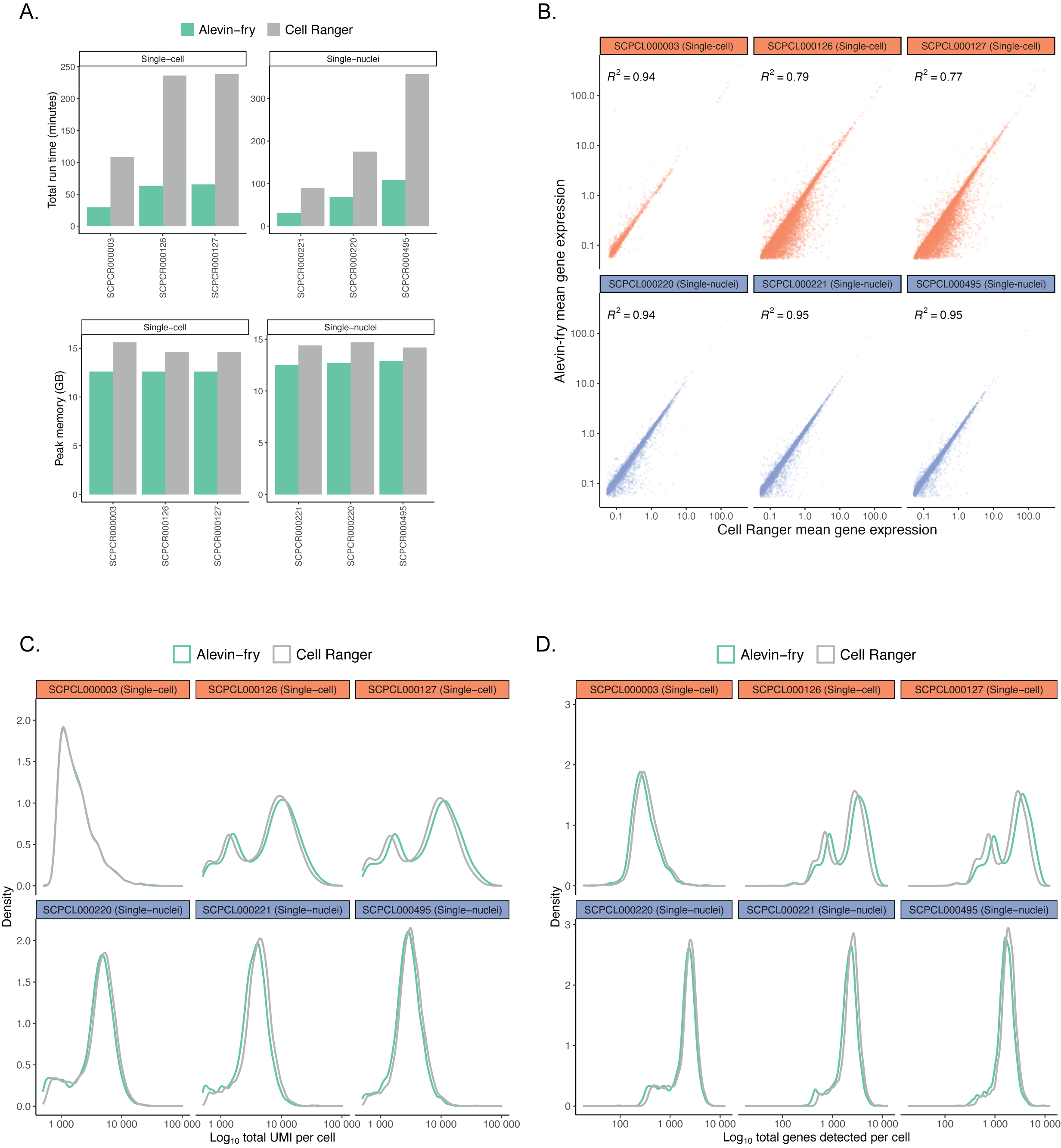
Each panel compares metrics for six ScPCA libraries, including three single-cell and three single-nuclei suspensions, obtained from processing libraries with `salmon alevin` and `alevin-fry` or `CellRanger`.
192
192
Results shown were generated with `CellRanger v6.1.2` using default parameters for single-cell libraries and use of the `--include_introns` flag to include intronic reads for single-nuclei libraries only.
@@ -211,7 +211,7 @@ Distributions reflect broad agreement between platforms in the total number of g
211
211
<br><br>
212
212
213
213
<!-- Figure S2 -->
214
-
{#fig:figS2 tag="S2" width="7in"}
214
+
{#fig:figS2 tag="S2" width="7in"}
215
215
216
216
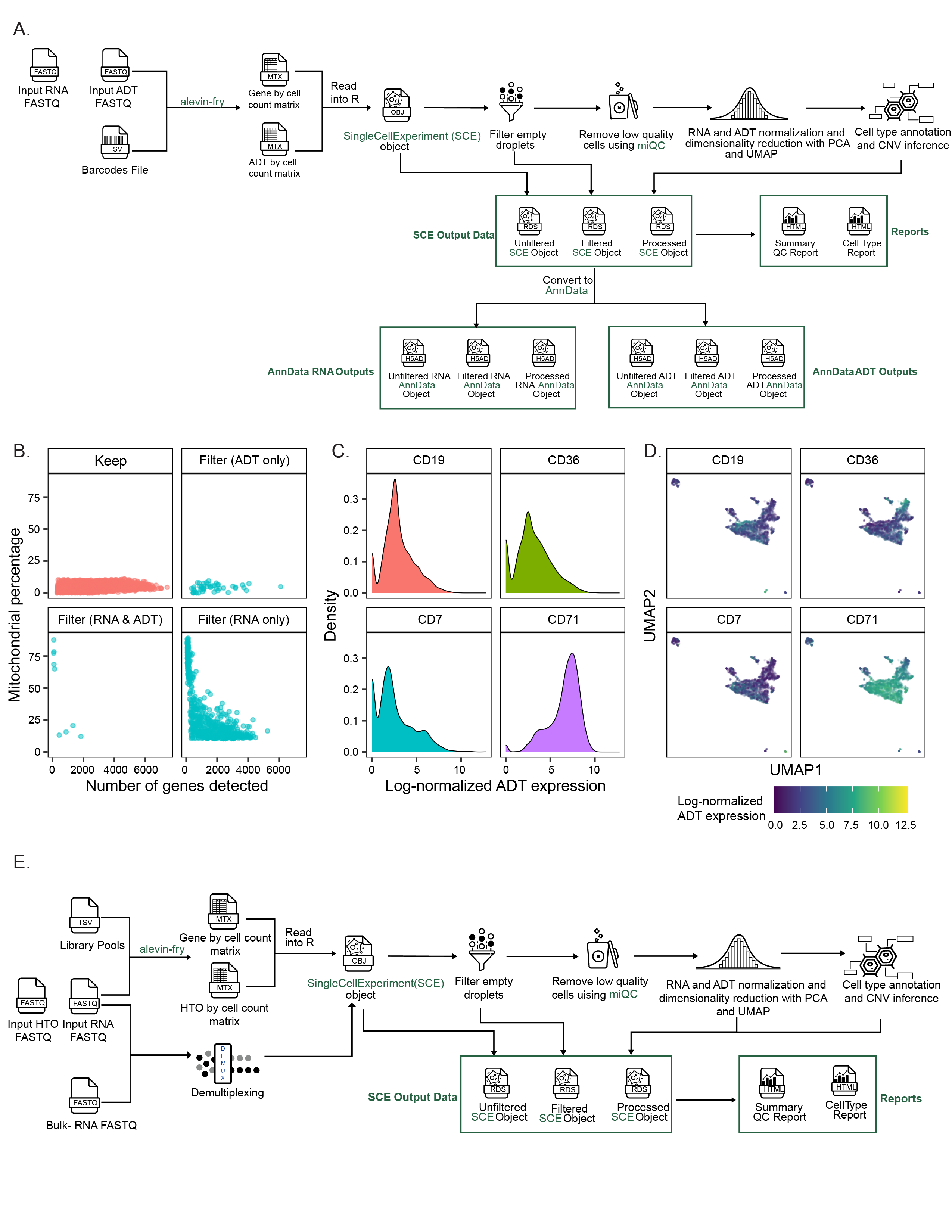
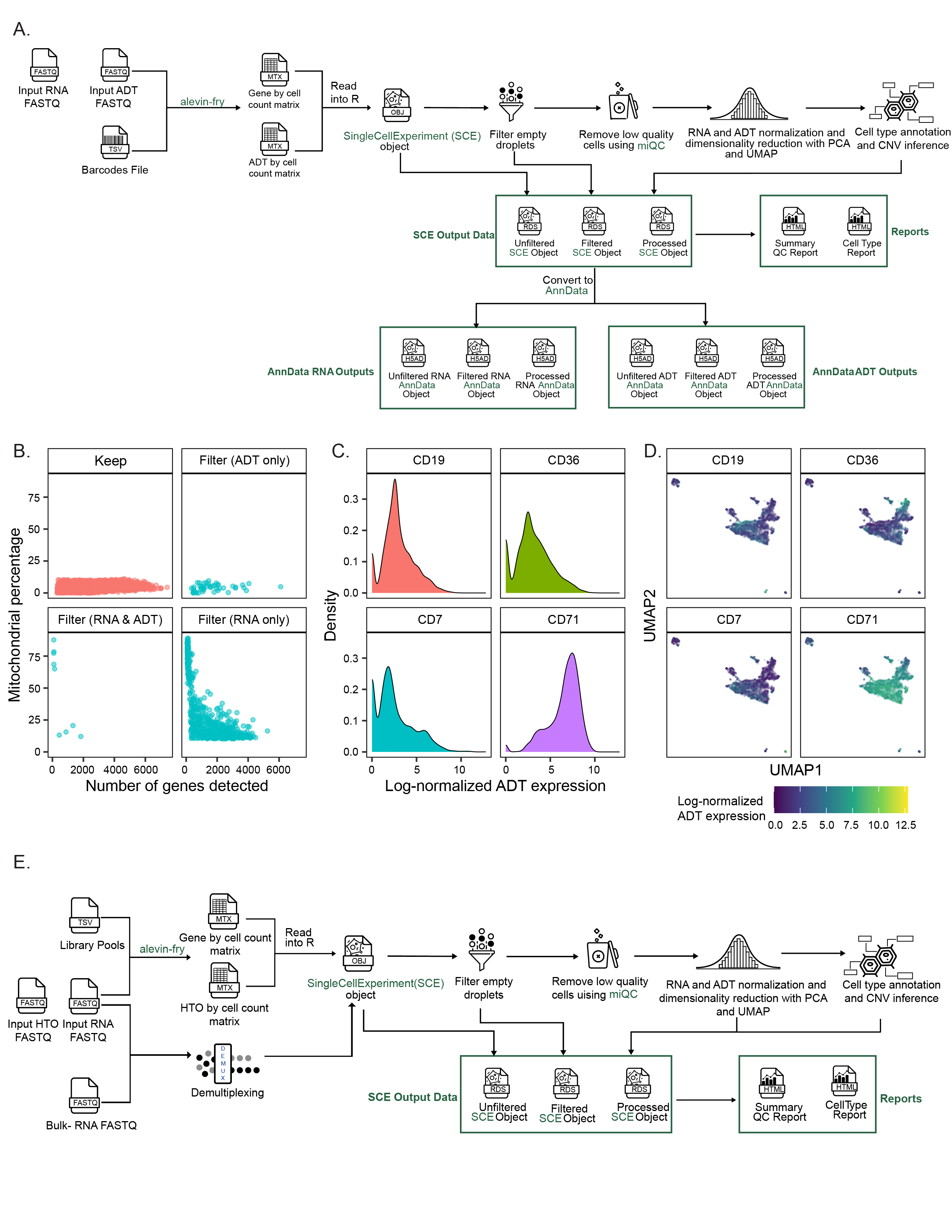
A. Overview of the `scpca-nf` workflow for processing libraries with CITE-seq or antibody-derived tag (ADT) data.
217
217
The workflow mirrors that shown in Figure {@fig:fig2}A with several differences accounting for the presence of ADT data.
@@ -245,7 +245,7 @@ Third, only `SCE` files are provided for multiplexed libraries; no corresponding
245
245
<br><br>
246
246
247
247
<!-- Figure S3 -->
248
-
{#fig:figS3 tag="S3" width="7in"}
248
+
{#fig:figS3 tag="S3" width="7in"}
249
249
250
250
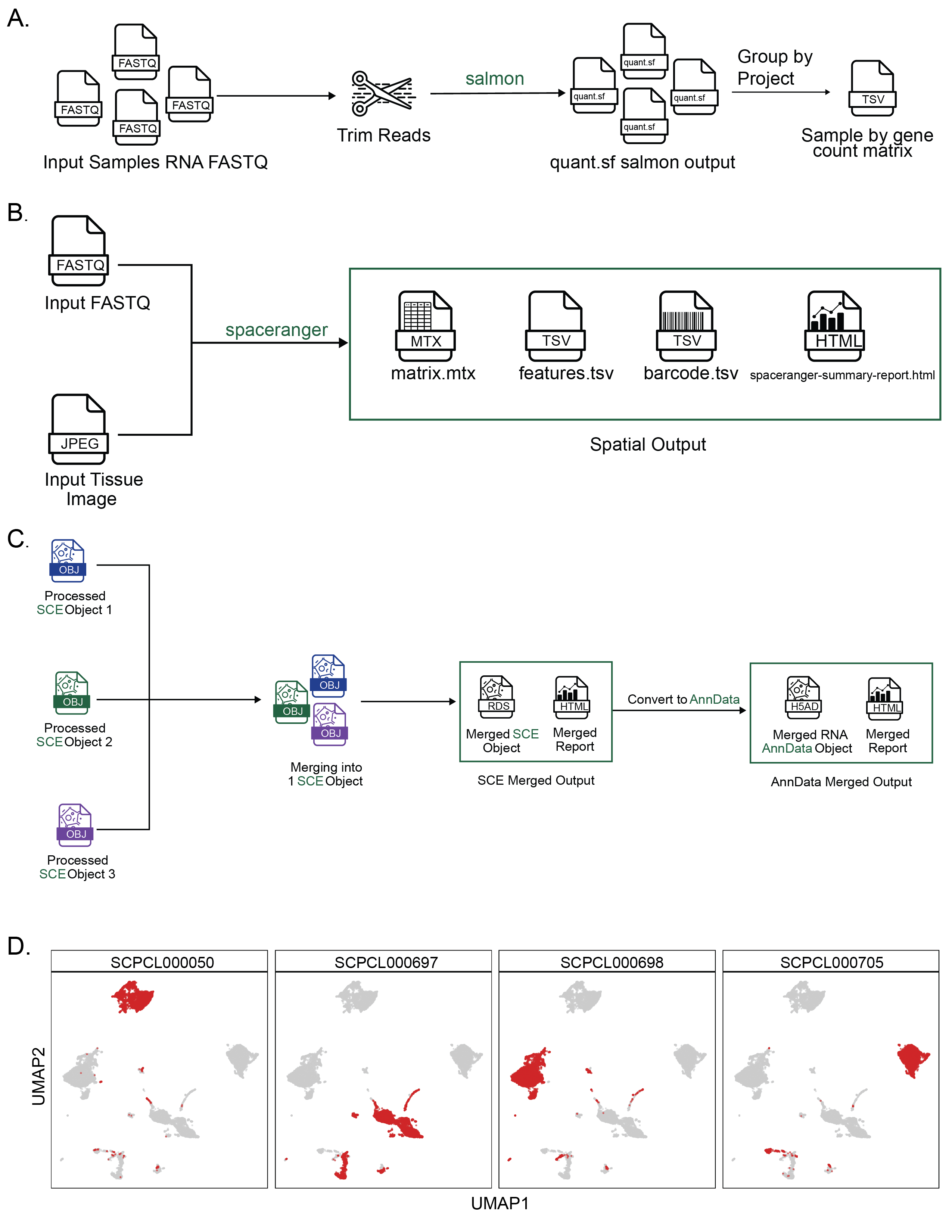
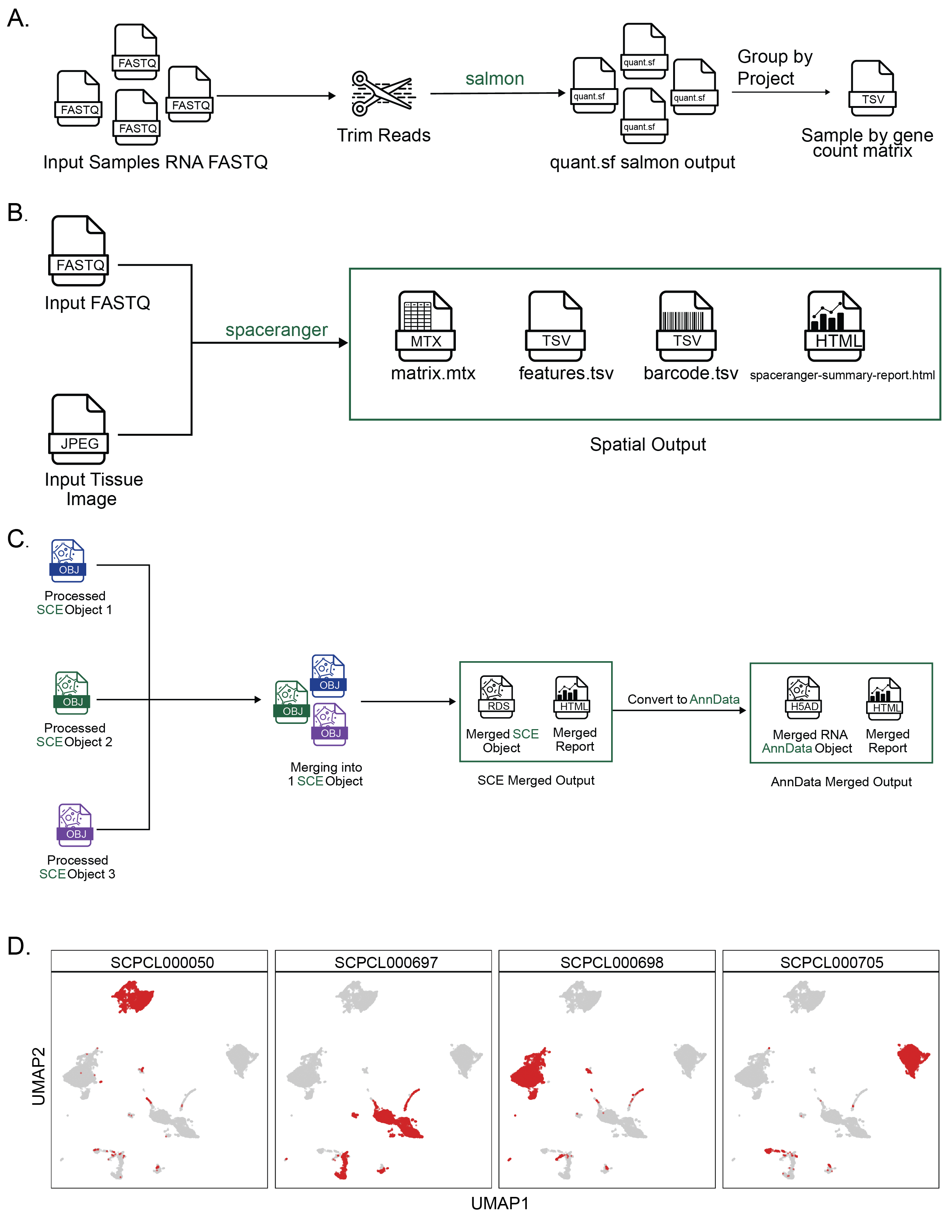
A. Overview of the bulk RNA-Seq workflow.
251
251
A set of FASTQ files from libraries sequenced with bulk RNA-seq are provided as input.
@@ -268,7 +268,7 @@ For this figure specifically, the merged UMAP was constructed from a merged obje
268
268
<br><br>
269
269
270
270
<!--Figure S4-->
271
-
{#fig:figS4 tag="S4" width="7in"}
271
+
{#fig:figS4 tag="S4" width="7in"}
272
272
273
273
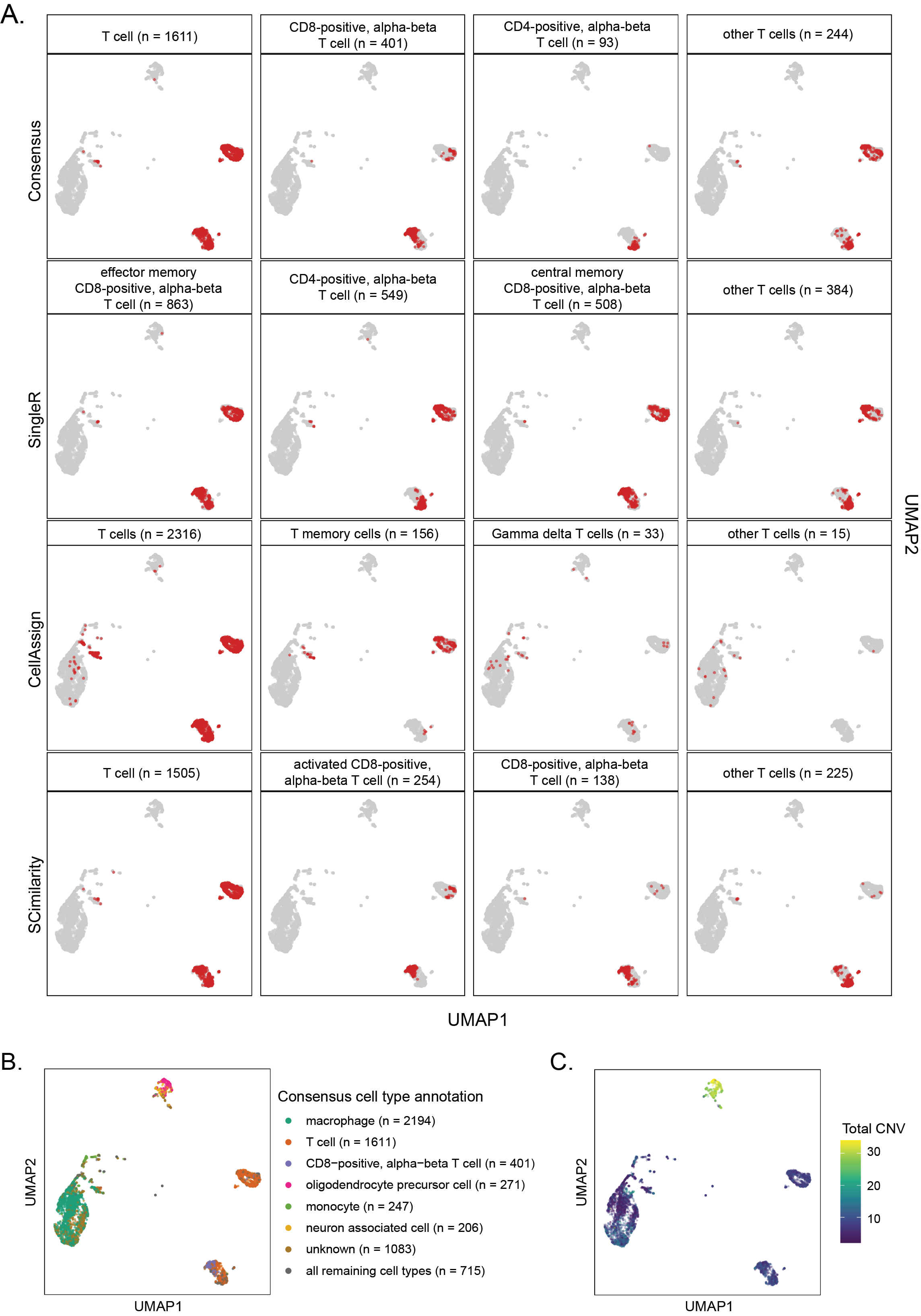
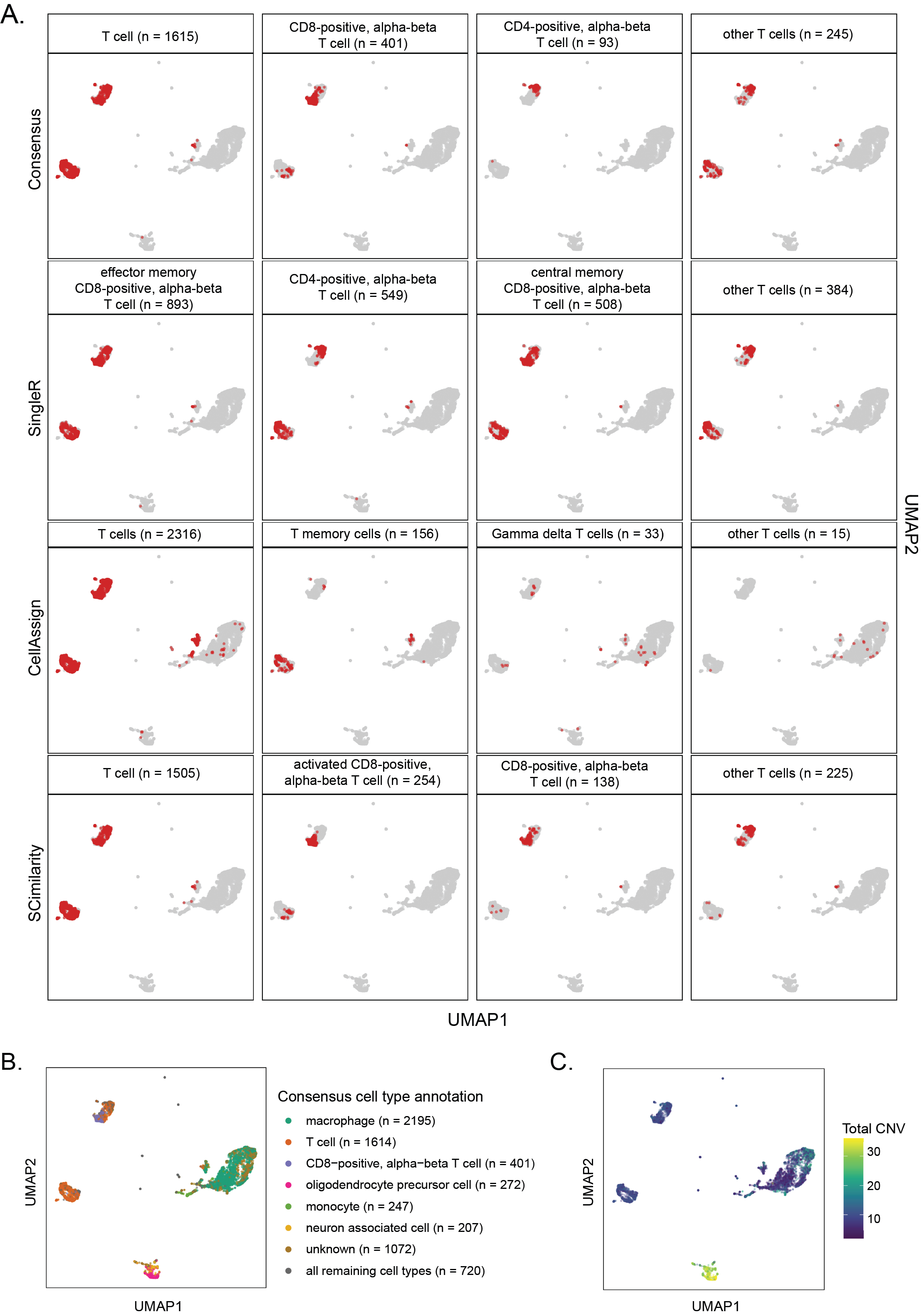
A. UMAP highlighting cells annotated as types of T cells with `SingleR`, `CellAssign`, `SCimilarity` as well as the associated consensus cell types for the library `SCPCL000049`.
274
274
All other cells are shown in gray.
@@ -283,7 +283,7 @@ The total per-cell CNV values were calculated by summing the total number of chr
283
283
284
284
285
285
<!-- Figure S5 -->
286
-
{#fig:figS5 tag="S5" width="9in"}
286
+
{#fig:figS5 tag="S5" width="9in"}
287
287
288
288
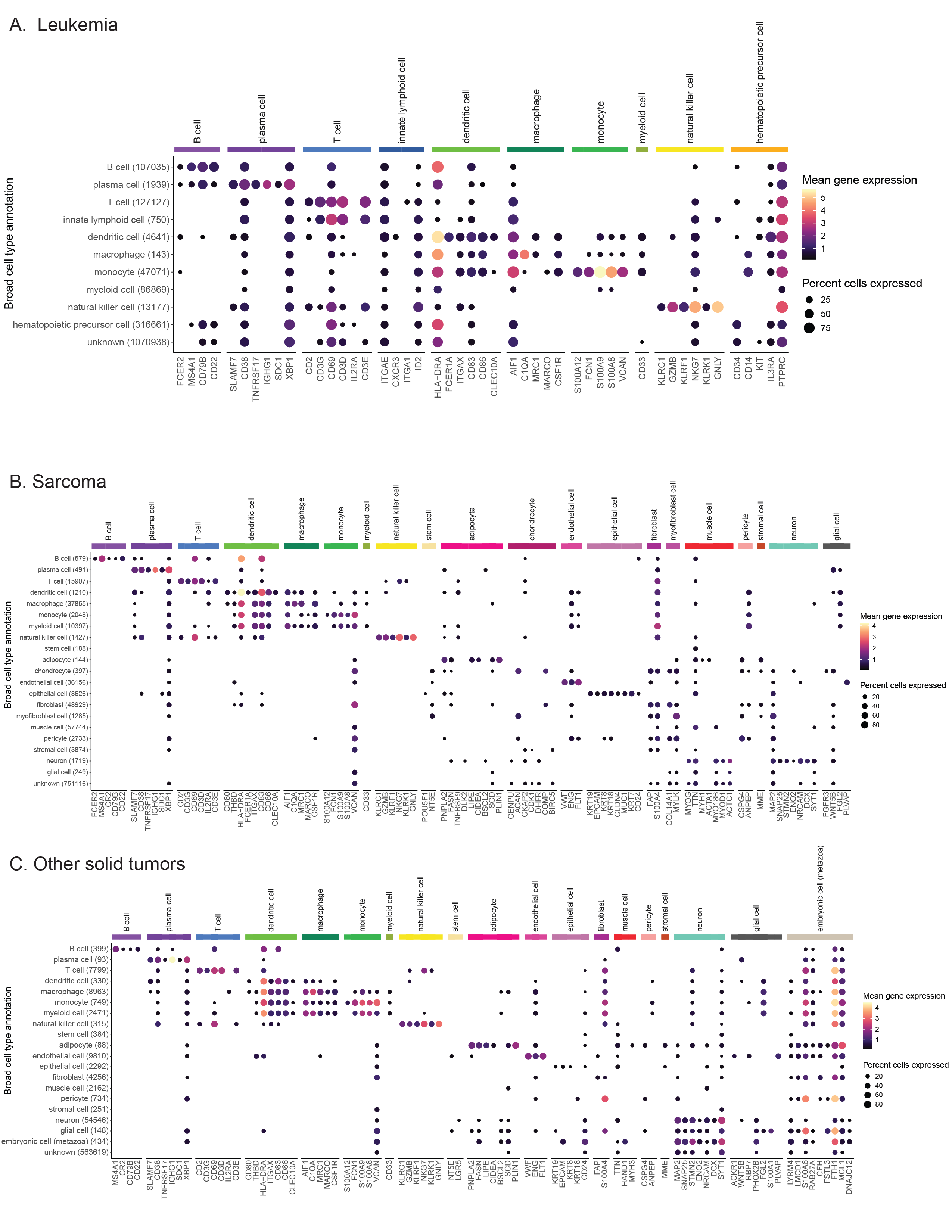
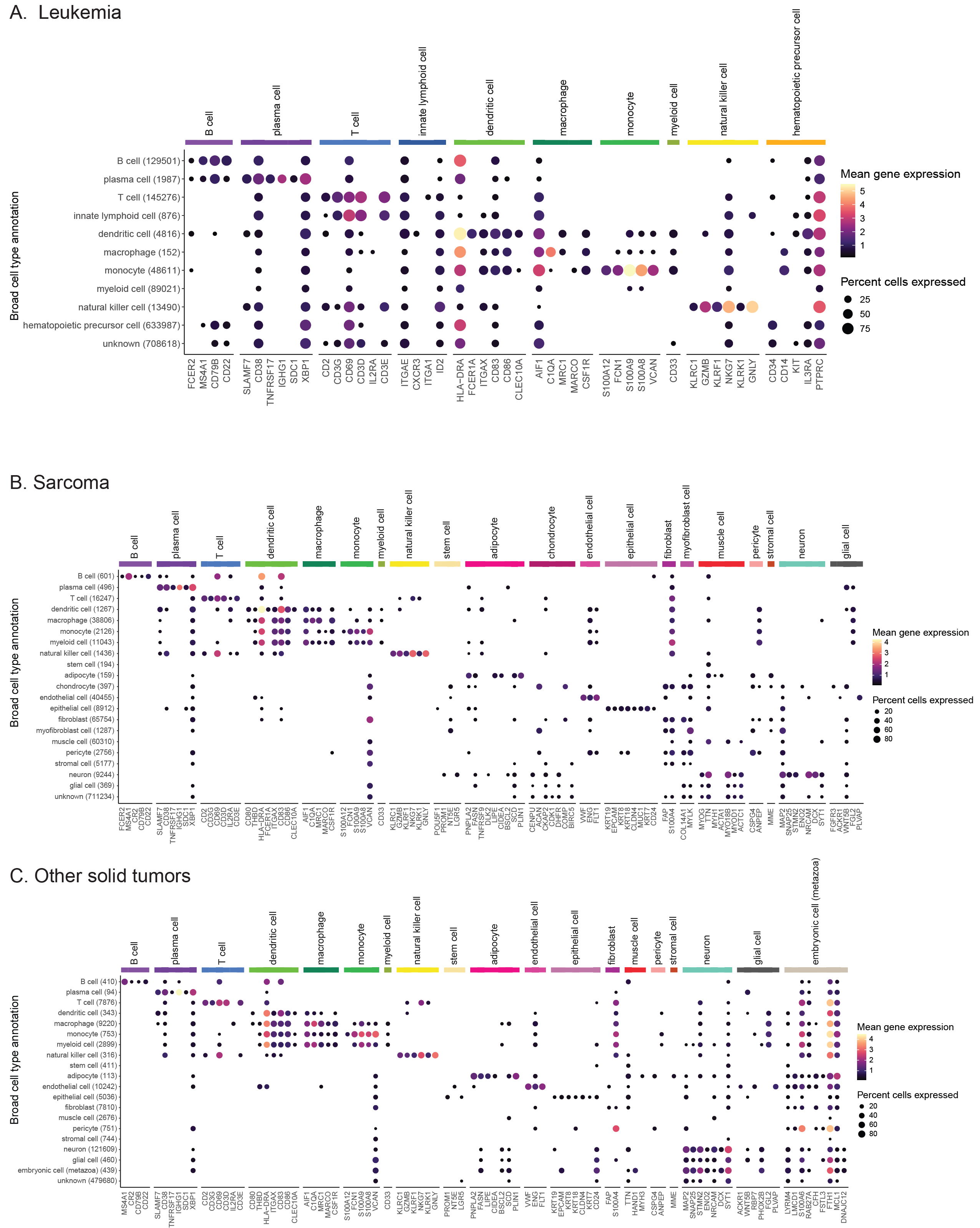
Dot plots showing expression of cell-type-specific marker genes across all libraries from Leukemia (A), Sarcoma (B), and Other solid tumors (C) diagnosis groups.
289
289
Expression is shown for each broad cell type annotation, where each broad cell type annotation is a collection of similar consensus cell type annotations.
@@ -296,7 +296,7 @@ Cell types without associated marker genes in `CellMarker 2.0` are not shown, in
296
296
<br><br>
297
297
298
298
<!-- Figure S6 -->
299
-
{#fig:figS6 tag="S6" width="7in"}
299
+
{#fig:figS6 tag="S6" width="7in"}
300
300
301
301
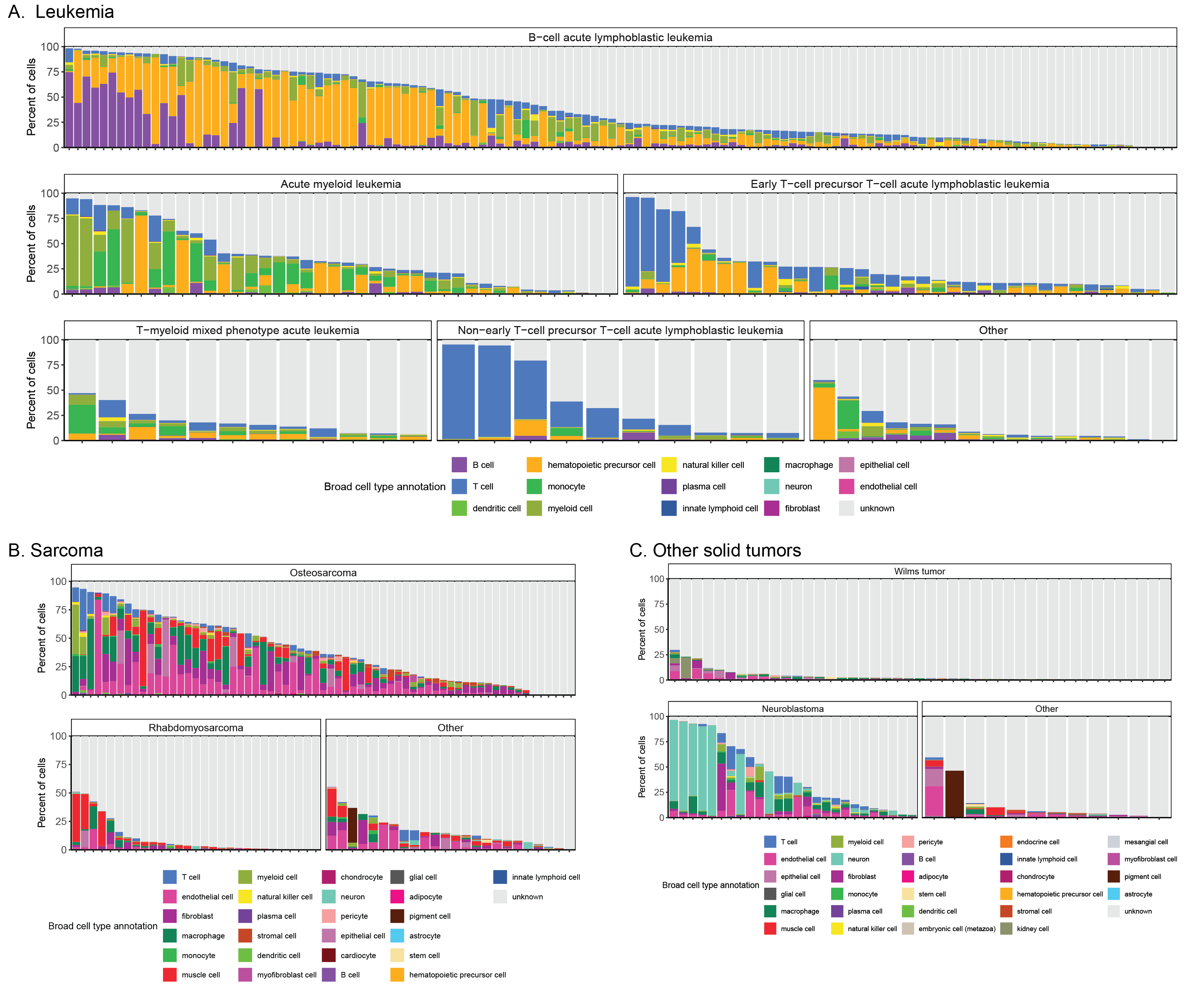
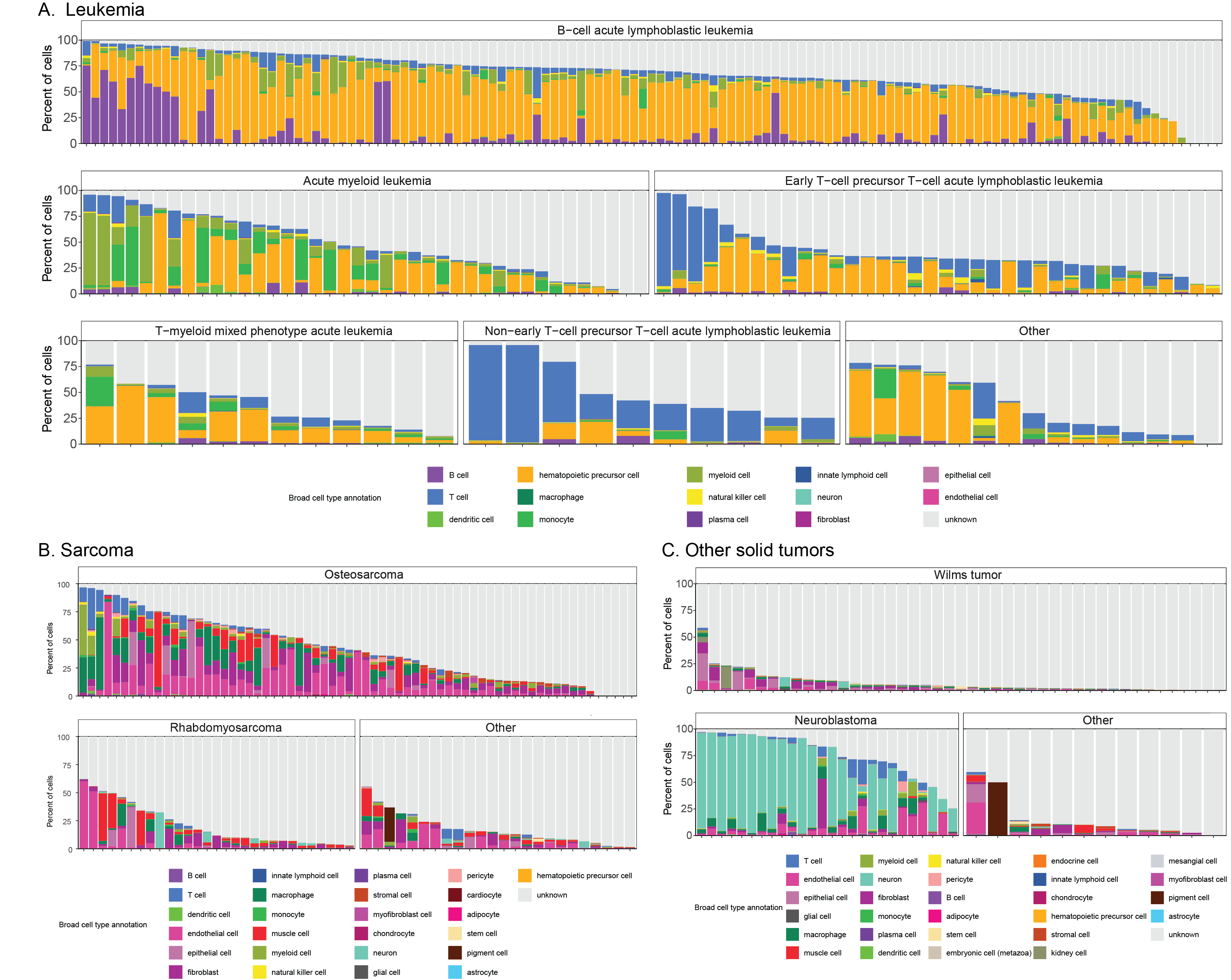
Barplots of the percentage of cells annotated as each broad consensus cell type annotation across all libraries from Leukemia (A), Sarcoma (B), and Other solid tumors (C) diagnosis groups.
302
302
Within each panel, libraries are shown grouped by diagnosis.
@@ -305,7 +305,7 @@ Only non-multiplexed libraries from patient tissue samples are shown.
305
305
<br><br>
306
306
307
307
<!-- Figure S7 -->
308
-
{#fig:figS7 tag="S7" width="7in"}
308
+
{#fig:figS7 tag="S7" width="7in"}
309
309
310
310
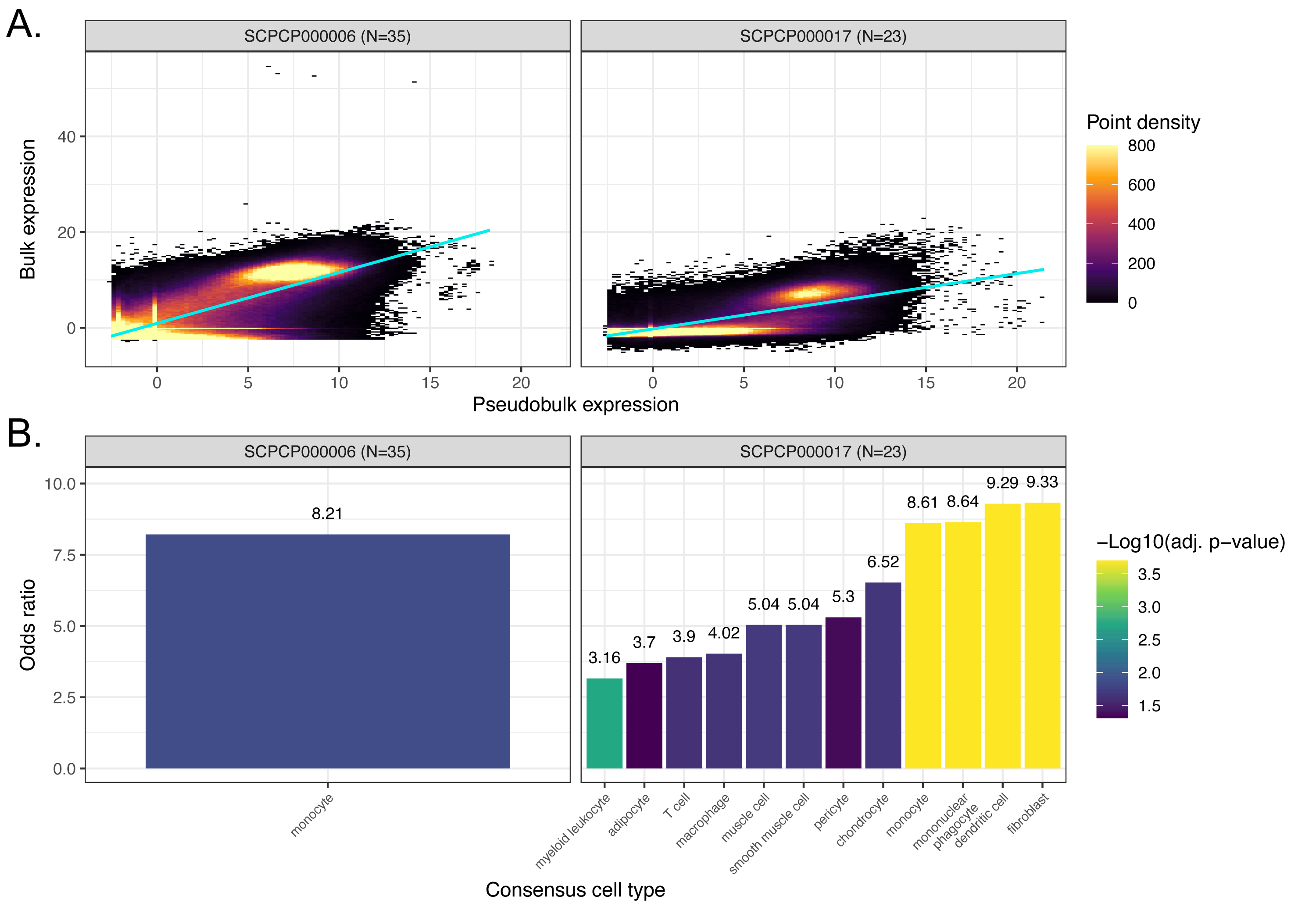
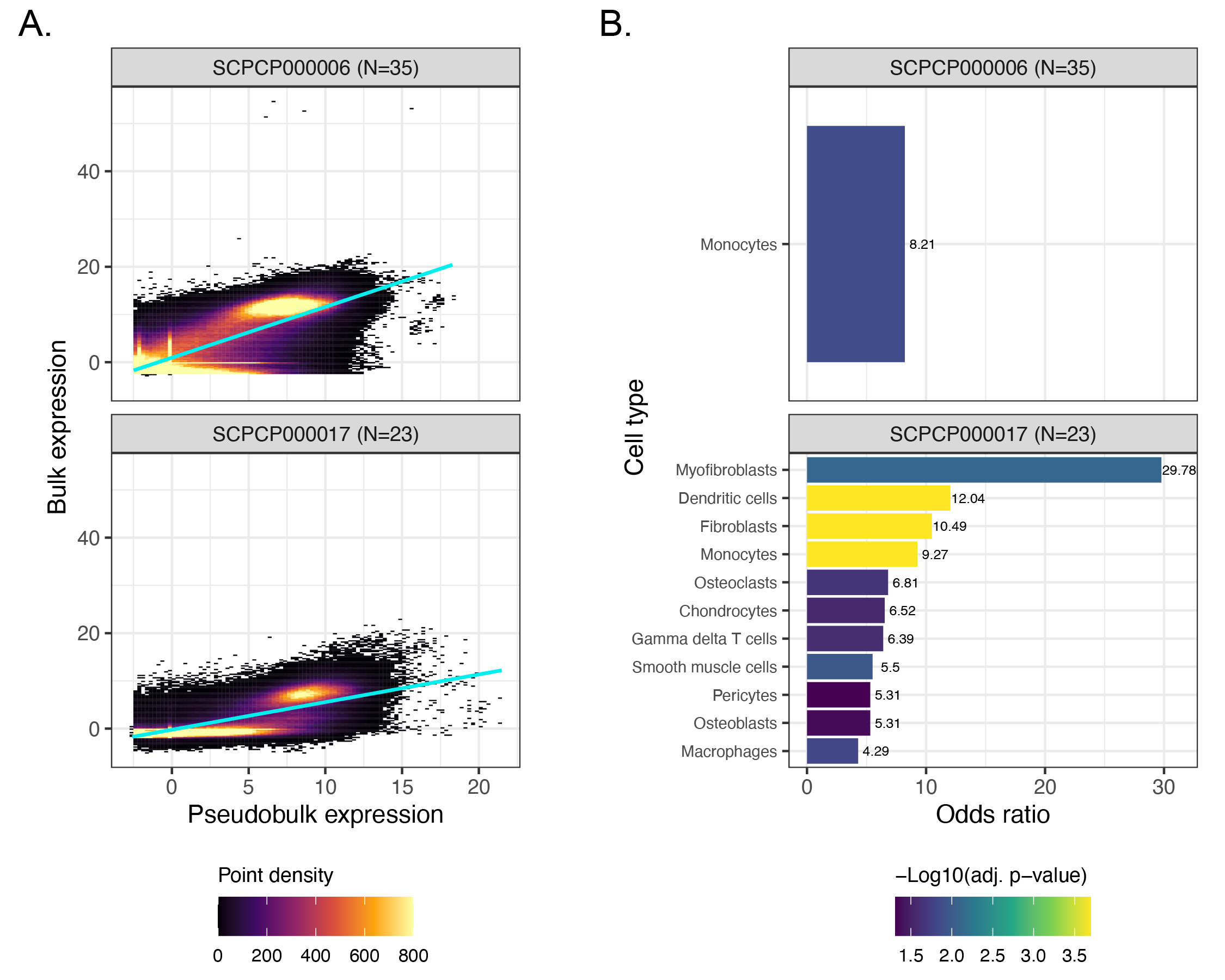
A. Scatter plots colored by point density of `DESeq2`-transformed and normalized bulk RNA-seq expression compared to pseudobulk expression from single-nuclei RNA-seq.
311
311
Projects with RNA-seq for both bulk and single-cell/nuclei modalities that are not displayed in Figure {@fig:fig6}A are shown.
0 commit comments