diff --git a/.cursor/rules/conda-environment.mdc b/.cursor/rules/conda-environment.mdc
new file mode 100644
index 0000000..ac4252c
--- /dev/null
+++ b/.cursor/rules/conda-environment.mdc
@@ -0,0 +1,10 @@
+---
+description:
+globs:
+alwaysApply: true
+---
+
+To run anything in the terminal or console, ALWAYS:
+1. Run: conda activate confopt_env
+2. Run: pip install .
+3. Run your command
diff --git a/.cursor/rules/python-best-practice-instructions.mdc b/.cursor/rules/python-best-practice-instructions.mdc
new file mode 100644
index 0000000..ed8187a
--- /dev/null
+++ b/.cursor/rules/python-best-practice-instructions.mdc
@@ -0,0 +1,25 @@
+---
+description:
+globs:
+alwaysApply: true
+---
+# Coding Style Guidelines
+
+- Adopt the DRY principle. If code is repeated in multiple places, it should be functionalized and called in those places.
+- Make all inputs explicit. Avoid relying on state or shared context unless encapsulated.
+- Avoid implicit behavior (e.g., mutation of input lists, in place dataframe modification).
+- Variable names should be descriptive and reduced to the shortest possible length.
+- If a function returns multiple types, refactor. Don't return Union[str, dict, None].
+- Use pytest.mark.parametrize for testing functions with categorical input values.
+- If mocking in unit testing is required, mock external APIs and I/O only.
+- No print() statements anywhere in the code. Use logging instead.
+- No single-letter variable names unless in mathematical contexts or loops.
+- No hard coded values. Use constants or configuration files.
+- Don't use early returns in if else statements.
+- Don't create classes if functions are sufficient.
+- Keep modules small and focused (under 500 lines).
+- Comments should explain why, not what. Keep comments under 10% of code.
+- Don't write docstrings.
+- Don't rely on default values for function arguments.
+- Avoid *args or **kwargs unless absolutely necessary.
+- Use pydantic models for configuration values.
diff --git a/.cursor/rules/software-engineering-best-practices.mdc b/.cursor/rules/software-engineering-best-practices.mdc
new file mode 100644
index 0000000..df86087
--- /dev/null
+++ b/.cursor/rules/software-engineering-best-practices.mdc
@@ -0,0 +1,10 @@
+---
+description:
+globs:
+alwaysApply: true
+---
+- Always comply with DRY and SOLID principles.
+- Use as little code as is necessary to carry out the desired functionality, do not over-engineer or over-validate your code.
+- Write easily testable and maintainable code.
+- Maximize separation of concerns.
+- Consider how your changes will affect the wider codebase, think several dependancies ahead.
diff --git a/.cursor/rules/system-permissions.mdc b/.cursor/rules/system-permissions.mdc
new file mode 100644
index 0000000..56139fb
--- /dev/null
+++ b/.cursor/rules/system-permissions.mdc
@@ -0,0 +1,8 @@
+---
+description:
+globs:
+alwaysApply: true
+---
+- NEVER commit any changes.
+- NEVER revert commits or affect the commit history.
+- NEVER push commits or pull from remote, or interact at all with remote branches.
diff --git a/.github/copilot-instructions.md b/.github/copilot-instructions.md
new file mode 100644
index 0000000..bc10dcb
--- /dev/null
+++ b/.github/copilot-instructions.md
@@ -0,0 +1,31 @@
+# Coding Style Guidelines
+
+- Adopt the DRY principle. If code is repeated in multiple places, it should be functionalized and called in those places.
+- Make all inputs explicit. Avoid relying on state or shared context unless encapsulated.
+- Avoid implicit behavior (e.g., mutation of input lists, in place dataframe modification).
+- Variable names should be descriptive and reduced to the shortest possible length.
+- If a function returns multiple types, refactor. Don't return Union[str, dict, None].
+- Use pytest.mark.parametrize for testing functions with categorical input values.
+- If mocking in unit testing is required, mock external APIs and I/O only.
+- No print() statements anywhere in the code. Use logging instead.
+- No single-letter variable names unless in mathematical contexts or loops.
+- No hard coded values. Use constants or configuration files.
+- Don't use early returns in if else statements.
+- Don't create classes if functions are sufficient.
+- Keep modules small and focused (under 500 lines).
+- Comments should explain why, not what. Keep comments under 10% of code.
+- Don't write docstrings.
+- Don't rely on default values for function arguments.
+- Avoid *args or **kwargs unless absolutely necessary.
+- Use pydantic models for configuration values.
+- Always comply with DRY and SOLID principles.
+- Use as little code as is necessary to carry out the desired functionality, do not over-engineer or over-validate your code.
+- Write easily testable and maintainable code.
+- Maximize separation of concerns.
+- Consider how your changes will affect the wider codebase, think several dependancies ahead.
+
+
+To run anything in the terminal or console, ALWAYS:
+1. Run: conda activate confopt_env
+2. Run: pip install .
+3. Run your command
diff --git a/.github/documentation-instructions.md b/.github/documentation-instructions.md
new file mode 100644
index 0000000..b43b5a4
--- /dev/null
+++ b/.github/documentation-instructions.md
@@ -0,0 +1,95 @@
+# Quantile Estimation Module Documentation Template
+
+This template provides a step-by-step guide and example prompt for documenting any Python module (e.g., `quantile_estimation.py`) in a style consistent with best practices for technical and developer documentation.
+
+---
+
+## 1. Docstring Requirements
+
+Add or update detailed and informative Google-style docstrings following these guidelines:
+
+### Module-level docstring:
+- Brief description of the module's purpose and core functionality
+- Key methodological approaches or architectural patterns used
+- Integration context within the broader framework
+- Focus on salient aspects, avoid trivial descriptions
+- Do not add any type hints in the doc strings.
+
+---
+
+## 2. Documentation File Requirements
+
+Create a comprehensive `.rst` documentation file in `docs/developer/components/[module_name].rst` with:
+
+### Structure Example:
+
+```
+[Module Name] Module
+===================
+
+Overview
+--------
+[Brief description and key features]
+
+Key Features
+------------
+[Bullet points of main capabilities]
+
+Architecture
+------------
+[Class hierarchy, design patterns, architectural decisions]
+
+[Methodology/Algorithm Sections]
+-------------------------------
+[Detailed explanations of key approaches, mathematical foundations where relevant]
+
+Usage Examples
+--------------
+[Practical code examples showing common usage patterns]
+
+Performance Considerations
+-------------------------
+[Computational complexity, scaling considerations, best practices]
+
+Integration Points
+-----------------
+[How this module connects with other framework components]
+
+Common Pitfalls
+---------------
+[Common mistakes and how to avoid them]
+
+See Also
+--------
+[Cross-references to related modules]
+```
+
+### Content requirements:
+- Technical depth appropriate for developers
+- Mathematical foundations with LaTeX equations where relevant (cite relevant papers if mainstream, do not hallucinate, if unsure do not cite any)
+- Practical usage examples with actual code
+- Performance and scalability guidance
+- Integration context within the framework
+- Best practices and common pitfalls
+- Cross-references to related components
+
+---
+
+## 3. Index Update
+
+Update `docs/developer/components/index.rst` to include the new module documentation in the appropriate section.
+
+---
+
+## 5. Best Practices
+- Documentation should be contextually relevant and technically accurate
+- Focus on methodology and implementation details that matter to developers
+- Provide both theoretical understanding and practical guidance
+- Ensure consistency with existing documentation style and organization
+- Make it easy for both newcomers and experienced developers to understand and use the module
+
+---
+
+## 6. Example Output (for quantile_estimation.py)
+
+See the current `quantile_estimation.py` for a fully documented example, and `docs/developer/components/quantile_estimation.rst` for a comprehensive documentation file.
diff --git a/.github/testing-instructions.md b/.github/testing-instructions.md
new file mode 100644
index 0000000..6dfdd50
--- /dev/null
+++ b/.github/testing-instructions.md
@@ -0,0 +1,22 @@
+# Coding Style Guidelines
+
+- Use pytest for all testing, use unittest for mocking.
+- Use pytest.mark.parametrize for testing functions with categorical input values:
+ - For literals, you should automatically cycle through all possible Literal values in your parametrization.
+ - For ordinal categories or discrete inputs (eg. n_observations, n_recommendations, etc.) pick some sensible ranges (eg. 0 if allowed, or minimum otherwise, then a sensible every day value, say 10, then a very large value, if it's not computationally expensive, say 1000).
+- Mock external APIs and I/O only, do not use mocking as a crutch to abstract away components who's behaviour you need to test.
+- Use fixtures to store toy data, mocked objects or any other object you plan to reference in the main tests, particularly if it will be used more than once. If the toy data is small and specific to the one test it's called in, it's ok to define it inside the test function.
+- Never define nested functions (function def is inside another function) unless explicitly required because of scope (eg. nested generator builders).
+- Avoid defining helper functions at the top of a test module, tests should be simple and mostly check existing methods' outputs. Very complex tests may require helper functions, but this should be limited.
+- ALL fixtures need to be defined in the tests/conftest.py file, NEVER define them directly in a test module.
+- Do not test initialization of classes. Do not use asserts that just check if an attribute of a class exists, or is equal to what you just defined it as, these are bloated tests that accomplish little, but add maintenance cost.
+- If you're testing a function or method that returns a shaped object, always check the shape (should it be the same as the input's? Should it be different? Should it be a specific size based on the inputs you passed to the function? etc. based on these questions formulate asserts that check those shape aspects)
+- Test the intent behind a function or method, not form or attributes. Read through the function or method carefully, understand its goals and approach, then write meaningful tests that check quality of outputs relative to intent.
+- Do not add strings after asserts, eg. do NOT do this:
+ assert len(final_alphas) == len(initial_alphas), "Alpha count should remain consistent"
+ after any assert statement, it should just be assert len(final_alphas) == len(initial_alphas)
+- Keep comments to a minimum, comments should just explain more obscure asserts or tests.
+- Each unit test should be a function, functions should not be grouped in testing classes and should not have self attributes.
+- When testing mathematical functions, understand the derivations and test assumptions and outputs given mathematical constraints and theory.
+- Do not write excessive amounts of tests, focus on the most important aspects of each function.
+- Avoid lenghty code repetition. If multiple tests share the same set ups or fixture processing but only differ in asserts, join them in a single test and add comments before each assert.
diff --git a/.github/workflows/ci-cd.yml b/.github/workflows/ci-cd.yml
new file mode 100644
index 0000000..7b1cf84
--- /dev/null
+++ b/.github/workflows/ci-cd.yml
@@ -0,0 +1,563 @@
+name: CI/CD Pipeline
+
+on:
+ push:
+ branches: [ '**' ]
+ pull_request:
+ types: [closed]
+ branches: [main]
+
+# Cancel in-progress workflows when a new commit is pushed
+concurrency:
+ group: ${{ github.workflow }}-${{ github.ref }}
+ cancel-in-progress: true
+
+env:
+ PYTHON_VERSION: "3.11"
+
+jobs:
+ test:
+ name: Test Suite
+ runs-on: ubuntu-latest
+ strategy:
+ fail-fast: false
+ matrix:
+ python-version: ["3.9", "3.10", "3.11", "3.12"]
+
+ steps:
+ - name: Checkout code
+ uses: actions/checkout@v4
+
+ - name: Set up Python ${{ matrix.python-version }}
+ uses: actions/setup-python@v5
+ with:
+ python-version: ${{ matrix.python-version }}
+
+ - name: Cache pip dependencies
+ uses: actions/cache@v4
+ with:
+ path: ~/.cache/pip
+ key: pip-${{ runner.os }}-${{ matrix.python-version }}-${{ hashFiles('**/pyproject.toml', '**/requirements*.txt') }}
+ restore-keys: |
+ pip-${{ runner.os }}-${{ matrix.python-version }}-
+
+ - name: Install dependencies
+ run: |
+ python -m pip install --upgrade pip
+ pip install -e ".[dev]"
+ pip install build twine
+
+ - name: Run tests
+ run: |
+ pytest tests/ -v --tb=short --junitxml=test-results-${{ matrix.python-version }}.xml -m "not slow"
+
+ - name: Upload test results
+ uses: actions/upload-artifact@v4
+ if: always()
+ with:
+ name: test-results-${{ matrix.python-version }}
+ path: test-results-${{ matrix.python-version }}.xml
+ retention-days: 2
+
+ lint:
+ name: Code Quality
+ runs-on: ubuntu-latest
+
+ steps:
+ - name: Checkout code
+ uses: actions/checkout@v4
+
+ - name: Set up Python
+ uses: actions/setup-python@v5
+ with:
+ python-version: ${{ env.PYTHON_VERSION }}
+
+ - name: Cache pre-commit
+ uses: actions/cache@v4
+ with:
+ path: ~/.cache/pre-commit
+ key: pre-commit-${{ runner.os }}-${{ hashFiles('.pre-commit-config.yaml') }}
+
+ - name: Install pre-commit
+ run: |
+ python -m pip install --upgrade pip
+ pip install pre-commit
+
+ - name: Run pre-commit hooks
+ run: pre-commit run --all-files
+
+ check-package-label:
+ name: Check Package Label
+ runs-on: ubuntu-latest
+ if: github.event_name == 'pull_request' && github.event.pull_request.merged == true
+ outputs:
+ has_package_label: ${{ steps.check_label.outputs.has_label }}
+ pr_number: ${{ github.event.pull_request.number }}
+
+ steps:
+ - name: Check for Package label
+ id: check_label
+ uses: actions/github-script@v7
+ with:
+ script: |
+ const labels = context.payload.pull_request.labels.map(label => label.name);
+ const has_package_label = labels.includes('package');
+
+ console.log('PR Labels:', labels);
+ console.log('Has package label:', has_package_label);
+
+ core.setOutput('has_label', has_package_label);
+
+ if (!has_package_label) {
+ console.log('⏭️ Skipping package deployment - no Package label found');
+ } else {
+ console.log('✅ Package label found - proceeding with deployment pipeline');
+ }
+
+ version-check:
+ name: Version Check
+ runs-on: ubuntu-latest
+ needs: [test, lint, check-package-label]
+ if: needs.check-package-label.outputs.has_package_label == 'true'
+ outputs:
+ version: ${{ steps.get_version.outputs.version }}
+ is_new_version: ${{ steps.check_pypi.outputs.is_new }}
+
+ steps:
+ - name: Checkout code
+ uses: actions/checkout@v4
+
+ - name: Set up Python
+ uses: actions/setup-python@v5
+ with:
+ python-version: ${{ env.PYTHON_VERSION }}
+
+ - name: Install dependencies for PyPI API
+ run: pip install requests packaging
+
+ - name: Get current version and compare with PyPI
+ id: version_check
+ run: |
+ python << 'EOF'
+ import re
+ import sys
+ import os
+ import requests
+ from packaging import version
+
+ def get_latest_pypi_version(package_name: str) -> str:
+ """Fetch the latest version of a PyPI package by name."""
+ url = f"https://pypi.org/pypi/{package_name}/json"
+ try:
+ response = requests.get(url, timeout=10)
+ if response.status_code == 200:
+ data = response.json()
+ return data["info"]["version"]
+ else:
+ raise Exception(f"Package '{package_name}' returned status {response.status_code}")
+ except requests.exceptions.RequestException as e:
+ raise Exception(f"Failed to fetch package info: {e}")
+
+ # Get current version from pyproject.toml
+ with open('pyproject.toml', 'r') as f:
+ content = f.read()
+ match = re.search(r'version = "([^"]+)"', content)
+
+ if not match:
+ print("❌ ERROR: Could not find version in pyproject.toml")
+ sys.exit(1)
+
+ current_version = match.group(1)
+ package_name = "confopt"
+
+ print(f"Current version from pyproject.toml: {current_version}")
+
+ # Get latest version from PyPI
+ try:
+ pypi_version = get_latest_pypi_version(package_name)
+ print(f"Latest version on PyPI: {pypi_version}")
+ except Exception as e:
+ print(f"❌ ERROR: Could not fetch PyPI version: {e}")
+ sys.exit(1)
+
+ # Compare versions
+ try:
+ current_ver = version.parse(current_version)
+ pypi_ver = version.parse(pypi_version)
+
+ if current_ver > pypi_ver:
+ print(f"✅ Version bump detected: {pypi_version} → {current_version}")
+ print("Proceeding with deployment")
+ is_new_version = True
+ elif current_ver == pypi_ver:
+ print(f"❌ No version bump: current version {current_version} equals PyPI version {pypi_version}")
+ print("Please bump the version in pyproject.toml before deploying")
+ is_new_version = False
+ sys.exit(1)
+ else:
+ print(f"❌ Version downgrade detected: {pypi_version} → {current_version}")
+ print("Current version is lower than PyPI version - this should not happen")
+ is_new_version = False
+ sys.exit(1)
+
+ except Exception as e:
+ print(f"❌ ERROR: Could not parse versions: {e}")
+ print(f"Current: {current_version}, PyPI: {pypi_version}")
+ sys.exit(1)
+
+ # Set outputs
+ with open(os.environ['GITHUB_OUTPUT'], 'a') as f:
+ f.write(f"version={current_version}\n")
+ f.write(f"pypi_version={pypi_version}\n")
+ f.write(f"is_new_version={'true' if is_new_version else 'false'}\n")
+ EOF
+
+ build:
+ name: Build Python Package
+ runs-on: ubuntu-latest
+ needs: [test, lint, version-check]
+
+ steps:
+ - name: Checkout code
+ uses: actions/checkout@v4
+
+ - name: Set up Python
+ uses: actions/setup-python@v5
+ with:
+ python-version: ${{ env.PYTHON_VERSION }}
+
+ - name: Install build dependencies
+ run: |
+ python -m pip install --upgrade pip
+ pip install build twine
+
+ - name: Build package (wheel and sdist)
+ run: python -m build
+
+ - name: Verify built packages
+ run: twine check dist/*
+
+ - name: Upload built packages
+ uses: actions/upload-artifact@v4
+ with:
+ name: python-package-distributions
+ path: dist/
+ retention-days: 2
+ verify_package:
+ name: Verify Package Installation
+ runs-on: ubuntu-latest
+ needs: [build]
+
+ steps:
+ - name: Checkout code
+ uses: actions/checkout@v4
+
+ - name: Download build artifacts
+ uses: actions/download-artifact@v4
+ with:
+ name: python-package-distributions
+ path: dist/
+
+ - name: Set up Python
+ uses: actions/setup-python@v5
+ with:
+ python-version: ${{ env.PYTHON_VERSION }}
+
+ - name: List built packages
+ run: |
+ echo "Built packages:"
+ ls -la dist/
+ echo ""
+ echo "Package summary:"
+ echo "Source distributions: $(ls dist/*.tar.gz 2>/dev/null | wc -l)"
+ echo "Wheels: $(ls dist/*.whl 2>/dev/null | wc -l)"
+
+ - name: Test wheel installation
+ run: |
+ # Test wheel installation
+ python -m venv test_wheel_env
+ source test_wheel_env/bin/activate
+ pip install --upgrade pip
+ pip install dist/*.whl
+
+ # Run minimal confopt test
+ python -c "
+ import numpy as np
+ from confopt.tuning import ConformalTuner
+ from confopt.wrapping import IntRange, FloatRange
+
+ # Minimal synthetic test
+ def simple_objective(configuration):
+ # Simple quadratic function with noise
+ x, y = configuration['x'], configuration['y']
+ return (x - 2)**2 + (y - 3)**2 + np.random.normal(0, 0.1)
+
+ search_space = {
+ 'x': FloatRange(min_value=0.0, max_value=5.0),
+ 'y': FloatRange(min_value=0.0, max_value=5.0)
+ }
+
+ tuner = ConformalTuner(
+ objective_function=simple_objective,
+ search_space=search_space,
+ minimize=True
+ )
+
+ tuner.tune(max_searches=50, n_random_searches=15, verbose=False)
+ best_params = tuner.get_best_params()
+ best_value = tuner.get_best_value()
+
+ print(f'Wheel installation and basic functionality test successful!')
+ print(f'Best params: {best_params}')
+ print(f'Best value: {best_value:.4f}')
+ "
+
+ deactivate
+ rm -rf test_wheel_env
+
+ - name: Test source distribution installation
+ run: |
+ # Test source installation
+ python -m venv test_sdist_env
+ source test_sdist_env/bin/activate
+ pip install --upgrade pip
+ pip install dist/*.tar.gz
+
+ # Run minimal confopt test
+ python -c "
+ import numpy as np
+ from confopt.tuning import ConformalTuner
+ from confopt.wrapping import IntRange, FloatRange
+
+ # Minimal synthetic test
+ def simple_objective(configuration):
+ # Simple quadratic function with noise
+ x, y = configuration['x'], configuration['y']
+ return (x - 2)**2 + (y - 3)**2 + np.random.normal(0, 0.1)
+
+ search_space = {
+ 'x': FloatRange(min_value=0.0, max_value=5.0),
+ 'y': FloatRange(min_value=0.0, max_value=5.0)
+ }
+
+ tuner = ConformalTuner(
+ objective_function=simple_objective,
+ search_space=search_space,
+ minimize=True
+ )
+
+ tuner.tune(max_searches=50, n_random_searches=15, verbose=False)
+ best_params = tuner.get_best_params()
+ best_value = tuner.get_best_value()
+
+ print(f'Source distribution installation and basic functionality test successful!')
+ print(f'Best params: {best_params}')
+ print(f'Best value: {best_value:.4f}')
+ "
+
+ deactivate
+ rm -rf test_sdist_env
+ test-publish:
+ name: Publish to TestPyPI
+ runs-on: ubuntu-latest
+ needs: [verify_package]
+
+ steps:
+ - name: Download build artifacts
+ uses: actions/download-artifact@v4
+ with:
+ name: python-package-distributions
+ path: dist/
+
+ - name: Publish to TestPyPI
+ uses: pypa/gh-action-pypi-publish@release/v1
+ with:
+ repository-url: https://test.pypi.org/legacy/
+ password: ${{ secrets.TEST_PYPI_API_TOKEN }}
+ skip-existing: true # Skip if version already exists
+
+ - name: Wait for TestPyPI propagation
+ run: sleep 30
+ verify-testpypi:
+ name: Verify TestPyPI Installation
+ runs-on: ubuntu-latest
+ needs: [test-publish]
+
+ steps:
+ - name: Checkout code
+ uses: actions/checkout@v4
+
+ - name: Set up Python
+ uses: actions/setup-python@v5
+ with:
+ python-version: ${{ env.PYTHON_VERSION }}
+
+ - name: Get version
+ id: get_version
+ run: |
+ # Get version from pyproject.toml
+ python << 'EOF'
+ import re
+ import sys
+ import os
+
+ with open('pyproject.toml', 'r') as f:
+ content = f.read()
+ match = re.search(r'version = "([^"]+)"', content)
+
+ if not match:
+ print("ERROR: Could not find version in pyproject.toml")
+ sys.exit(1)
+
+ version = match.group(1)
+ with open(os.environ['GITHUB_OUTPUT'], 'a') as f:
+ f.write(f"version={version}\n")
+ print(f"Current version: {version}")
+ EOF
+
+ - name: Test installation scenarios from TestPyPI
+ run: |
+ VERSION=${{ steps.get_version.outputs.version }}
+
+ # Test wheel installation from TestPyPI
+ echo "Test: Wheel installation from TestPyPI..."
+ python -m venv test_wheel_env
+ source test_wheel_env/bin/activate
+ pip install --upgrade pip
+
+ pip install --index-url https://test.pypi.org/simple/ --extra-index-url https://pypi.org/simple/ confopt==$VERSION
+
+ python -c "
+ import numpy as np
+ from confopt.tuning import ConformalTuner
+ from confopt.wrapping import IntRange, FloatRange
+
+ # Minimal synthetic test
+ def simple_objective(configuration):
+ # Simple quadratic function with noise
+ x, y = configuration['x'], configuration['y']
+ return (x - 2)**2 + (y - 3)**2 + np.random.normal(0, 0.1)
+
+ search_space = {
+ 'x': FloatRange(min_value=0.0, max_value=5.0),
+ 'y': FloatRange(min_value=0.0, max_value=5.0)
+ }
+
+ tuner = ConformalTuner(
+ objective_function=simple_objective,
+ search_space=search_space,
+ minimize=True
+ )
+
+ tuner.tune(max_searches=50, n_random_searches=15, verbose=False)
+ best_params = tuner.get_best_params()
+ best_value = tuner.get_best_value()
+
+ print('TestPyPI wheel installation and functionality test successful!')
+ print(f'Best params: {best_params}')
+ print(f'Best value: {best_value:.4f}')
+ "
+
+ deactivate
+ rm -rf test_wheel_env
+
+
+
+ # Test source installation from TestPyPI
+ echo "Test: Source installation from TestPyPI..."
+ python -m venv test_source_env
+ source test_source_env/bin/activate
+ pip install --upgrade pip
+
+ pip install --no-binary=confopt --index-url https://test.pypi.org/simple/ --extra-index-url https://pypi.org/simple/ confopt==$VERSION
+
+ python -c "
+ import numpy as np
+ from confopt.tuning import ConformalTuner
+ from confopt.wrapping import IntRange, FloatRange
+
+ # Minimal synthetic test
+ def simple_objective(configuration):
+ # Simple quadratic function with noise
+ x, y = configuration['x'], configuration['y']
+ return (x - 2)**2 + (y - 3)**2 + np.random.normal(0, 0.1)
+
+ search_space = {
+ 'x': FloatRange(min_value=0.0, max_value=5.0),
+ 'y': FloatRange(min_value=0.0, max_value=5.0)
+ }
+
+ tuner = ConformalTuner(
+ objective_function=simple_objective,
+ search_space=search_space,
+ minimize=True
+ )
+
+ tuner.tune(max_searches=50, n_random_searches=15, verbose=False)
+ best_params = tuner.get_best_params()
+ best_value = tuner.get_best_value()
+
+ print('TestPyPI source installation and functionality test successful!')
+ print(f'Best params: {best_params}')
+ print(f'Best value: {best_value:.4f}')
+ "
+
+ deactivate
+ rm -rf test_source_env
+
+ echo "All TestPyPI installation scenarios validated successfully!"
+ publish:
+ name: Publish to PyPI
+ runs-on: ubuntu-latest
+ needs: [verify-testpypi]
+
+ steps:
+ - name: Download build artifacts
+ uses: actions/download-artifact@v4
+ with:
+ name: python-package-distributions
+ path: dist/
+
+ - name: Publish to PyPI
+ uses: pypa/gh-action-pypi-publish@release/v1
+ with:
+ password: ${{ secrets.PYPI_API_TOKEN }}
+
+ release:
+ name: Create GitHub Release
+ runs-on: ubuntu-latest
+ needs: [publish]
+ permissions:
+ contents: write
+
+ steps:
+ - name: Checkout code
+ uses: actions/checkout@v4
+
+ - name: Download build artifacts
+ uses: actions/download-artifact@v4
+ with:
+ name: python-package-distributions
+ path: dist/
+
+ - name: Create GitHub Release Draft
+ uses: softprops/action-gh-release@v2
+ with:
+ tag_name: v${{ needs.version-check.outputs.version }}
+ name: Release v${{ needs.version-check.outputs.version }}
+ body: |
+ ## Package Information
+ - **Version**: ${{ needs.version-check.outputs.version }}
+ - **PyPI**: https://pypi.org/project/confopt/${{ needs.version-check.outputs.version }}/
+
+ ## Changes
+ *Please add release notes and changelog information here before publishing.*
+
+ ---
+
+ **Build Information:**
+ - Commit: ${{ github.sha }}
+ - PR: #${{ needs.check-package-label.outputs.pr_number }}
+ files: dist/*
+ draft: true
+ prerelease: false
diff --git a/.gitignore b/.gitignore
index f2c98bd..e0bb25d 100644
--- a/.gitignore
+++ b/.gitignore
@@ -24,3 +24,14 @@ var/
*.egg-info/
.installed.cfg
*.egg
+
+# Compiled extension modules
+*.pyd
+*.so
+*.c
+*.whl
+
+# Dev
+cache/
+_build/
+benchmarks/
diff --git a/.pre-commit-config.yaml b/.pre-commit-config.yaml
index 0e904be..4bc081a 100644
--- a/.pre-commit-config.yaml
+++ b/.pre-commit-config.yaml
@@ -6,6 +6,7 @@ repos:
- id: end-of-file-fixer
- id: check-yaml
- id: check-added-large-files
+ exclude: \.c$
- id: debug-statements
- id: detect-private-key
- repo: https://github.com/psf/black
@@ -16,4 +17,9 @@ repos:
rev: 7.0.0
hooks:
- id: flake8
- args: ['--max-line-length=131', '--ignore=E203,W503']
+ args: ['--max-line-length=131', '--ignore=E203,W503,E501']
+- repo: https://github.com/PyCQA/autoflake
+ rev: v2.2.0 # Use the latest stable version
+ hooks:
+ - id: autoflake
+ args: ["--remove-all-unused-imports", "--remove-unused-variables", "--in-place"]
diff --git a/.readthedocs.yml b/.readthedocs.yml
new file mode 100644
index 0000000..03690f0
--- /dev/null
+++ b/.readthedocs.yml
@@ -0,0 +1,32 @@
+# Read the Docs configuration file
+# See https://docs.readthedocs.io/en/stable/config-file/v2.html for details
+
+version: 2
+
+# Set the version of Python and other tools you might need
+build:
+ os: ubuntu-22.04
+ tools:
+ python: "3.11"
+ jobs:
+ post_install:
+ # Install the package with documentation dependencies
+ - pip install -e ".[docs]"
+
+# Build documentation in the docs/ directory with Sphinx
+sphinx:
+ configuration: docs/conf.py
+ fail_on_warning: true
+
+# Build formats
+formats:
+ - pdf
+ - epub
+
+# Declare the Python requirements required to build your documentation
+python:
+ install:
+ - method: pip
+ path: .
+ extra_requirements:
+ - docs
diff --git a/MANIFEST.in b/MANIFEST.in
new file mode 100644
index 0000000..b876f29
--- /dev/null
+++ b/MANIFEST.in
@@ -0,0 +1,35 @@
+# Include essential files
+include LICENSE
+include README.md
+include requirements.txt
+
+# Exclude build artifacts and temporary files
+prune build
+prune dist
+prune *.egg-info
+prune __pycache__
+global-exclude *.pyc
+global-exclude *.pyo
+
+global-exclude .DS_Store
+
+# Exclude development and testing directories
+prune tests
+prune examples
+prune misc
+prune benchmarks
+prune cache
+prune docs
+prune assets
+
+# Exclude result and profile directories
+prune benchmark_results
+prune code_optimization_results
+prune comparison_results
+prune method_profiles
+prune optimization_results
+
+# Exclude development files
+exclude requirements-dev.txt
+exclude .pre-commit-config.yaml
+exclude .readthedocs.yml
diff --git a/README.md b/README.md
index d145f0f..a9528b1 100644
--- a/README.md
+++ b/README.md
@@ -1,92 +1,165 @@
-# ConfOpt
+
-[](https://opensource.org/licenses/Apache-2.0)
-[](https://doi.org/10.48550/arXiv.2207.03017)
+

+
-The package currently implements Adaptive Conformal Hyperparameter Optimization (ACHO), as detailed
-in [the original paper](https://doi.org/10.48550/arXiv.2207.03017).
+
-## Installation
+[](https://pepy.tech/project/confopt)
+[](https://pepy.tech/project/confopt)
+[](https://badge.fury.io/py/confopt)
+[](https://confopt.readthedocs.io/)
+[](https://pypi.org/project/confopt/)
+
-You can install ConfOpt from [PyPI](https://pypi.org/project/confopt) using `pip`:
+
+
+---
+
+Built for machine learning practitioners requiring flexible and robust hyperparameter tuning, **ConfOpt** delivers superior optimization performance through conformal uncertainty quantification and a wide selection of surrogate models.
+
+## 📦 Installation
+
+Install ConfOpt from PyPI using pip:
```bash
pip install confopt
```
-## Getting Started
+For the latest development version:
+
+```bash
+git clone https://github.com/rick12000/confopt.git
+cd confopt
+pip install -e .
+```
+
+## 🎯 Getting Started
-As an example, we'll tune a Random Forest model with data from a regression task.
+The example below shows how to optimize hyperparameters for a RandomForest classifier. You can find more examples in the [documentation](https://confopt.readthedocs.io/).
-Start by setting up your training and validation data:
+### Step 1: Import Required Libraries
```python
-from sklearn.datasets import fetch_california_housing
-
-X, y = fetch_california_housing(return_X_y=True)
-split_idx = int(len(X) * 0.5)
-X_train, y_train = X[:split_idx, :], y[:split_idx]
-X_val, y_val = X[split_idx:, :], y[split_idx:]
+from confopt.tuning import ConformalTuner
+from confopt.wrapping import IntRange, FloatRange, CategoricalRange
+from sklearn.ensemble import RandomForestClassifier
+from sklearn.datasets import load_wine
+from sklearn.model_selection import train_test_split
+from sklearn.metrics import accuracy_score
```
+We import the necessary libraries for tuning and model evaluation. The `load_wine` function is used to load the wine dataset, which serves as our example data for optimizing the hyperparameters of the RandomForest classifier (the dataset is trivial and we can easily reach 100% accuracy, this is for example purposes only).
-Then import the Random Forest model to tune and define a search space for
-its parameters (must be a dictionary mapping the model's parameter names to
-possible values of that parameter to search):
+### Step 2: Define the Objective Function
```python
-from sklearn.ensemble import RandomForestRegressor
-
-parameter_search_space = {
- "n_estimators": [10, 30, 50, 100, 150, 200, 300, 400],
- "min_samples_split": [0.005, 0.01, 0.1, 0.2, 0.3],
- "min_samples_leaf": [0.005, 0.01, 0.1, 0.2, 0.3],
- "max_features": [None, 0.8, 0.9, 1],
-}
+def objective_function(configuration):
+ X, y = load_wine(return_X_y=True)
+ X_train, X_test, y_train, y_test = train_test_split(
+ X, y, test_size=0.3, random_state=42, stratify=y
+ )
+
+ model = RandomForestClassifier(
+ n_estimators=configuration['n_estimators'],
+ max_features=configuration['max_features'],
+ criterion=configuration['criterion'],
+ random_state=42
+ )
+ model.fit(X_train, y_train)
+ predictions = model.predict(X_test)
+
+ return accuracy_score(y_test, predictions)
```
+This function defines the objective we want to optimize. It loads the wine dataset, splits it into training and testing sets, and trains a RandomForest model using the provided configuration. The function returns test accuracy, which will be the objective value ConfOpt will optimize for.
-Now import the `ConformalSearcher` class and initialize it with:
+### Step 3: Define the Search Space
-- The model to tune.
-- The raw X and y data.
-- The parameter search space.
-- An extra variable clarifying whether this is a regression or classification problem.
+```python
+search_space = {
+ 'n_estimators': IntRange(min_value=50, max_value=200),
+ 'max_features': FloatRange(min_value=0.1, max_value=1.0),
+ 'criterion': CategoricalRange(choices=['gini', 'entropy', 'log_loss'])
+}
+```
+Here, we specify the search space for hyperparameters. In this Random Forest example, this includes defining the range for the number of estimators, the proportion of features to consider when looking for the best split, and the criterion for measuring the quality of a split.
-Hyperparameter tuning can be kicked off with the `search` method and a specification
-of how long the tuning should run for (in seconds):
+### Step 4: Create and Run the Tuner
```python
-from confopt.tuning import ConformalSearcher
-
-searcher = ConformalSearcher(
- model=RandomForestRegressor(),
- X_train=X_train,
- y_train=y_train,
- X_val=X_val,
- y_val=y_val,
- search_space=parameter_search_space,
- prediction_type="regression",
-)
-
-searcher.search(
- runtime_budget=120 # How many seconds to run the search for
+tuner = ConformalTuner(
+ objective_function=objective_function,
+ search_space=search_space,
+ minimize=False
)
+tuner.tune(max_searches=50, n_random_searches=10)
```
+We initialize the `ConformalTuner` with the objective function and search space. The `tune` method then kickstarts hyperparameter search and finds the hyperparameters that maximize test accuracy.
-Once done, you can retrieve the best parameters obtained during tuning using:
+### Step 5: Retrieve and Display Results
```python
-best_params = searcher.get_best_params()
+best_params = tuner.get_best_params()
+best_score = tuner.get_best_value()
+
+print(f"Best accuracy: {best_score:.4f}")
+print(f"Best parameters: {best_params}")
```
+Finally, we retrieve the optimization's best parameters and test accuracy score and print them to the console for review.
-Or automatically retrain your model on full data and optimal parameters with:
+For detailed examples and explanations see the [documentation](https://confopt.readthedocs.io/).
-```python
-best_model = searcher.fit_best_model()
-```
+## 📚 Documentation
+
+### **User Guide**
+- **[Classification Example](https://confopt.readthedocs.io/en/latest/basic_usage/classification_example.html)**: RandomForest hyperparameter tuning on a classification task.
+- **[Regression Example](https://confopt.readthedocs.io/en/latest/basic_usage/regression_example.html)**: RandomForest hyperparameter tuning on a regression task.
+
+### **Developer Resources**
+- **[Architecture Overview](https://confopt.readthedocs.io/en/latest/architecture.html)**: System design and module interactions.
+- **[API Reference](https://confopt.readthedocs.io/en/latest/api_reference.html)**:
+Complete reference for main classes, methods, and parameters.
+
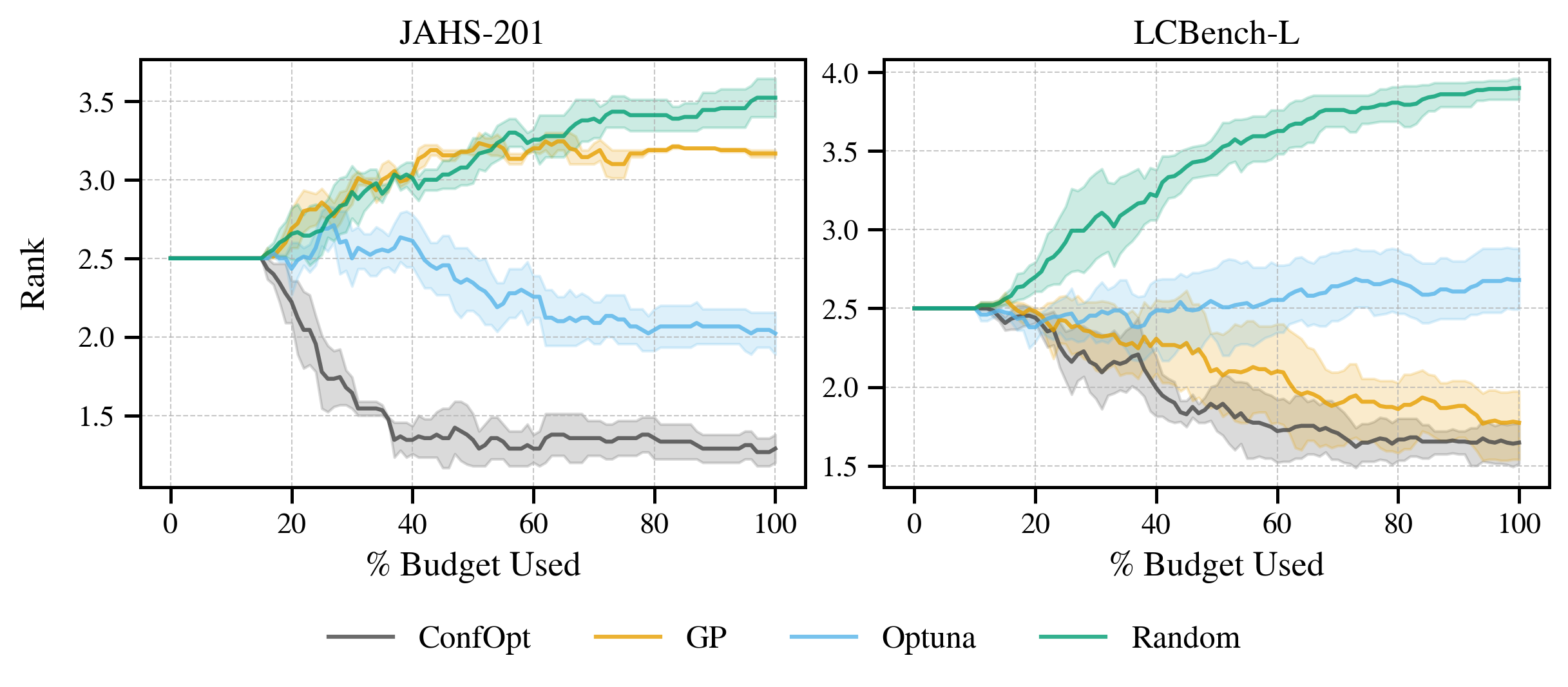
+## 📈 Benchmarks
+
+
+
+
+
Ready to take your hyperparameter optimization to the next level?
+
Get Started |
+
API Docs
+
tune()
random_search()
conformal_search()
_evaluate_configuration()"]
+ STOP["stop_search()
check_objective_function()"]
+ end
+
+ subgraph "Experiment Management"
+ STUDY["Study
append_trial()
batch_append_trials()
get_best_configuration()
get_best_performance()
get_searched_configurations()
get_searched_performances()
get_average_target_model_runtime()"]
+ TRIAL["Trial
iteration
timestamp
configuration
performance
acquisition_source
lower_bound
upper_bound
searcher_runtime
target_model_runtime"]
+ RT["RuntimeTracker
pause_runtime()
resume_runtime()
return_runtime()"]
+ PBM["ProgressBarManager
create_progress_bar()
update_progress()
close_progress_bar()"]
+ end
+
+ subgraph "Configuration Management"
+ BCM["BaseConfigurationManager
mark_as_searched()
tabularize_configs()
listify_configs()
add_to_banned_configurations()"]
+ SCM["StaticConfigurationManager
get_searchable_configurations()
get_searchable_configurations_count()"]
+ DCM["DynamicConfigurationManager
get_searchable_configurations()
get_searchable_configurations_count()"]
+ CE["ConfigurationEncoder
transform()
_build_encoding_schema()
_create_feature_matrix()"]
+ GTC["get_tuning_configurations()
_uniform_sampling()
_sobol_sampling()"]
+ CCH["create_config_hash()
hash_generation()"]
+ end
+
+ subgraph "Acquisition Layer"
+ BCS["BaseConformalSearcher
predict()
update()
get_interval()
_calculate_betas()"]
+ QCS["QuantileConformalSearcher
fit()
_predict_with_ucb()
_predict_with_thompson()
_predict_with_pessimistic_lower_bound()
_predict_with_expected_improvement()"]
+ end
+
+ subgraph "Conformal Prediction"
+ QCE["QuantileConformalEstimator
fit()
predict_intervals()
calculate_betas()
update_alphas()
_fit_non_conformal()
_fit_cv_plus()
_fit_train_test_split()"]
+ DTACI["DtACI
update()
pinball_loss()"]
+ SACS["set_calibration_split()
alpha_to_quantiles()"]
+ end
+
+ subgraph "Hyperparameter Tuning"
+ RT_TUNER["RandomTuner
tune()
_create_fold_indices()
_score_configurations()
_fit_model()
_evaluate_model()"]
+ PT["PointTuner
tune()
_fit_model()
_evaluate_model()"]
+ QT["QuantileTuner
_fit_model()
_evaluate_model()"]
+ IE["initialize_estimator()
estimator_creation()"]
+ ASCF["average_scores_across_folds()
score_aggregation()"]
+ end
+
+ subgraph "Estimator Registry"
+ ER["ESTIMATOR_REGISTRY
rf, gbm, kr, knn
qgbm, qrf, qknn, ql, qgp, qleaf
qens1, qens2, qens3, qens4, qens5"]
+ EC["EstimatorConfig
estimator_name
estimator_class
default_params
estimator_parameter_space
ensemble_components
is_ensemble_estimator()
is_quantile_estimator()"]
+ end
+
+ subgraph "Quantile Estimators"
+ BMFQE["BaseMultiFitQuantileEstimator
fit()
_fit_quantile_estimator()"]
+ BSFQE["BaseSingleFitQuantileEstimator
fit()
_fit_implementation()"]
+ QL["QuantileLasso
fit()
predict_quantiles()"]
+ QG["QuantileGBM
fit()
predict_quantiles()"]
+ QF["QuantileForest
fit()
predict_quantiles()"]
+ QK["QuantileKNN
fit()
predict_quantiles()"]
+ QGP["QuantileGP
fit()
predict_quantiles()"]
+ QLeaf["QuantileLeaf
fit()
predict_quantiles()"]
+ end
+
+ subgraph "Ensemble Methods"
+ BEE["BaseEnsembleEstimator
fit()
predict()"]
+ PEE["PointEnsembleEstimator
fit()
predict()
_compute_point_weights()
_compute_linear_stack_weights()
_get_stacking_training_data()"]
+ QEE["QuantileEnsembleEstimator
fit()
predict()
_compute_quantile_weights()
_compute_linear_stack_weights()
_get_stacking_training_data()"]
+ QLM["QuantileLassoMeta
fit()
predict()
_quantile_loss_objective()"]
+ end
+
+ subgraph "Sampling Strategies"
+ LBS["LowerBoundSampler
calculate_ucb_predictions()
update_exploration_step()
fetch_alphas()
update_interval_width()"]
+ PLBS["PessimisticLowerBoundSampler
fetch_alphas()
update_interval_width()"]
+ TS["ThompsonSampler
calculate_thompson_predictions()
fetch_alphas()
update_interval_width()"]
+ EIS["ExpectedImprovementSampler
calculate_expected_improvement()
update_best_value()
fetch_alphas()
update_interval_width()"]
+ end
+
+ subgraph "Sampling Utilities"
+ IQA["initialize_quantile_alphas()
alpha_generation()"]
+ IMA["initialize_multi_adapters()
adapter_creation()"]
+ ISA["initialize_single_adapter()
single_adapter_setup()"]
+ UMIW["update_multi_interval_widths()
width_updates()"]
+ USIW["update_single_interval_width()
single_width_update()"]
+ FCB["flatten_conformal_bounds()
bounds_flattening()"]
+ VEQ["validate_even_quantiles()
quantile_validation()"]
+ end
+
+ subgraph "Data Processing"
+ TVS["train_val_split()
data_splitting()"]
+ end
+
+ subgraph "Searcher Optimization"
+ DSO["DecayingSearcherOptimizer
select_arm()
update()
_calculate_current_interval()"]
+ FSO["FixedSearcherOptimizer
select_arm()
update()"]
+ end
+
+ subgraph "Parameter Structures"
+ PR["ParameterRange
IntRange
FloatRange
CategoricalRange"]
+ CB["ConformalBounds
lower_bounds
upper_bounds"]
+ end
+
+ %% Main Flow Connections
+ CT --> STUDY
+ CT --> RT
+ CT --> PBM
+ CT --> SCM
+ CT --> DCM
+ CT --> QCS
+ CT --> TVS
+ CT --> DSO
+ CT --> FSO
+ CT --> STOP
+
+ %% Configuration Management Flow
+ STUDY --> TRIAL
+ STUDY --> CE
+ STUDY --> GTC
+ STUDY --> CCH
+ BCM --> SCM
+ BCM --> DCM
+ SCM --> GTC
+ DCM --> GTC
+ DCM --> DSO
+
+ %% Acquisition Flow
+ QCS --> QCE
+ BCS --> LBS
+ BCS --> PLBS
+ BCS --> TS
+ BCS --> EIS
+
+ %% Conformal Prediction Flow
+ QCE --> QT
+ QCE --> IE
+ QCE --> DTACI
+ QCE --> SACS
+ QCS --> SACS
+ DTACI --> SACS
+
+ %% Hyperparameter Tuning Flow
+ RT_TUNER --> IE
+ RT_TUNER --> ASCF
+ PT --> RT_TUNER
+ PT --> ER
+ QT --> RT_TUNER
+ QT --> ER
+ IE --> ER
+ IE --> EC
+
+ %% Estimator Flow
+ ER --> EC
+ EC --> BMFQE
+ EC --> BSFQE
+ BMFQE --> QL
+ BMFQE --> QG
+ BSFQE --> QF
+ BSFQE --> QK
+ BSFQE --> QGP
+ BSFQE --> QLeaf
+ EC --> BEE
+ BEE --> PEE
+ BEE --> QEE
+
+ %% Ensemble Flow
+ PEE --> BMFQE
+ PEE --> BSFQE
+ QEE --> BMFQE
+ QEE --> BSFQE
+ QEE --> QLM
+
+ %% Sampling Utilities Flow
+ LBS --> IQA
+ LBS --> VEQ
+ PLBS --> IQA
+ PLBS --> VEQ
+ TS --> IQA
+ TS --> IMA
+ TS --> ISA
+ TS --> VEQ
+ EIS --> IQA
+ EIS --> UMIW
+ EIS --> USIW
+ EIS --> VEQ
+
+ %% Adaptive Flow
+ IMA --> DTACI
+ ISA --> DTACI
+ UMIW --> DTACI
+ USIW --> DTACI
+
+ %% Data Structure Flow
+ CT --> PR
+ QCE --> CB
+ LBS --> CB
+ PLBS --> CB
+ TS --> CB
+ EIS --> CB
+
+ %% Styling
+ style CT fill:#ff6b6b
+ style QCS fill:#4ecdc4
+ style QCE fill:#45b7d1
+ style DSO fill:#96ceb4
+ style STUDY fill:#feca57
+
+End-to-End Execution Flow
+~~~~~~~~~~~~~~~~~~~~~~~~~
+
+**Step 1: Initialization and Setup**
+
+When ``ConformalTuner.tune()`` starts, it creates a ``Study`` object to track all trials and results. The study initializes a ``RuntimeTracker`` for timing and ``ProgressBarManager`` for user feedback. Parameter spaces are defined using ``ParameterRange`` objects (``IntRange``, ``FloatRange``, ``CategoricalRange``) which specify search bounds and types.
+
+Configuration management happens through either ``StaticConfigurationManager`` (for predefined configurations) or ``DynamicConfigurationManager`` (for adaptive suggestions). The ``ConfigurationEncoder`` handles conversion between different parameter representations, while ``get_tuning_configurations()`` generates initial parameter samples using uniform or Sobol sequences.
+
+**Step 2: Acquisition Function Setup**
+
+The system uses quantile-based conformal prediction for acquisition:
+
+* ``QuantileConformalSearcher`` - uses direct quantile estimation
+
+This inherits from ``BaseConformalSearcher`` which provides the common interface for ``predict()``, ``update()``, and ``get_interval()`` methods.
+
+**Conformal Estimator Initialization:**
+
+``QuantileConformalEstimator`` implements quantile-based conformal prediction using direct quantile estimation with conformal adjustment for coverage guarantees.
+
+**Step 3: Data Processing Pipeline**
+
+Raw input data flows through ``train_val_split()`` which creates training, validation, and calibration sets. This split data structure maintains proper separation required for conformal prediction coverage guarantees.
+
+For ``QuantileConformalEstimator``, the training data gets processed as:
+
+* Quantile estimation → trains quantile regression models for prediction intervals
+* Validation set → generates nonconformity scores for conformal calibration
+
+**Step 4: Hyperparameter Tuning Layer**
+
+The tuning hierarchy works as follows:
+
+.. code-block:: text
+
+ RandomTuner (base class)
+ ├── PointTuner (for point estimation)
+ └── QuantileTuner (for quantile estimation)
+
+``tune()`` handles the optimization process:
+
+1. Creates cross-validation folds through ``_create_fold_indices()``
+2. Scores configurations using ``_score_configurations()``
+3. Uses ``initialize_estimator()`` to create estimator instances from ``ESTIMATOR_REGISTRY``
+4. Performs cross-validation through ``_fit_model()`` and ``_evaluate_model()``
+5. Aggregates results using ``average_scores_across_folds()``
+6. Returns fitted estimator and best hyperparameters
+
+The ``ESTIMATOR_REGISTRY`` contains ``EstimatorConfig`` objects that define:
+
+* Architecture identifiers
+* Parameter ranges for hyperparameter search
+* Default parameter values
+* Estimator class references
+
+**Step 5: Estimator Implementation Layer**
+
+The system supports multiple quantile estimator types:
+
+**Individual Quantile Estimators:**
+
+* ``QuantileLasso`` - L1-regularized quantile regression
+* ``QuantileGBM`` - Gradient boosting for quantile estimation
+* ``QuantileForest`` - Random forest with quantile prediction
+* ``QuantileKNN`` - K-nearest neighbors for quantile estimation
+* ``QuantileGP`` - Gaussian process with quantile likelihood
+* ``QuantileLeaf`` - Leaf-based quantile estimation
+
+**Ensemble Estimators:**
+
+* ``BaseEnsembleEstimator`` - abstract base class for ensemble methods
+* ``PointEnsembleEstimator`` - combines multiple point estimators using weighted averaging with uniform or linear stacking strategies
+* ``QuantileEnsembleEstimator`` - combines multiple quantile estimators using uniform or linear stacking approaches
+* ``QuantileLassoMeta`` - specialized meta-learner for quantile ensemble optimization using Lasso regression
+
+Ensemble implementations support multiple weighting strategies:
+- Uniform weighting for simple averaging
+- Linear stacking with cross-validation optimization
+- Lasso-based meta-learning for optimal weight computation
+
+**Step 6: Acquisition Strategy Execution**
+
+The ``BaseConformalSearcher.predict()`` method routes to strategy-specific implementations:
+
+**Acquisition Function Hierarchy:**
+
+.. code-block:: text
+
+ Acquisition Strategies
+ ├── LowerBoundSampler (Upper Confidence Bound)
+ ├── PessimisticLowerBoundSampler (Conservative Lower Bound)
+ ├── ThompsonSampler (Posterior Sampling)
+ └── ExpectedImprovementSampler (Expected Improvement)
+
+Each strategy calls specific methods:
+
+* ``LowerBoundSampler`` → ``calculate_ucb_predictions()``
+* ``ThompsonSampler`` → ``calculate_thompson_predictions()``
+* ``ExpectedImprovementSampler`` → ``_calculate_expected_improvement()``
+
+
+All strategies use shared utilities from ``selection.sampling.utils``:
+
+* ``initialize_quantile_alphas()`` - sets up alpha levels
+* ``initialize_multi_adapters()`` / ``initialize_single_adapter()`` - configures adaptive mechanisms
+* ``update_multi_interval_widths()`` / ``update_single_interval_width()`` - adjusts interval sizes
+* ``flatten_conformal_bounds()`` - converts bounds to usable format
+
+**Step 7: Conformal Prediction and Interval Generation**
+
+The conformal estimators generate prediction intervals:
+
+1. ``fit()`` method trains on calibration data
+2. ``predict_intervals()`` generates ``ConformalBounds`` objects containing lower_bound, upper_bound, and alpha values
+3. ``calculate_betas()`` computes coverage feedback for adaptive adjustment
+
+**Step 8: Adaptive Feedback Loop**
+
+After each evaluation, the system updates:
+
+1. ``get_interval()`` retrieves prediction interval bounds for storage and analysis
+2. ``_calculate_betas()`` computes coverage statistics
+3. ``DtACI.update()`` adjusts significance levels based on coverage feedback
+4. ``pinball_loss()`` provides loss-based adaptation signals
+
+**Step 9: Trial Management and Optimization**
+
+Results flow back through the trial management system:
+
+1. ``_evaluate_configuration()`` executes the objective function
+2. ``append_trial()`` records results in the study
+3. ``get_best_configuration()`` retrieves current optimal configuration
+4. ``conformal_search()`` continues the optimization loop
+
+**Conformal Searcher Optimization**
+
+All conformal searchers require training on the accumulated configuration-to-performance pairs during search. The system provides different optimization strategies for determining when and how frequently to retrain the searchers:
+
+* ``DecayingSearcherOptimizer`` - increases tuning intervals over time using linear, exponential, or logarithmic decay functions
+* ``FixedSearcherOptimizer`` - maintains constant retraining intervals and tuning trial counts
+
+The system also supports disabling searcher optimization entirely for simpler use cases.
diff --git a/docs/basic_usage/classification_example.rst b/docs/basic_usage/classification_example.rst
new file mode 100644
index 0000000..cda1d98
--- /dev/null
+++ b/docs/basic_usage/classification_example.rst
@@ -0,0 +1,202 @@
+Classification Example
+=======================
+
+This example will show you how to use ConfOpt to optimize hyperparameters for a classification task.
+
+If you already used hyperparameter tuning packages, the "Code Example" section below will give you a quick run through of how to use ConfOpt. If not, don't worry, the "Detailed Walkthrough" section will explain everything step-by-step.
+
+Code Example
+------------
+
+1. Set up search space and objective function:
+
+.. code-block:: python
+
+
+ from confopt.tuning import ConformalTuner
+ from confopt.wrapping import IntRange, FloatRange, CategoricalRange
+
+ from sklearn.ensemble import RandomForestClassifier
+
+ from sklearn.datasets import load_wine
+ from sklearn.model_selection import train_test_split
+ from sklearn.metrics import accuracy_score
+
+ search_space = {
+ 'n_estimators': IntRange(min_value=50, max_value=200),
+ 'max_features': FloatRange(min_value=0.1, max_value=1.0),
+ 'criterion': CategoricalRange(choices=['gini', 'entropy', 'log_loss'])
+ }
+
+ def objective_function(configuration):
+ X, y = load_wine(return_X_y=True)
+ X_train, X_test, y_train, y_test = train_test_split(
+ X, y, test_size=0.3, random_state=42, stratify=y
+ )
+
+ model = RandomForestClassifier(
+ n_estimators=configuration['n_estimators'],
+ max_features=configuration['max_features'],
+ criterion=configuration['criterion'],
+ random_state=42
+ )
+
+ model.fit(X_train, y_train)
+ predictions = model.predict(X_test)
+ score = accuracy_score(y_test, predictions)
+
+ return score
+
+2. Call ConfOpt to tune hyperparameters:
+
+.. code-block:: python
+
+ tuner = ConformalTuner(
+ objective_function=objective_function,
+ search_space=search_space,
+ minimize=False
+ )
+
+ tuner.tune(
+ max_searches=50,
+ n_random_searches=10,
+ verbose=True
+ )
+
+3. Extract results:
+
+.. code-block:: python
+
+ best_params = tuner.get_best_params()
+ best_accuracy = tuner.get_best_value()
+
+ tuned_model = RandomForestClassifier(**best_params, random_state=42)
+
+
+Detailed Walkthrough
+--------------------
+
+Imports
+~~~~~~~
+
+First, let's import everything we'll be needing:
+
+.. code-block:: python
+
+ from confopt.tuning import ConformalTuner
+ from confopt.wrapping import IntRange, FloatRange, CategoricalRange
+
+ from sklearn.ensemble import RandomForestClassifier
+
+ from sklearn.datasets import load_wine
+ from sklearn.model_selection import train_test_split
+ from sklearn.metrics import accuracy_score
+
+For this tutorial, we'll be using the sklearn Wine dataset and trying to tune the hyperparameters of a ``RandomForestClassifier``.
+
+Search Space
+~~~~~~~~~~~~
+
+Next, we need to define the hyperparameter space we want ``confopt`` to optimize over.
+
+This is done using the :ref:`IntRange `, :ref:`FloatRange `, and :ref:`CategoricalRange ` classes, which specify the ranges for each hyperparameter.
+Below let's define a simple example with one of each type of hyperparameter:
+
+.. code-block:: python
+
+ search_space = {
+ 'n_estimators': IntRange(min_value=50, max_value=200),
+ 'max_features': FloatRange(min_value=0.1, max_value=1.0),
+ 'criterion': CategoricalRange(choices=['gini', 'entropy', 'log_loss'])
+ }
+
+
+This tells ``confopt`` to explore the following hyperparameter ranges:
+
+* ``n_estimators``: Number of trees in the forest (all integer values from 50 to 200)
+* ``max_features``: Fraction of features to consider at each split (any float between 0.1 and 1.0)
+* ``criterion``: Function to measure the quality of a split (choose from 'gini', 'entropy', or 'log_loss')
+
+
+Objective Function
+~~~~~~~~~~~~~~~~~~
+
+The objective function defines how the model trains and what metric you want to optimize for during hyperparameter search:
+
+.. code-block:: python
+
+ def objective_function(configuration):
+ X, y = load_wine(return_X_y=True)
+ X_train, X_test, y_train, y_test = train_test_split(
+ X, y, test_size=0.3, random_state=42, stratify=y
+ )
+
+ model = RandomForestClassifier(
+ n_estimators=configuration['n_estimators'],
+ max_features=configuration['max_features'],
+ criterion=configuration['criterion'],
+ random_state=42
+ )
+
+ model.fit(X_train, y_train)
+ predictions = model.predict(X_test)
+ score = accuracy_score(y_test, predictions)
+
+ return score
+
+The objective function must take a single argument called ``configuration``, which is a dictionary containing a hyperparameter value for each hyperparameter name specified in your ``search_space``. The values will be chosen automatically by the tuner during optimization.
+
+The ``score`` can be any metric of your choosing (e.g., accuracy, log loss, F1 score, etc.). This is the value that ``confopt`` will try to optimize for.
+
+In this example, the data is loaded and split inside the objective function for simplicity, but you may prefer to load the data outside (to avoid reloading it for each configuration) and
+either pass the training and test sets as arguments using ``partial`` from the ``functools`` library, or reference them from the global scope.
+
+Running the Optimization
+~~~~~~~~~~~~~~~~~~~~~~~~
+
+
+To start optimizing, first instantiate a :ref:`ConformalTuner ` by providing your objective function, search space, and the optimization direction:
+
+.. code-block:: python
+
+ tuner = ConformalTuner(
+ objective_function=objective_function,
+ search_space=search_space,
+ minimize=False # Use True for metrics like log loss
+ )
+
+The ``minimize`` parameter should be set to ``False`` if you want to maximize your metric (e.g., accuracy), or ``True`` if you want to minimize it (e.g., log loss).
+
+To actually kickstart the hyperparameter search, call:
+
+.. code-block:: python
+
+ tuner.tune(
+ max_searches=50,
+ n_random_searches=10,
+ verbose=True
+ )
+
+Where:
+
+* ``max_searches`` controls how many different hyperparameter configurations will be tried in total.
+* ``n_random_searches`` sets how many of those will be chosen randomly before the tuner switches to using smart optimization (eg. ``max_searches=50`` and ``n_random_searches=10`` means the tuner will sample 10 random configurations, then 40 smart configurations).
+
+
+Getting the Results
+~~~~~~~~~~~~~~~~~~~
+
+
+After that runs, you can retrieve the best hyperparameters or the best score found respectively using ``get_best_params()`` and ``get_best_value()``:
+
+.. code-block:: python
+
+ best_params = tuner.get_best_params()
+ best_accuracy = tuner.get_best_value()
+
+Which you can use to instantiate a tuned version of your model:
+
+.. code-block:: python
+
+
+ tuned_model = RandomForestClassifier(**best_params, random_state=42)
diff --git a/docs/basic_usage/regression_example.rst b/docs/basic_usage/regression_example.rst
new file mode 100644
index 0000000..71ab574
--- /dev/null
+++ b/docs/basic_usage/regression_example.rst
@@ -0,0 +1,194 @@
+Regression Example
+==================
+
+This example will show you how to use ConfOpt to optimize hyperparameters for a regression task.
+
+If you already used hyperparameter tuning packages, the "Code Example" section below will give you a quick run through of how to use ConfOpt. If not, don't worry, the "Detailed Walkthrough" section will explain everything step-by-step.
+
+Code Example
+------------
+
+1. Set up search space and objective function:
+
+.. code-block:: python
+
+
+ from confopt.tuning import ConformalTuner
+ from confopt.wrapping import IntRange, FloatRange, CategoricalRange
+ from sklearn.ensemble import RandomForestRegressor
+ from sklearn.datasets import load_diabetes
+ from sklearn.model_selection import train_test_split
+ from sklearn.metrics import mean_squared_error, r2_score
+
+ search_space = {
+ 'n_estimators': IntRange(min_value=50, max_value=200),
+ 'max_depth': IntRange(min_value=3, max_value=15),
+ 'min_samples_split': IntRange(min_value=2, max_value=10)
+ }
+
+ def objective_function(configuration):
+ X, y = load_diabetes(return_X_y=True)
+ X_train, X_test, y_train, y_test = train_test_split(
+ X, y, test_size=0.3, random_state=42
+ )
+
+ model = RandomForestRegressor(
+ n_estimators=configuration['n_estimators'],
+ max_depth=configuration['max_depth'],
+ min_samples_split=configuration['min_samples_split'],
+ random_state=42
+ )
+
+ model.fit(X_train, y_train)
+ predictions = model.predict(X_test)
+ mse = mean_squared_error(y_test, predictions)
+ return mse # Lower is better (minimize MSE)
+
+2. Call ConfOpt to tune hyperparameters:
+
+.. code-block:: python
+
+ tuner = ConformalTuner(
+ objective_function=objective_function,
+ search_space=search_space,
+ minimize=True # Minimizing MSE
+ )
+
+ tuner.tune(
+ max_searches=50,
+ n_random_searches=10,
+ verbose=True
+ )
+
+3. Extract results:
+
+.. code-block:: python
+
+ best_params = tuner.get_best_params()
+ best_mse = tuner.get_best_value()
+
+ tuned_model = RandomForestRegressor(**best_params, random_state=42)
+
+Detailed Walkthrough
+--------------------
+
+Imports
+~~~~~~~
+
+First, let's import everything we'll be needing:
+
+.. code-block:: python
+
+ from confopt.tuning import ConformalTuner
+ from confopt.wrapping import IntRange, FloatRange, CategoricalRange
+
+ from sklearn.ensemble import RandomForestRegressor
+
+ from sklearn.datasets import load_diabetes
+ from sklearn.model_selection import train_test_split
+ from sklearn.metrics import mean_squared_error
+
+For this tutorial, we'll be using the sklearn Diabetes dataset and trying to tune the hyperparameters of a ``RandomForestRegressor``.
+
+Search Space
+~~~~~~~~~~~~
+
+Next, we need to define the hyperparameter space we want ``confopt`` to optimize over.
+
+This is done using the :ref:`IntRange `, :ref:`FloatRange `, and :ref:`CategoricalRange ` classes, which specify the ranges for each hyperparameter.
+
+Below let's define a simple example with a few typical hyperparameters for regression:
+
+.. code-block:: python
+
+ search_space = {
+ 'n_estimators': IntRange(min_value=50, max_value=200),
+ 'max_depth': IntRange(min_value=3, max_value=15),
+ 'min_samples_split': IntRange(min_value=2, max_value=10)
+ }
+
+This tells ``confopt`` to explore the following hyperparameter ranges:
+
+* ``n_estimators``: Number of trees in the forest (all integer values from 50 to 200)
+* ``max_depth``: Maximum tree depth (all integer values from 3 to 15)
+* ``min_samples_split``: Minimum samples to split a node (all integer values from 2 to 10)
+
+Objective Function
+~~~~~~~~~~~~~~~~~~
+
+The objective function defines how the model trains and what metric you want to optimize for during hyperparameter search:
+
+.. code-block:: python
+
+ def objective_function(configuration):
+ X, y = load_diabetes(return_X_y=True)
+ X_train, X_test, y_train, y_test = train_test_split(
+ X, y, test_size=0.3, random_state=42
+ )
+
+ model = RandomForestRegressor(
+ n_estimators=configuration['n_estimators'],
+ max_depth=configuration['max_depth'],
+ min_samples_split=configuration['min_samples_split'],
+ random_state=42
+ )
+
+ model.fit(X_train, y_train)
+ predictions = model.predict(X_test)
+ mse = mean_squared_error(y_test, predictions)
+ return mse # Lower is better (minimize MSE)
+
+
+The objective function must take a single argument called ``configuration``, which is a dictionary containing a value for each hyperparameter name specified in your ``search_space``. The values will be chosen automatically by the tuner during optimization.
+
+The ``score`` can be any metric of your choosing (e.g., MSE, R², MAE, etc.). This is the value that ``confopt`` will try to optimize for. For MSE, lower is better, so we minimize it.
+
+In this example, the data is loaded and split inside the objective function for simplicity, but you may prefer to load the data outside (to avoid reloading it for each configuration) and either pass the training and test sets as arguments using ``partial`` from the ``functools`` library, or reference them from the global scope.
+
+Running the Optimization
+~~~~~~~~~~~~~~~~~~~~~~~~
+
+
+To start optimizing, first instantiate a :ref:`ConformalTuner ` by providing your objective function, search space, and the optimization direction:
+
+.. code-block:: python
+
+ tuner = ConformalTuner(
+ objective_function=objective_function,
+ search_space=search_space,
+ minimize=True # Minimizing MSE
+ )
+
+The ``minimize`` parameter should be set to ``True`` to minimize metrics where lower is better (e.g., MSE, MAE), or ``False`` to maximize metrics where higher is better (e.g., R²).
+
+To actually kickstart the hyperparameter search, call:
+
+.. code-block:: python
+
+ tuner.tune(
+ max_searches=50,
+ n_random_searches=10,
+ verbose=True
+ )
+
+Where:
+
+* ``max_searches`` controls how many different hyperparameter configurations will be tried in total.
+* ``n_random_searches`` sets how many of those will be chosen randomly before the tuner switches to using smart optimization (e.g., ``max_searches=50`` and ``n_random_searches=10`` means the tuner will sample 10 random configurations, then 40 smart configurations).
+
+Getting the Results
+~~~~~~~~~~~~~~~~~~~
+
+
+After that runs, you can retrieve the best hyperparameters or the best score found respectively using :meth:`~confopt.tuning.ConformalTuner.get_best_params` and :meth:`~confopt.tuning.ConformalTuner.get_best_value`:
+
+.. code-block:: python
+
+ best_params = tuner.get_best_params()
+ best_mse = tuner.get_best_value()
+
+Which you can use to instantiate a tuned version of your model:
+
+.. code-block:: python
+
+ tuned_model = RandomForestRegressor(**best_params, random_state=42)
diff --git a/docs/conf.py b/docs/conf.py
new file mode 100644
index 0000000..084bc03
--- /dev/null
+++ b/docs/conf.py
@@ -0,0 +1,214 @@
+# Configuration file for the Sphinx documentation builder.
+# For the full list of built-in configuration values, see the documentation:
+# https://www.sphinx-doc.org/en/master/usage/configuration.html
+
+import os
+import sys
+
+sys.path.insert(0, os.path.abspath(".."))
+
+# RTD environment detection (optional, for any future customizations)
+on_rtd = os.environ.get("READTHEDOCS", None) == "True"
+rtd_version = os.environ.get("READTHEDOCS_VERSION", "latest")
+
+# -- Project information -----------------------------------------------------
+
+project = "ConfOpt"
+copyright = "2025, Riccardo Doyle"
+author = "Riccardo Doyle"
+release = "2.0.0"
+version = "2.0.0"
+
+# -- General configuration ---------------------------------------------------
+
+extensions = [
+ "sphinx.ext.autodoc",
+ "sphinx.ext.autosummary",
+ "sphinx.ext.napoleon",
+ "sphinx.ext.viewcode",
+ "sphinx.ext.intersphinx",
+ "sphinx.ext.githubpages",
+ "myst_parser",
+ "sphinx_copybutton",
+ "sphinxcontrib.mermaid",
+]
+
+# MyST parser configuration
+myst_enable_extensions = [
+ "colon_fence",
+ "deflist",
+ "html_admonition",
+ "html_image",
+ "linkify",
+ "replacements",
+ "smartquotes",
+ "tasklist",
+]
+
+# Napoleon settings for Google-style docstrings
+napoleon_google_docstring = True
+napoleon_numpy_docstring = False
+napoleon_include_init_with_doc = False
+napoleon_include_private_with_doc = False
+napoleon_include_special_with_doc = True
+napoleon_use_admonition_for_examples = False
+napoleon_use_admonition_for_notes = False
+napoleon_use_admonition_for_references = False
+napoleon_use_ivar = False
+napoleon_use_param = True
+napoleon_use_rtype = True
+
+# Autodoc settings
+
+autodoc_default_options = {
+ "members": True,
+ "member-order": "bysource",
+ "special-members": "__init__",
+ "undoc-members": True,
+ "exclude-members": "__weakref__",
+}
+autodoc_typehints = "description"
+autodoc_class_attributes = False
+
+# Autosummary settings
+autosummary_generate = True
+autosummary_generate_overwrite = True
+
+# Intersphinx mapping
+intersphinx_mapping = {
+ "python": ("https://docs.python.org/3", None),
+ "numpy": ("https://numpy.org/doc/stable/", None),
+ "scipy": ("https://docs.scipy.org/doc/scipy/", None),
+ "sklearn": ("https://scikit-learn.org/stable/", None),
+}
+
+templates_path = ["_templates"]
+exclude_patterns = ["_build", "Thumbs.db", ".DS_Store"]
+
+# -- Options for HTML output -------------------------------------------------
+
+html_theme = "sphinx_rtd_theme"
+html_theme_options = {
+ "canonical_url": "https://confopt.readthedocs.io/",
+ "logo_only": True, # Show project title alongside logo
+ "prev_next_buttons_location": "bottom",
+ "style_external_links": True,
+ "style_nav_header_background": "#db2777", # Match our pink theme
+ # Navigation options - optimized for usability
+ "collapse_navigation": False,
+ "sticky_navigation": True,
+ "navigation_depth": 3,
+ "includehidden": True,
+ "titles_only": False,
+ # Additional RTD theme options
+ "vcs_pageview_mode": "blob",
+ "navigation_with_keys": True,
+}
+
+html_static_path = ["_static"]
+html_css_files = ["custom.css"]
+html_js_files = ["layout-manager.js"]
+
+# GitHub integration
+html_context = {
+ "display_github": True,
+ "github_user": "rick12000",
+ "github_repo": "confopt",
+ "github_version": "main",
+ "conf_py_path": "/docs/",
+}
+
+# Custom logo and favicon
+html_logo = "../assets/logo.png"
+html_favicon = None # RTD will handle this
+
+# The root toctree document (updated from deprecated master_doc)
+root_doc = "index"
+
+# The name of the Pygments (syntax highlighting) style to use
+pygments_style = "sphinx"
+
+# If true, `todo` and `todoList` produce output, else they produce nothing
+todo_include_todos = False
+
+# Security and performance improvements
+tls_verify = True
+tls_cacerts = ""
+
+# Suppress warnings for external references that may not always be available
+suppress_warnings = [
+ "ref.doc",
+ "ref.ref",
+ "epub.unknown_project_files",
+]
+
+# Enable nitpicky mode for better link validation (but suppress known issues)
+nitpicky = True
+nitpick_ignore = [
+ ("py:class", "type"),
+ ("py:class", "object"),
+ ("py:class", "callable"),
+ ("py:class", "default=100"),
+ ("py:class", "default=None"),
+ ("py:class", "default=15"),
+ ("py:class", "default=1"),
+ ("py:class", "default=True"),
+ ("py:class", "confopt.wrapping.IntRange"),
+ ("py:class", "confopt.wrapping.FloatRange"),
+ ("py:class", "confopt.wrapping.CategoricalRange"),
+ ("py:class", "BaseConformalSearcher"),
+ ("py:class", "numpy.array"),
+ ("py:class", "sklearn.preprocessing._data.StandardScaler"),
+ ("py:class", "confopt.selection.acquisition.BaseConformalSearcher"),
+ ("py:class", "confopt.utils.tracking.ProgressBarManager"),
+ ("py:class", "confopt.selection.acquisition.QuantileConformalSearcher"),
+ ("py:class", "confopt.selection.sampling.bound_samplers.LowerBoundSampler"),
+ ("py:class", "confopt.selection.sampling.thompson_samplers.ThompsonSampler"),
+ (
+ "py:class",
+ "confopt.selection.sampling.bound_samplers.PessimisticLowerBoundSampler",
+ ),
+ (
+ "py:class",
+ "confopt.selection.sampling.expected_improvement_samplers.ExpectedImprovementSampler",
+ ),
+ ("py:class", "confopt.wrapping.ConformalBounds"),
+ ("py:class", "pydantic_core.core_schema.ValidationInfo"),
+ ("py:class", "ConfigDict"),
+ ("py:meth", "confopt.tuning.ConformalTuner.get_best_params"),
+ ("py:meth", "confopt.tuning.ConformalTuner.get_best_value"),
+]
+
+# -- Options for LaTeX output ------------------------------------------------
+
+latex_elements = {
+ "papersize": "letterpaper",
+ "pointsize": "10pt",
+}
+
+latex_documents = [
+ (root_doc, "confopt.tex", "ConfOpt Documentation", author, "manual"),
+]
+
+# -- Options for manual page output ------------------------------------------
+
+man_pages = [(root_doc, "confopt", "ConfOpt Documentation", [author], 1)]
+
+# -- Options for Texinfo output ----------------------------------------------
+
+texinfo_documents = [
+ (
+ root_doc,
+ "confopt",
+ "ConfOpt Documentation",
+ author,
+ "confopt",
+ "Voice Command Assistant for Accessibility.",
+ "Miscellaneous",
+ ),
+]
+
+# -- Options for Epub output -------------------------------------------------
+
+epub_title = project
+epub_exclude_files = ["search.html", ".nojekyll", ".doctrees", "environment.pickle"]
diff --git a/docs/contact.rst b/docs/contact.rst
new file mode 100644
index 0000000..22c0761
--- /dev/null
+++ b/docs/contact.rst
@@ -0,0 +1,8 @@
+Contact
+=======
+
+🌟 **GitHub:** https://github.com/rick12000/confopt
+
+🛠️ **Support:** https://github.com/rick12000/confopt/issues
+
+📧 **Contribution Requests or Feedback:** r.doyle.edu@gmail.com
diff --git a/docs/getting_started.rst b/docs/getting_started.rst
new file mode 100644
index 0000000..906fd7e
--- /dev/null
+++ b/docs/getting_started.rst
@@ -0,0 +1,13 @@
+Getting Started
+===============
+
+This section provides practical examples of using ConfOpt for different types of machine learning tasks.
+
+Each example provides a full code example, followed by a step by step explanation.
+
+.. toctree::
+ :maxdepth: 1
+ :caption: Examples
+
+ basic_usage/classification_example
+ basic_usage/regression_example
diff --git a/docs/index.rst b/docs/index.rst
new file mode 100644
index 0000000..5815ccc
--- /dev/null
+++ b/docs/index.rst
@@ -0,0 +1,57 @@
+.. image:: ../assets/logo.png
+ :align: center
+ :width: 250px
+
+
+`ConfOpt `_ is a flexible hyperparameter optimization library, blending the strenghts of quantile regression with the calibration of conformal prediction.
+
+Find out how to **include it in your ML workflow** below! 👇
+
+.. toctree::
+ :maxdepth: 1
+ :caption: User Guide
+
+ installation
+ getting_started
+ advanced_usage
+
+.. toctree::
+ :maxdepth: 1
+ :caption: Developer Guide
+ :hidden:
+
+ api_reference
+ architecture
+
+.. toctree::
+ :maxdepth: 1
+ :caption: Other
+ :hidden:
+
+ roadmap
+ contact
+
+📈 Benchmarks
+=============
+
+.. image:: ../assets/benchmark_results.png
+ :align: center
+ :width: 450px
+ :alt: Benchmark Results
+
+**ConfOpt** is significantly better than plain old random search, but it also beats established tools like **Optuna** or traditional **Gaussian Processes**!
+
+The above benchmark considers neural architecture search on complex image recognition datasets (JAHS-201) and neural network tuning on tabular classification datasets (LCBench-L).
+
+For a fuller analysis of caveats and benchmarking results, refer to the latest methodological paper.
+
+🔬 Theory
+==========
+
+ConfOpt implements surrogate models and acquisition functions from the following papers:
+
+- **Adaptive Conformal Hyperparameter Optimization**: `arXiv, 2022 `_
+
+- **Optimizing Hyperparameters with Conformal Quantile Regression**: `PMLR, 2023 `_
+
+- **Enhancing Performance and Calibration in Quantile Hyperparameter Optimization**: `arXiv, 2025 `_
diff --git a/docs/installation.rst b/docs/installation.rst
new file mode 100644
index 0000000..abb007e
--- /dev/null
+++ b/docs/installation.rst
@@ -0,0 +1,16 @@
+Installation
+============
+
+Install `ConfOpt `_ using pip:
+
+.. code-block:: bash
+
+ pip install confopt
+
+Alternatively, for the latest development version, clone the repository and install it in editable mode:
+
+.. code-block:: bash
+
+ git clone https://github.com/rick12000/confopt.git
+ cd confopt
+ pip install -e .
diff --git a/docs/make.bat b/docs/make.bat
new file mode 100644
index 0000000..2f00427
--- /dev/null
+++ b/docs/make.bat
@@ -0,0 +1,68 @@
+@ECHO OFF
+
+pushd %~dp0
+
+REM Command file for Sphinx documentation
+
+if "%SPHINXBUILD%" == "" (
+ set SPHINXBUILD=sphinx-build
+)
+set SOURCEDIR=.
+set BUILDDIR=_build
+
+%SPHINXBUILD% >NUL 2>NUL
+if errorlevel 9009 (
+ echo.
+ echo.The 'sphinx-build' command was not found. Make sure you have Sphinx
+ echo.installed, then set the SPHINXBUILD environment variable to point
+ echo.to the full path of the 'sphinx-build' executable. Alternatively you
+ echo.may add the Sphinx directory to PATH.
+ echo.
+ echo.If you don't have Sphinx installed, grab it from
+ echo.https://sphinx-doc.org/
+ exit /b 1
+)
+
+if "%1" == "" goto help
+if "%1" == "livehtml" goto livehtml
+if "%1" == "cleanhtml" goto cleanhtml
+
+%SPHINXBUILD% -M %1 %SOURCEDIR% %BUILDDIR% %SPHINXOPTS% %O%
+goto end
+
+:help
+%SPHINXBUILD% -M help %SOURCEDIR% %BUILDDIR% %SPHINXOPTS% %O%
+echo.
+echo.Additional targets:
+echo. livehtml Start live rebuild server using sphinx-autobuild
+echo. cleanhtml Clean build cache and rebuild HTML documentation
+goto end
+
+:livehtml
+echo Starting live documentation server...
+echo Clearing build cache...
+if exist "%BUILDDIR%" rd /s /q "%BUILDDIR%" 2>nul
+echo Performing initial clean build...
+%SPHINXBUILD% -E -a %SOURCEDIR% %BUILDDIR%\html %SPHINXOPTS% %O%
+echo Open http://localhost:8000 in your browser
+echo Press Ctrl+C to stop the server
+sphinx-autobuild %SOURCEDIR% %BUILDDIR%\html %SPHINXOPTS% %O% --host 0.0.0.0 --port 8000 --ignore "*.tmp" --ignore "*.swp" --ignore "*~" --watch %SOURCEDIR%
+if errorlevel 1 (
+ echo.
+ echo.sphinx-autobuild not found. Install with: pip install sphinx-autobuild
+ echo.Or use: build_docs.bat live
+ exit /b 1
+)
+goto end
+
+:cleanhtml
+echo Clearing build cache...
+if exist "%BUILDDIR%" rd /s /q "%BUILDDIR%" 2>nul
+echo Building HTML documentation with clean cache...
+%SPHINXBUILD% -E -a %SOURCEDIR% %BUILDDIR%\html %SPHINXOPTS% %O%
+echo.
+echo.Build finished. The HTML pages are in %BUILDDIR%\html.
+goto end
+
+:end
+popd
diff --git a/docs/requirements.txt b/docs/requirements.txt
new file mode 100644
index 0000000..44090df
--- /dev/null
+++ b/docs/requirements.txt
@@ -0,0 +1,7 @@
+sphinx>=8.1.0
+sphinx-rtd-theme>=2.0.0
+myst-parser>=3.0.0
+sphinx-copybutton>=0.5.2
+sphinxcontrib-mermaid>=0.9.2
+linkify-it-py>=2.0.0
+sphinx-autobuild>=2024.2.4
diff --git a/docs/roadmap.rst b/docs/roadmap.rst
new file mode 100644
index 0000000..2aa17bd
--- /dev/null
+++ b/docs/roadmap.rst
@@ -0,0 +1,21 @@
+========
+Roadmap
+========
+
+Upcoming Features
+=================
+
+Functionality
+------------------------
+
+* **Multi Fidelity Support**: Enable single fidelity conformal searchers to adapt to multi-fidelity settings, allowing them to be competitive in settings where models can be partially trained and lower fidelities are predictive of full fidelity performance.
+* **Multi Objective Support**: Allow searchers to optimize for more than one objective (eg. accuracy and runtime).
+* **Transfer Learning Support**: Allow searchers to use a pretrained model or an observation matcher as a starting point for tuning.
+* **Local Search**: Expected Improvement sampler currently only performs one off configuration scoring. Local search (where a local neighbourhood around the initial EI optimum is explored as a second pass refinement) can significantly improve performance.
+* **Hierarchical Hyperparameters**: Improved handling for hierarchical hyperparameter spaces (currently supported, via flattening of the hyperparameters, but potentially suboptimal for surrogate learning)
+
+Resource Management
+---------------------
+
+* **Parallel Search Support**: Allow searchers to evaluate multiple configurations in parallel if compute allows.
+* **Smart Resource Usage**: Auto detect best amount of parallelism based on available resources and expected load.
diff --git a/examples/tabular_tuning.py b/examples/tabular_tuning.py
deleted file mode 100644
index e40490e..0000000
--- a/examples/tabular_tuning.py
+++ /dev/null
@@ -1,46 +0,0 @@
-from sklearn.datasets import fetch_california_housing
-from sklearn.ensemble import RandomForestRegressor
-from confopt.tuning import ConformalSearcher
-
-# Set up toy data:
-X, y = fetch_california_housing(return_X_y=True)
-split_idx = int(len(X) * 0.5)
-X_train, y_train = X[:split_idx, :], y[:split_idx]
-X_val, y_val = X[split_idx:, :], y[split_idx:]
-
-# Define parameter search space:
-parameter_search_space = {
- "n_estimators": [10, 30, 50, 100, 150, 200, 300, 400],
- "min_samples_split": [0.005, 0.01, 0.1, 0.2, 0.3],
- "min_samples_leaf": [0.005, 0.01, 0.1, 0.2, 0.3],
- "max_features": [None, 0.8, 0.9, 1],
-}
-
-# Set up conformal searcher instance:
-searcher = ConformalSearcher(
- model=RandomForestRegressor(),
- X_train=X_train,
- y_train=y_train,
- X_val=X_val,
- y_val=y_val,
- search_space=parameter_search_space,
- prediction_type="regression",
-)
-
-# Carry out hyperparameter search:
-searcher.search(
- runtime_budget=120,
-)
-
-# Extract results, in the form of either:
-
-# 1. The best hyperparamter configuration found during search
-best_params = searcher.get_best_params()
-
-# 2. An initialized (but not trained) model object with the
-# best hyperparameter configuration found during search
-model_init = searcher.configure_best_model()
-
-# 3. A trained model with the best hyperparameter configuration
-# found during search
-model = searcher.fit_best_model()
diff --git a/pyproject.toml b/pyproject.toml
new file mode 100644
index 0000000..7222e74
--- /dev/null
+++ b/pyproject.toml
@@ -0,0 +1,54 @@
+[build-system]
+requires = ["setuptools>=61.0", "wheel"]
+build-backend = "setuptools.build_meta"
+
+[project]
+name = "confopt"
+version = "2.0.0"
+description = "Conformal hyperparameter optimization tool"
+readme = "README.md"
+authors = [
+ {name = "Riccardo Doyle", email = "r.doyle.edu@gmail.com"}
+]
+requires-python = ">=3.9"
+classifiers = [
+ "Programming Language :: Python :: 3.9",
+ "Programming Language :: Python :: 3.10",
+ "Programming Language :: Python :: 3.11",
+ "Programming Language :: Python :: 3.12",
+]
+dependencies = [
+ "numpy>=1.20.0",
+ "scikit-learn>=1.0.0",
+ "scipy>=1.7.0",
+ "pandas>=1.3.0",
+ "tqdm>=4.60.0",
+ "pydantic>=2.0.0",
+ "joblib>=1.0.0",
+ "statsmodels>=0.13.0"
+]
+
+[project.urls]
+Source = "https://github.com/rick12000/confopt"
+Documentation = "https://confopt.readthedocs.io"
+Changelog = "https://github.com/rick12000/confopt/releases"
+
+[project.optional-dependencies]
+dev = [
+ "pytest>=7.4.0",
+ "pytest-xdist>=3.0.0",
+ "pre-commit>=3.4.0",
+ "autoflake>=2.0.0",
+]
+docs = [
+ "sphinx>=5.0.0",
+ "sphinx-rtd-theme>=1.3.0",
+ "myst-parser>=2.0.0",
+ "sphinx-copybutton>=0.5.0",
+ "sphinxcontrib-mermaid>=0.8.0",
+ "sphinx-autobuild>=2024.10.3"
+]
+
+[tool.setuptools]
+packages = { find = { where = ["."] , include = ["confopt*"] } }
+include-package-data = true
diff --git a/pytest.ini b/pytest.ini
new file mode 100644
index 0000000..7617853
--- /dev/null
+++ b/pytest.ini
@@ -0,0 +1,3 @@
+[pytest]
+markers =
+ slow: marks tests as slow (deselect with '-m "not slow"')
diff --git a/requirements-dev.txt b/requirements-dev.txt
index 8c19c2a..98353cf 100644
--- a/requirements-dev.txt
+++ b/requirements-dev.txt
@@ -1,2 +1,4 @@
-pytest==7.4.2
-pre-commit==3.4.0
+pytest>=7.4.0
+pytest-xdist>=3.0.0
+pre-commit>=3.4.0
+autoflake>=2.0.0
diff --git a/requirements.txt b/requirements.txt
index b30be02..de42639 100644
--- a/requirements.txt
+++ b/requirements.txt
@@ -1,5 +1,8 @@
-numpy>=1.24.4
-scikit-learn>=1.3.2
-quantile-forest>=1.2.4
-tqdm>=4.66.1
-pandas>=2.0.3
+numpy>=1.20.0
+scikit-learn>=1.0.0
+scipy>=1.7.0
+pandas>=1.3.0
+tqdm>=4.60.0
+pydantic>=2.0.0
+joblib>=1.0.0
+statsmodels>=0.13.0
diff --git a/setup.py b/setup.py
deleted file mode 100644
index fe08e4c..0000000
--- a/setup.py
+++ /dev/null
@@ -1,26 +0,0 @@
-from setuptools import setup, find_packages
-
-with open("README.md", "r") as f:
- long_description = f.read()
-
-setup(
- name="confopt",
- description="Conformal hyperparameter optimization tool",
- long_description=long_description,
- long_description_content_type="text/markdown",
- url="https://github.com/rick12000/confopt",
- author="Riccardo Doyle",
- author_email="r.doyle.edu@gmail.com",
- packages=find_packages(),
- version="1.0.2",
- license="Apache License 2.0",
- install_requires=[line.strip() for line in open("requirements.txt").readlines()],
- # TODO: Replace this with explicits
- classifiers=[
- "Programming Language :: Python :: 3.8",
- "Programming Language :: Python :: 3.9",
- "Programming Language :: Python :: 3.10",
- "Programming Language :: Python :: 3.11",
- "Programming Language :: Python :: 3.12",
- ],
-)
diff --git a/tests/conftest.py b/tests/conftest.py
index 28c11ee..0630043 100644
--- a/tests/conftest.py
+++ b/tests/conftest.py
@@ -1,164 +1,923 @@
import random
-from typing import Dict
import numpy as np
import pytest
-from sklearn.ensemble import GradientBoostingRegressor
-
-from confopt.estimation import (
- QuantileConformalRegression,
- LocallyWeightedConformalRegression,
+from typing import Dict
+from confopt.tuning import (
+ ConformalTuner,
+)
+from confopt.utils.configurations.sampling import get_tuning_configurations
+from confopt.selection.acquisition import QuantileConformalSearcher
+from confopt.selection.sampling.thompson_samplers import ThompsonSampler
+from confopt.wrapping import FloatRange, IntRange, CategoricalRange, ConformalBounds
+from sklearn.base import BaseEstimator
+from confopt.selection.estimator_configuration import (
+ ESTIMATOR_REGISTRY,
)
-from confopt.tuning import ConformalSearcher
-from confopt.utils import get_tuning_configurations
+from confopt.selection.estimators.quantile_estimation import (
+ BaseSingleFitQuantileEstimator,
+ BaseMultiFitQuantileEstimator,
+)
+from confopt.selection.estimators.ensembling import (
+ QuantileEnsembleEstimator,
+ PointEnsembleEstimator,
+)
+from unittest.mock import Mock
+from confopt.selection.adaptation import DtACI
DEFAULT_SEED = 1234
-# Dummy made up search space:
-DUMMY_PARAMETER_GRID: Dict = {
- "int_parameter": [1, 2, 3, 4, 5],
- "float_parameter": [1.1, 2.2, 3.3, 4.4],
-}
-# Dummy search space for a GBM model:
-DUMMY_GBM_PARAMETER_GRID: Dict = {
- "n_estimators": [5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20],
- "learning_rate": [0.1, 0.2, 0.3, 0.4, 0.5],
-}
+def build_estimator_architectures(amended: bool = False):
+ """Build estimator architecture lists from ESTIMATOR_REGISTRY.
+
+ Args:
+ amended: If True, creates modified versions with n_estimators=25 for faster testing.
+ If False, creates standard architecture lists.
+
+ Returns:
+ Tuple containing:
+ - point_estimator_architectures: List of point estimator names
+ - single_fit_quantile_estimator_architectures: List of single-fit quantile estimator names
+ - multi_fit_quantile_estimator_architectures: List of multi-fit quantile estimator names
+ - quantile_estimator_architectures: List of all quantile estimator names
+ - estimator_registry: Registry of estimator configurations (amended if requested)
+ """
+ from copy import deepcopy
+
+ point_estimator_architectures = []
+ single_fit_quantile_estimator_architectures = []
+ multi_fit_quantile_estimator_architectures = []
+ quantile_estimator_architectures = []
+
+ # Create registry (amended if requested)
+ if amended:
+ estimator_registry = {}
+ for estimator_name, estimator_config in ESTIMATOR_REGISTRY.items():
+ amended_config = deepcopy(estimator_config)
+
+ # Check if the estimator has n_estimators parameter
+ if (
+ hasattr(amended_config, "default_params")
+ and "n_estimators" in amended_config.default_params
+ ):
+ amended_config.default_params["n_estimators"] = 15
+
+ # Also check ensemble components if it's an ensemble estimator
+ if (
+ hasattr(amended_config, "ensemble_components")
+ and amended_config.ensemble_components
+ ):
+ for component in amended_config.ensemble_components:
+ if "params" in component and "n_estimators" in component["params"]:
+ component["params"]["n_estimators"] = 15
+
+ if estimator_name in ["gp", "qgp"]:
+ continue
+
+ if "qens" in estimator_name:
+ continue
+
+ estimator_registry[estimator_name] = amended_config
+ else:
+ estimator_registry = ESTIMATOR_REGISTRY
+
+ # Build architecture lists
+ for estimator_name, estimator_config in estimator_registry.items():
+ if issubclass(
+ estimator_config.estimator_class,
+ (
+ BaseMultiFitQuantileEstimator,
+ BaseSingleFitQuantileEstimator,
+ QuantileEnsembleEstimator,
+ ),
+ ):
+ quantile_estimator_architectures.append(estimator_name)
+ if issubclass(
+ estimator_config.estimator_class,
+ (BaseMultiFitQuantileEstimator),
+ ):
+ multi_fit_quantile_estimator_architectures.append(estimator_name)
+ elif issubclass(
+ estimator_config.estimator_class,
+ (BaseSingleFitQuantileEstimator),
+ ):
+ single_fit_quantile_estimator_architectures.append(estimator_name)
+ elif issubclass(
+ estimator_config.estimator_class, (BaseEstimator, PointEnsembleEstimator)
+ ):
+ point_estimator_architectures.append(estimator_name)
+
+ return (
+ point_estimator_architectures,
+ single_fit_quantile_estimator_architectures,
+ multi_fit_quantile_estimator_architectures,
+ quantile_estimator_architectures,
+ estimator_registry,
+ )
+
+
+# Create original architecture lists
+(
+ POINT_ESTIMATOR_ARCHITECTURES,
+ SINGLE_FIT_QUANTILE_ESTIMATOR_ARCHITECTURES,
+ MULTI_FIT_QUANTILE_ESTIMATOR_ARCHITECTURES,
+ QUANTILE_ESTIMATOR_ARCHITECTURES,
+ _,
+) = build_estimator_architectures(amended=False)
+
+# Create amended architecture lists for faster testing
+(
+ AMENDED_POINT_ESTIMATOR_ARCHITECTURES,
+ AMENDED_SINGLE_FIT_QUANTILE_ESTIMATOR_ARCHITECTURES,
+ AMENDED_MULTI_FIT_QUANTILE_ESTIMATOR_ARCHITECTURES,
+ AMENDED_QUANTILE_ESTIMATOR_ARCHITECTURES,
+ AMENDED_ESTIMATOR_REGISTRY,
+) = build_estimator_architectures(amended=True)
+
+
+def simple_quadratic_minimization(x):
+ """Simple quadratic function for minimization testing.
+
+ Global minimum at x = [2, -1] with value 0.
+ This creates a clear, smooth objective surface that conformal prediction
+ can easily learn and exploit, unlike random search.
+ """
+ x = np.asarray(x)
+ # Shifted quadratic with minimum at [2, -1]
+ return (x[0] - 2) ** 2 + (x[1] + 1) ** 2
+
+
+def simple_quadratic_maximization(x):
+ """Simple negative quadratic function for maximization testing.
+
+ Global maximum at x = [1, 0.5] with value 0.
+ This creates a clear, smooth objective surface that conformal prediction
+ can easily learn and exploit, unlike random search.
+ """
+ x = np.asarray(x)
+ # Negative shifted quadratic with maximum at [1, 0.5]
+ return -((x[0] - 1) ** 2 + (x[1] - 0.5) ** 2)
+
+
+def rastrigin(x, A=20):
+ n = len(x)
+ rastrigin_value = A * n + np.sum(x**2 - A * np.cos(2 * np.pi * x))
+ return rastrigin_value
+
+
+def ackley(x, a=20, b=0.2, c=2 * np.pi):
+ """Ackley function - commonly used maximization benchmark.
+
+ Global minimum is at x = [0, 0, ..., 0] with value 0.
+ For maximization, we negate this so global maximum is 0 at origin.
+ """
+ x = np.asarray(x)
+ n = len(x)
+ sum1 = np.sum(x**2)
+ sum2 = np.sum(np.cos(c * x))
+ ackley_value = (
+ -a * np.exp(-b * np.sqrt(sum1 / n)) - np.exp(sum2 / n) + a + np.exp(1)
+ )
+ return -ackley_value # Negate for maximization
+
+
+class ObjectiveSurfaceGenerator:
+ def __init__(self, generator: str):
+ self.generator = generator
+
+ def predict(self, params):
+ x = np.array(list(params.values()), dtype=float)
+
+ if self.generator == "rastrigin":
+ y = rastrigin(x=x)
+
+ return y
@pytest.fixture
-def dummy_stationary_gaussian_dataset():
- np.random.seed(DEFAULT_SEED)
- random.seed(DEFAULT_SEED)
+def mock_constant_objective_function():
+ def objective(configuration: Dict):
+ return 2
- X, y = [], []
- for x_observation in range(1, 11):
- for _ in range(0, 1000):
- X.append(x_observation)
- y.append(np.random.normal(0, 101))
- dataset = np.column_stack([X, y])
- np.random.shuffle(dataset)
- return dataset
+ return objective
@pytest.fixture
-def dummy_fixed_quantile_dataset():
+def toy_dataset():
+ # Create a small toy dataset with deterministic values
+ X = np.array([[1, 2], [3, 4], [5, 6], [7, 8]])
+ y = np.array([2, 4, 6, 8])
+ return X, y
+
+
+@pytest.fixture
+def big_toy_dataset():
+ # Create a larger toy dataset with 80 observations and 2 features
+ X = np.linspace(0, 10, 80).reshape(-1, 1) # Capped at 80
+ X = np.hstack([X, X + np.random.normal(0, 1, 80).reshape(-1, 1)]) # Capped at 80
+ # Make y always negative by using negative coefficients and subtracting a constant
+ y = -5 * X[:, 0] - 3 * X[:, 1] - 10 + np.random.normal(0, 1, 80) # Capped at 80
+ return X, y
+
+
+@pytest.fixture
+def quantiles():
+ return [0.1, 0.5, 0.9]
+
+
+@pytest.fixture
+def dummy_expanding_quantile_gaussian_dataset():
np.random.seed(DEFAULT_SEED)
random.seed(DEFAULT_SEED)
X, y = [], []
- for x_observation in range(1, 11):
- for _ in range(0, 1000):
+ # Reduce to 80 total observations (16 per x_observation)
+ for x_observation in range(1, 6):
+ for _ in range(0, 20): # Adjusted to make total 80
X.append(x_observation)
- y.append(random.choice(range(1, 101)))
- dataset = np.column_stack([X, y])
- np.random.shuffle(dataset)
- return dataset
+ y.append(x_observation * np.random.normal(0, 10))
+
+ X_array = np.array(X).reshape(-1, 1)
+ # Normalize X to have zero mean and unit variance
+ X_normalized = (X_array - np.mean(X_array)) / np.std(X_array)
+ return X_normalized, np.array(y)
@pytest.fixture
-def dummy_init_quantile_regression():
- qcr = QuantileConformalRegression(quantile_estimator_architecture="qgbm")
- return qcr
+def dummy_parameter_grid():
+ return {
+ "param_1": FloatRange(min_value=0.01, max_value=100, log_scale=True),
+ "param_2": IntRange(min_value=1, max_value=100),
+ "param_3": CategoricalRange(choices=["option1", "option2", "option3"]),
+ }
@pytest.fixture
-def dummy_init_locally_weighted_regression():
- lwr = LocallyWeightedConformalRegression(
- point_estimator_architecture="gbm",
- demeaning_estimator_architecture="gbm",
- variance_estimator_architecture="gbm",
- )
- return lwr
+def simple_minimization_parameter_grid():
+ """Parameter grid for simple quadratic minimization function.
+
+ Optimum is at x1=2, x2=-1. This grid covers the optimum with reasonable bounds
+ that allow the conformal prediction algorithm to learn the pattern efficiently.
+ """
+ return {
+ "x1": FloatRange(min_value=-2.0, max_value=6.0),
+ "x2": FloatRange(min_value=-5.0, max_value=3.0),
+ }
@pytest.fixture
-def dummy_configuration_performance_bounds():
- """
- Dummy performance bounds, where each set of
- bounds is meant to represent upper and lower
- expectations of a certain parameter configuration's
- performance.
+def simple_maximization_parameter_grid():
+ """Parameter grid for simple quadratic maximization function.
+
+ Optimum is at x1=1, x2=0.5. This grid covers the optimum with reasonable bounds
+ that allow the conformal prediction algorithm to learn the pattern efficiently.
"""
- performance_lower_bounds = np.arange(0, 100, 0.5)
- performance_upper_bounds = performance_lower_bounds + 10
- return performance_lower_bounds, performance_upper_bounds
+ return {
+ "x1": FloatRange(min_value=-2.0, max_value=4.0),
+ "x2": FloatRange(min_value=-2.5, max_value=3.5),
+ }
@pytest.fixture
-def dummy_parameter_grid():
- return DUMMY_PARAMETER_GRID
+def rastrigin_parameter_grid():
+ """Parameter grid for 6-dimensional Rastrigin function optimization."""
+ return {
+ "x1": FloatRange(min_value=-5.12, max_value=5.12),
+ "x2": FloatRange(min_value=-5.12, max_value=5.12),
+ "x3": FloatRange(min_value=-5.12, max_value=5.12),
+ "x4": FloatRange(min_value=-5.12, max_value=5.12),
+ "x5": FloatRange(min_value=-5.12, max_value=5.12),
+ "x6": FloatRange(min_value=-5.12, max_value=5.12),
+ }
@pytest.fixture
-def dummy_configurations(dummy_parameter_grid):
- """
- Samples unique configurations from broader
- possible values in dummy hyperparameter search space.
- """
- max_configurations = 100
- tuning_configurations = get_tuning_configurations(
- parameter_grid=dummy_parameter_grid,
- n_configurations=max_configurations,
- random_state=DEFAULT_SEED,
+def ackley_parameter_grid():
+ """Parameter grid for 6-dimensional Ackley function optimization."""
+ return {
+ "x1": FloatRange(min_value=-32.768, max_value=32.768),
+ "x2": FloatRange(min_value=-32.768, max_value=32.768),
+ "x3": FloatRange(min_value=-32.768, max_value=32.768),
+ "x4": FloatRange(min_value=-32.768, max_value=32.768),
+ "x5": FloatRange(min_value=-32.768, max_value=32.768),
+ "x6": FloatRange(min_value=-32.768, max_value=32.768),
+ }
+
+
+@pytest.fixture
+def linear_data_drift():
+ np.random.seed(42)
+ n = 500
+ X = np.linspace(0, 10, n).reshape(-1, 1)
+
+ noise_level = np.linspace(0.5, 3, n)
+ noise = np.random.normal(0, 1, n) * noise_level
+
+ y = np.zeros(n)
+
+ first_segment = int(0.3 * n)
+ y[:first_segment] = 2 * X[:first_segment].flatten() + 5 + noise[:first_segment]
+
+ second_segment = int(0.6 * n)
+ y[first_segment:second_segment] = (
+ 3 * X[first_segment:second_segment].flatten()
+ + 2
+ + noise[first_segment:second_segment]
)
- return tuning_configurations
+
+ y[second_segment:] = 2.5 * X[second_segment:].flatten() + 8 + noise[second_segment:]
+
+ return X, y
+
+
+@pytest.fixture
+def simple_conformal_bounds():
+ lower_bounds1 = np.array([0.1, 0.3, 0.5])
+ upper_bounds1 = np.array([0.4, 0.6, 0.8])
+
+ lower_bounds2 = np.array([0.2, 0.4, 0.6])
+ upper_bounds2 = np.array([0.5, 0.7, 0.9])
+
+ return [
+ ConformalBounds(lower_bounds=lower_bounds1, upper_bounds=upper_bounds1),
+ ConformalBounds(lower_bounds=lower_bounds2, upper_bounds=upper_bounds2),
+ ]
+
+
+@pytest.fixture
+def estimator1():
+ """Mock point estimator that returns deterministic values scaled to input size."""
+ mock = Mock()
+
+ def scaled_predict(X):
+ # Return values that scale based on input length
+ n_samples = len(X)
+ return np.arange(1, n_samples + 1) * 2 # [2, 4, 6, 8, ...] based on input size
+
+ mock.predict = Mock(side_effect=scaled_predict)
+ mock.fit = Mock(return_value=mock)
+ return mock
+
+
+@pytest.fixture
+def estimator2():
+ """Mock point estimator that returns different deterministic values scaled to input size."""
+ mock = Mock()
+
+ def scaled_predict(X):
+ # Return values that scale based on input length
+ n_samples = len(X)
+ return np.arange(2, n_samples + 2) * 2 # [4, 6, 8, 10, ...] based on input size
+
+ mock.predict = Mock(side_effect=scaled_predict)
+ mock.fit = Mock(return_value=mock)
+ return mock
+
+
+@pytest.fixture
+def quantile_estimator1(quantiles):
+ """Mock quantile estimator that returns deterministic quantile predictions for any input size."""
+ mock = Mock()
+
+ def scaled_predict(X):
+ # Return values for any size of X
+ n_samples = len(X)
+ result = np.zeros((n_samples, len(quantiles)))
+ for i, q in enumerate(quantiles):
+ result[:, i] = (i + 1) * 2 # Values 2, 4, 6 for quantiles
+ return result
+
+ mock.fit = Mock(return_value=mock)
+ mock.predict = Mock(side_effect=scaled_predict)
+ return mock
@pytest.fixture
-def dummy_gbm_parameter_grid():
- return DUMMY_GBM_PARAMETER_GRID
+def quantile_estimator2(quantiles):
+ """Mock quantile estimator that returns constant values across quantiles."""
+ mock = Mock()
+
+ def scaled_predict(X):
+ # Return values for any size of X
+ n_samples = len(X)
+ return np.ones((n_samples, len(quantiles))) * 4
+
+ mock.fit = Mock(return_value=mock)
+ mock.predict = Mock(side_effect=scaled_predict)
+ return mock
+
+
+@pytest.fixture
+def competing_estimator():
+ """Mock point estimator with different performance characteristics."""
+ mock = Mock()
+
+ def scaled_predict(X):
+ # Return values that scale based on input length
+ n_samples = len(X)
+ return (
+ np.arange(0.5, n_samples + 0.5) * 2
+ ) # [1, 3, 5, 7, ...] based on input size
+
+ mock.predict = Mock(side_effect=scaled_predict)
+ mock.fit = Mock(return_value=mock)
+ return mock
@pytest.fixture
-def dummy_gbm_configurations(dummy_gbm_parameter_grid):
- max_configurations = 60
- gbm_tuning_configurations = get_tuning_configurations(
- parameter_grid=dummy_gbm_parameter_grid,
- n_configurations=max_configurations,
- random_state=DEFAULT_SEED,
+def tuner(mock_constant_objective_function, dummy_parameter_grid):
+ # Create a standard tuner instance that can be reused across tests
+ return ConformalTuner(
+ objective_function=mock_constant_objective_function,
+ search_space=dummy_parameter_grid,
+ minimize=True,
+ n_candidates=100,
)
- return gbm_tuning_configurations
@pytest.fixture
-def dummy_initialized_conformal_searcher__gbm_mse(
- dummy_stationary_gaussian_dataset, dummy_gbm_parameter_grid
-):
- """
- Creates a conformal searcher instance from dummy raw X, y data
- and a dummy parameter grid.
+def small_parameter_grid():
+ """Small parameter grid for focused configuration testing"""
+ return {
+ "x": FloatRange(min_value=0.0, max_value=1.0),
+ "y": IntRange(min_value=1, max_value=3),
+ "z": CategoricalRange(choices=["A", "B"]),
+ }
- This particular fixture is set to optimize a GBM base model on
- regression data, using an MSE objective. The model architecture
- and type of data are arbitrarily pinned; more fixtures could
- be created to test other model or data types.
- """
- custom_loss_function = "mean_squared_error"
- prediction_type = "regression"
- model = GradientBoostingRegressor()
- X, y = (
- dummy_stationary_gaussian_dataset[:, 0].reshape(-1, 1),
- dummy_stationary_gaussian_dataset[:, 1],
+@pytest.fixture
+def dynamic_tuner(mock_constant_objective_function, small_parameter_grid):
+ """Tuner configured for dynamic sampling with small candidate count"""
+ return ConformalTuner(
+ objective_function=mock_constant_objective_function,
+ search_space=small_parameter_grid,
+ minimize=True,
+ n_candidates=5,
+ dynamic_sampling=True,
)
- train_split = 0.5
- X_train, y_train = (
- X[: round(len(X) * train_split), :],
- y[: round(len(y) * train_split)],
+
+
+@pytest.fixture
+def static_tuner(mock_constant_objective_function, small_parameter_grid):
+ """Tuner configured for static sampling with small candidate count"""
+ return ConformalTuner(
+ objective_function=mock_constant_objective_function,
+ search_space=small_parameter_grid,
+ minimize=True,
+ n_candidates=10,
+ dynamic_sampling=False,
)
- X_val, y_val = X[round(len(X) * train_split) :, :], y[round(len(y) * train_split) :]
-
- searcher = ConformalSearcher(
- model=model,
- X_train=X_train,
- y_train=y_train,
- X_val=X_val,
- y_val=y_val,
- search_space=dummy_gbm_parameter_grid,
- prediction_type=prediction_type,
- custom_loss_function=custom_loss_function,
+
+
+# Fixtures for quantile estimation testing
+
+
+@pytest.fixture
+def uniform_regression_data():
+ """Generate uniform regression data for quantile testing."""
+ np.random.seed(42)
+ n_samples = 300
+ n_features = 3
+
+ X = np.random.uniform(-1, 1, size=(n_samples, n_features))
+ y = np.random.uniform(0, 1, size=n_samples)
+
+ return X, y
+
+
+@pytest.fixture
+def heteroscedastic_regression_data():
+ """Generate heteroscedastic regression data where variance changes with X."""
+ np.random.seed(42)
+ n_samples = 200
+ X = np.linspace(-3, 3, n_samples).reshape(-1, 1)
+
+ # Heteroscedastic noise: variance increases with |X|
+ noise_std = 0.5 + 1.5 * np.abs(X.flatten())
+ noise = np.random.normal(0, 1, n_samples) * noise_std
+
+ # True function: quadratic with heteroscedastic noise
+ y = 2 * X.flatten() ** 2 + 1.5 * X.flatten() + noise
+
+ return X, y
+
+
+@pytest.fixture
+def multimodal_regression_data():
+ """Generate multimodal regression data with multiple peaks and valleys."""
+ np.random.seed(42)
+ n_samples = 300
+ X = np.linspace(-4, 4, n_samples).reshape(-1, 1)
+
+ # Multimodal function: mixture of Gaussians
+ y = (
+ 2 * np.exp(-0.5 * (X.flatten() + 2) ** 2)
+ + 1.5 * np.exp(-0.5 * (X.flatten() - 1) ** 2)
+ + np.exp(-0.5 * (X.flatten() - 3) ** 2)
+ + np.random.normal(0, 0.3, n_samples)
)
- return searcher
+ return X, y
+
+
+@pytest.fixture
+def skewed_regression_data():
+ """Generate regression data with skewed noise distribution."""
+ np.random.seed(42)
+ n_samples = 250
+ X = np.linspace(0, 5, n_samples).reshape(-1, 1)
+
+ # Skewed noise using exponential distribution
+ skewed_noise = np.random.exponential(0.5, n_samples) - 0.5
+
+ # True function with skewed residuals
+ y = np.sin(X.flatten()) + 0.5 * X.flatten() + skewed_noise
+
+ return X, y
+
+
+@pytest.fixture
+def high_dimensional_regression_data():
+ """Generate high-dimensional regression data for testing scalability."""
+ np.random.seed(42)
+ n_samples = 150
+ n_features = 8
+ X = np.random.randn(n_samples, n_features)
+
+ # Linear combination with interaction terms
+ true_coef = np.array([2, -1, 0.5, -0.5, 1, 0, -0.3, 0.8])
+ y = X @ true_coef + 0.5 * X[:, 0] * X[:, 1] + np.random.normal(0, 0.5, n_samples)
+
+ return X, y
+
+
+@pytest.fixture
+def sparse_regression_data():
+ """Generate sparse regression data with few informative features."""
+ np.random.seed(42)
+ n_samples = 100
+ n_features = 10
+ X = np.random.randn(n_samples, n_features)
+
+ # Only first 3 features are informative
+ true_coef = np.zeros(n_features)
+ true_coef[:3] = [3, -2, 1.5]
+ y = X @ true_coef + np.random.normal(0, 0.3, n_samples)
+
+ return X, y
+
+
+@pytest.fixture
+def toy_regression_data():
+ """Generate simple toy regression data for basic testing."""
+
+ def _generate_data(n_samples=100, n_features=2, noise_std=0.1, random_state=42):
+ np.random.seed(random_state)
+ X = np.random.randn(n_samples, n_features)
+ true_coef = np.ones(n_features)
+ y = X @ true_coef + np.random.normal(0, noise_std, n_samples)
+ return X, y
+
+ return _generate_data
+
+
+@pytest.fixture
+def quantile_test_data():
+ """Generate data with known quantile structure for validation."""
+ np.random.seed(42)
+ n_samples = 500
+ X = np.linspace(-3, 3, n_samples).reshape(-1, 1)
+
+ # Create data where we know the true quantiles
+ # Use a location-scale model: Y = μ(X) + σ(X) * ε
+ mu = 2 * X.flatten() # Mean function
+ sigma = 0.5 + 0.3 * np.abs(X.flatten()) # Scale function
+ epsilon = np.random.normal(0, 1, n_samples) # Standard normal noise
+
+ y = mu + sigma * epsilon
+
+ # Store true quantiles for validation
+ true_quantiles = {}
+ for q in [0.1, 0.25, 0.5, 0.75, 0.9]:
+ from scipy.stats import norm
+
+ true_quantiles[q] = mu + sigma * norm.ppf(q)
+
+ return X, y, true_quantiles
+
+
+@pytest.fixture
+def monotonicity_test_quantiles():
+ """Standard quantiles for monotonicity testing."""
+ return [0.1, 0.25, 0.5, 0.75, 0.9]
+
+
+@pytest.fixture
+def alpha_levels_for_conformalization():
+ """Standard alpha levels for conformalization testing."""
+ return [0.1, 0.2, 0.3] # Corresponding to 90%, 80%, 70% coverage
+
+
+@pytest.fixture
+def estimation_test_data():
+ """Generate test data for estimation module tests."""
+ np.random.seed(42)
+ X = np.random.rand(50, 5)
+ y = X.sum(axis=1) + np.random.normal(0, 0.1, 50)
+ from sklearn.model_selection import train_test_split
+
+ return train_test_split(X, y, test_size=0.25, random_state=42)
+
+
+@pytest.fixture
+def point_tuner():
+ """Create a PointTuner instance for testing."""
+ from confopt.selection.estimation import PointTuner
+
+ return PointTuner(random_state=42)
+
+
+@pytest.fixture
+def quantile_tuner_with_quantiles():
+ """Create a QuantileTuner instance with quantiles for testing."""
+ from confopt.selection.estimation import QuantileTuner
+
+ quantiles = [0.1, 0.9]
+ return QuantileTuner(quantiles=quantiles, random_state=42), quantiles
+
+
+@pytest.fixture
+def multi_interval_bounds():
+ """Create multiple ConformalBounds objects for multi-interval testing."""
+ n_obs = 30
+ bounds_list = []
+ for i in range(3):
+ width_factor = (i + 1) * 0.5
+ lower = np.random.uniform(-1, 0, n_obs)
+ upper = lower + np.random.uniform(0.2 * width_factor, 1.0 * width_factor, n_obs)
+ bounds_list.append(ConformalBounds(lower_bounds=lower, upper_bounds=upper))
+ return bounds_list
+
+
+@pytest.fixture
+def nested_intervals():
+ """Create properly nested intervals for testing interval relationships."""
+ n_obs = 20
+ # Create nested intervals: each inner interval contained within outer
+ center = np.random.uniform(-1, 1, n_obs)
+
+ # Widest interval (lowest confidence)
+ wide_lower = center - 2.0
+ wide_upper = center + 2.0
+
+ # Medium interval
+ med_lower = center - 1.0
+ med_upper = center + 1.0
+
+ # Narrowest interval (highest confidence)
+ narrow_lower = center - 0.5
+ narrow_upper = center + 0.5
+
+ return [
+ ConformalBounds(lower_bounds=wide_lower, upper_bounds=wide_upper),
+ ConformalBounds(lower_bounds=med_lower, upper_bounds=med_upper),
+ ConformalBounds(lower_bounds=narrow_lower, upper_bounds=narrow_upper),
+ ]
+
+
+@pytest.fixture
+def coverage_feedback():
+ """Sample coverage feedback for adaptation testing."""
+ return [0.85, 0.78, 0.92]
+
+
+@pytest.fixture
+def small_dataset():
+ """Small dataset for computational testing."""
+ n_obs = 10
+ bounds = []
+ for _ in range(2):
+ lower = np.random.uniform(-0.5, 0, n_obs)
+ upper = lower + np.random.uniform(0.1, 0.5, n_obs)
+ bounds.append(ConformalBounds(lower_bounds=lower, upper_bounds=upper))
+ return bounds
+
+
+@pytest.fixture
+def test_predictions_and_widths():
+ """Combined point predictions and interval widths for LCB testing."""
+ np.random.seed(42)
+ n_points = 15
+ point_estimates = np.random.uniform(-2, 2, n_points)
+ interval_widths = np.random.uniform(0.2, 1.5, n_points)
+ return point_estimates, interval_widths
+
+
+@pytest.fixture
+def conformal_bounds_deterministic():
+ """Deterministic conformal bounds for reproducible testing."""
+ lower_bounds1 = np.array([1.0, 2.0, 3.0, 4.0])
+ upper_bounds1 = np.array([1.5, 2.5, 3.5, 4.5])
+
+ lower_bounds2 = np.array([0.8, 1.8, 2.8, 3.8])
+ upper_bounds2 = np.array([1.3, 2.3, 3.3, 4.3])
+
+ return [
+ ConformalBounds(lower_bounds=lower_bounds1, upper_bounds=upper_bounds1),
+ ConformalBounds(lower_bounds=lower_bounds2, upper_bounds=upper_bounds2),
+ ]
+
+
+@pytest.fixture
+def comprehensive_minimizing_tuning_setup(simple_minimization_parameter_grid):
+ """Fixture for comprehensive integration test setup (objective, warm starts, tuner, searcher).
+
+ Uses a simple quadratic minimization function that's easy for conformal prediction to learn,
+ ensuring the test validates that conformal search outperforms random search.
+ """
+
+ def optimization_objective(configuration: Dict) -> float:
+ # Extract 2-dimensional vector from configuration
+ x = np.array(
+ [
+ configuration["x1"],
+ configuration["x2"],
+ ]
+ )
+
+ # Use simple quadratic function for minimization (minimum at [2, -1])
+ return simple_quadratic_minimization(x)
+
+ warm_start_configs_raw = get_tuning_configurations(
+ parameter_grid=simple_minimization_parameter_grid,
+ n_configurations=5,
+ random_state=123,
+ sampling_method="uniform",
+ )
+ warm_start_configs = []
+ for config in warm_start_configs_raw:
+ performance = optimization_objective(config)
+ warm_start_configs.append((config, performance))
+
+ def make_tuner_and_searcher(dynamic_sampling):
+ tuner = ConformalTuner(
+ objective_function=optimization_objective,
+ search_space=simple_minimization_parameter_grid,
+ minimize=True,
+ n_candidates=1000,
+ warm_starts=warm_start_configs,
+ dynamic_sampling=dynamic_sampling,
+ )
+ searcher = QuantileConformalSearcher(
+ quantile_estimator_architecture="qgbm",
+ sampler=ThompsonSampler(
+ n_quantiles=4,
+ adapter="DtACI",
+ enable_optimistic_sampling=False,
+ ),
+ n_pre_conformal_trials=32,
+ calibration_split_strategy="train_test_split",
+ )
+ return tuner, searcher, warm_start_configs, optimization_objective
+
+ return make_tuner_and_searcher
+
+
+@pytest.fixture
+def comprehensive_maximizing_tuning_setup(simple_maximization_parameter_grid):
+ """Fixture for comprehensive integration test setup for maximization (objective, warm starts, tuner, searcher).
+
+ Uses a simple quadratic maximization function that's easy for conformal prediction to learn,
+ ensuring the test validates that conformal search outperforms random search.
+ """
+
+ def optimization_objective(configuration: Dict) -> float:
+ # Extract 2-dimensional vector from configuration
+ x = np.array(
+ [
+ configuration["x1"],
+ configuration["x2"],
+ ]
+ )
+
+ # Use simple quadratic function for maximization (maximum at [1, 0.5])
+ return simple_quadratic_maximization(x)
+
+ warm_start_configs_raw = get_tuning_configurations(
+ parameter_grid=simple_maximization_parameter_grid,
+ n_configurations=5,
+ random_state=123,
+ sampling_method="uniform",
+ )
+ warm_start_configs = []
+ for config in warm_start_configs_raw:
+ performance = optimization_objective(config)
+ warm_start_configs.append((config, performance))
+
+ def make_tuner_and_searcher(dynamic_sampling):
+ tuner = ConformalTuner(
+ objective_function=optimization_objective,
+ search_space=simple_maximization_parameter_grid,
+ minimize=False, # Set to False for maximization
+ n_candidates=1000,
+ warm_starts=warm_start_configs,
+ dynamic_sampling=dynamic_sampling,
+ )
+ searcher = QuantileConformalSearcher(
+ quantile_estimator_architecture="qgbm",
+ sampler=ThompsonSampler(
+ n_quantiles=4,
+ adapter="DtACI",
+ enable_optimistic_sampling=False,
+ ),
+ n_pre_conformal_trials=32,
+ calibration_split_strategy="train_test_split",
+ )
+ return tuner, searcher, warm_start_configs, optimization_objective
+
+ return make_tuner_and_searcher
+
+
+@pytest.fixture
+def moderate_shift_data():
+ """Create data with moderate distribution shift (0.1 -> 0.5 noise)."""
+ np.random.seed(42)
+ n_points = 200
+ shift_point = 100
+
+ X1 = np.random.randn(shift_point, 2)
+ y1 = X1.sum(axis=1) + 0.1 * np.random.randn(shift_point)
+
+ X2 = np.random.randn(n_points - shift_point, 2)
+ y2 = X2.sum(axis=1) + 0.5 * np.random.randn(n_points - shift_point)
+
+ return np.vstack([X1, X2]), np.hstack([y1, y2])
+
+
+@pytest.fixture
+def high_shift_data():
+ """Create data with high distribution shift (0.1 -> 0.8 -> 0.1 noise)."""
+ np.random.seed(42)
+ n_points = 300
+ shift_points = [100, 200]
+
+ X1 = np.random.randn(shift_points[0], 2)
+ y1 = X1.sum(axis=1) + 0.1 * np.random.randn(shift_points[0])
+
+ X2 = np.random.randn(shift_points[1] - shift_points[0], 2)
+ y2 = X2.sum(axis=1) + 0.8 * np.random.randn(shift_points[1] - shift_points[0])
+
+ X3 = np.random.randn(n_points - shift_points[1], 2)
+ y3 = X3.sum(axis=1) + 0.1 * np.random.randn(n_points - shift_points[1])
+
+ return np.vstack([X1, X2, X3]), np.hstack([y1, y2, y3])
+
+
+@pytest.fixture
+def dtaci_instance():
+ """Standard DtACI instance for testing."""
+ return DtACI(alpha=0.1, gamma_values=[0.01, 0.05, 0.1])
+
+
+# Quantile Estimation Test Data Fixtures
+@pytest.fixture
+def linear_regression_data():
+ """Simple linear regression with homoscedastic noise."""
+ np.random.seed(42)
+ n_samples = 200
+ X = np.linspace(-2, 2, n_samples).reshape(-1, 1)
+ y = 2.5 * X.flatten() + 1.0 + np.random.normal(0, 0.5, n_samples)
+ return X, y
+
+
+@pytest.fixture
+def heteroscedastic_data():
+ """Heteroscedastic data where variance increases with |X|."""
+ np.random.seed(42)
+ n_samples = 300
+ X = np.linspace(-3, 3, n_samples).reshape(-1, 1)
+ noise_std = 0.3 + 1.2 * np.abs(X.flatten())
+ noise = np.random.normal(0, 1, n_samples) * noise_std
+ y = 1.5 * X.flatten() ** 2 + 0.5 * X.flatten() + noise
+ return X, y
+
+
+@pytest.fixture
+def diabetes_data():
+ """Scikit-learn diabetes dataset for regression testing."""
+ from sklearn.datasets import load_diabetes
+
+ diabetes = load_diabetes()
+ return diabetes.data, diabetes.target
+
+
+@pytest.fixture
+def comprehensive_test_quantiles():
+ """Comprehensive set of quantiles for testing."""
+ return [0.05, 0.25, 0.5, 0.75, 0.95]
+
+
+@pytest.fixture
+def ensemble_test_quantiles():
+ return [0.25, 0.5, 0.75]
diff --git a/tests/integration_tests/tuning_integration.py b/tests/integration_tests/tuning_integration.py
new file mode 100644
index 0000000..3f75040
--- /dev/null
+++ b/tests/integration_tests/tuning_integration.py
@@ -0,0 +1,206 @@
+import pytest
+import numpy as np
+from typing import Dict, Tuple, Optional
+
+from confopt.tuning import ConformalTuner
+from confopt.wrapping import CategoricalRange
+from confopt.selection.acquisition import QuantileConformalSearcher, LowerBoundSampler
+
+DRAW_OR_WIN_RATE_THRESHOLD = 0.75
+WINDOW_SIZE = 20
+TARGET_ALPHAS = [0.25, 0.5, 0.75]
+ADAPTER_TYPES = ["DtACI", "ACI"]
+
+
+def complex_objective(configuration: Dict) -> float:
+ x1 = configuration["x1"]
+ x2 = configuration["x2"]
+ categorical_val = {"A": 1.0, "B": 2.5, "C": 4.0}[configuration["categorical"]]
+
+ term1 = np.sin(x1 * np.pi) * np.cos(x2 * np.pi)
+ term2 = 0.5 * (x1 - 0.3) ** 2 + 0.8 * (x2 - 0.7) ** 2
+ term3 = categorical_val * np.exp(-((x1 - 0.5) ** 2 + (x2 - 0.5) ** 2))
+
+ return term1 + term2 + term3 + np.random.normal(0, 0.05)
+
+
+def calculate_coverage_rate_from_study(study) -> float:
+ breach_count = 0
+ total_intervals = 0
+
+ for trial in study.trials:
+ if trial.lower_bound is not None and trial.upper_bound is not None:
+ total_intervals += 1
+ if not (trial.lower_bound <= trial.performance <= trial.upper_bound):
+ breach_count += 1
+
+ return 1 - (breach_count / total_intervals)
+
+
+def calculate_windowed_deviations_from_study(
+ study, alpha: float, window_size: int
+) -> float:
+ target_coverage = 1 - alpha
+ trials = [
+ t
+ for t in study.trials
+ if t.lower_bound is not None and t.upper_bound is not None
+ ]
+
+ if len(trials) < window_size:
+ return 0.0
+
+ n_windows = len(trials) // window_size
+ deviations = []
+
+ for i in range(n_windows):
+ start_idx = i * window_size
+ end_idx = start_idx + window_size
+ window_trials = trials[start_idx:end_idx]
+
+ breaches = sum(
+ 1
+ for t in window_trials
+ if not (t.lower_bound <= t.performance <= t.upper_bound)
+ )
+ window_coverage = 1 - (breaches / window_size)
+ deviation = abs(window_coverage - target_coverage)
+ deviations.append(deviation)
+
+ return np.mean(deviations)
+
+
+def run_experiment(
+ adapter_type: Optional[str], seed: int, alpha: float
+) -> Tuple[float, float]:
+ np.random.seed(seed)
+
+ search_space = {
+ "x1": CategoricalRange(choices=np.linspace(0, 1, 15).tolist()),
+ "x2": CategoricalRange(choices=np.linspace(0, 1, 15).tolist()),
+ "categorical": CategoricalRange(choices=["A", "B", "C"]),
+ }
+
+ interval_width = 1 - alpha
+
+ sampler = LowerBoundSampler(
+ interval_width=interval_width,
+ adapter=adapter_type,
+ c=0,
+ )
+
+ searcher = QuantileConformalSearcher(
+ quantile_estimator_architecture="qgbm",
+ sampler=sampler,
+ n_pre_conformal_trials=32,
+ calibration_split_strategy="train_test_split",
+ )
+
+ tuner = ConformalTuner(
+ objective_function=complex_objective,
+ search_space=search_space,
+ minimize=True,
+ n_candidates=2000,
+ dynamic_sampling=True,
+ )
+
+ tuner.tune(
+ n_random_searches=15,
+ searcher=searcher,
+ random_state=seed,
+ max_searches=60,
+ verbose=False,
+ )
+
+ coverage_rate = calculate_coverage_rate_from_study(tuner.study)
+ windowed_deviation = calculate_windowed_deviations_from_study(
+ tuner.study, alpha, WINDOW_SIZE
+ )
+
+ return coverage_rate, windowed_deviation
+
+
+@pytest.mark.slow
+@pytest.mark.parametrize("target_alpha", TARGET_ALPHAS)
+@pytest.mark.parametrize("adapter_type", ADAPTER_TYPES)
+def test_adaptive_vs_nonadaptive_coverage(target_alpha, adapter_type):
+ print(f"Testing {adapter_type} with target alpha {target_alpha}")
+ n_seeds = 5
+ adaptive_wins_global = 0
+ adaptive_wins_local = 0
+
+ for seed in range(n_seeds):
+ adaptive_coverage, adaptive_local_dev = run_experiment(
+ adapter_type, seed, target_alpha
+ )
+ nonadaptive_coverage, nonadaptive_local_dev = run_experiment(
+ None, seed, target_alpha
+ )
+
+ target_coverage = 1 - target_alpha
+ adaptive_global_dev = abs(adaptive_coverage - target_coverage)
+ nonadaptive_global_dev = abs(nonadaptive_coverage - target_coverage)
+
+ if adaptive_global_dev <= nonadaptive_global_dev:
+ adaptive_wins_global += 1
+ if adaptive_local_dev <= nonadaptive_local_dev:
+ adaptive_wins_local += 1
+
+ global_win_rate = adaptive_wins_global / n_seeds
+ local_win_rate = adaptive_wins_local / n_seeds
+
+ print(f"Global win rate: {global_win_rate}, Local win rate: {local_win_rate}")
+
+ if adapter_type is not None:
+ assert (
+ global_win_rate >= DRAW_OR_WIN_RATE_THRESHOLD
+ ), f"Global win rate: {global_win_rate}"
+ assert (
+ local_win_rate >= DRAW_OR_WIN_RATE_THRESHOLD
+ ), f"Local win rate: {local_win_rate}"
+
+
+def test_dtaci_parameter_evolution():
+ search_space = {
+ "x1": CategoricalRange(choices=np.linspace(0, 1, 8).tolist()),
+ "x2": CategoricalRange(choices=np.linspace(0, 1, 8).tolist()),
+ "categorical": CategoricalRange(choices=["A", "B", "C"]),
+ }
+
+ sampler = LowerBoundSampler(
+ interval_width=0.8,
+ adapter="DtACI",
+ c=0,
+ )
+
+ searcher = QuantileConformalSearcher(
+ quantile_estimator_architecture="ql",
+ sampler=sampler,
+ n_pre_conformal_trials=32,
+ )
+
+ tuner = ConformalTuner(
+ objective_function=complex_objective,
+ search_space=search_space,
+ minimize=True,
+ n_candidates=500,
+ )
+
+ tuner.tune(
+ n_random_searches=15,
+ searcher=searcher,
+ random_state=42,
+ max_searches=100,
+ verbose=False,
+ )
+
+ adapter = sampler.adapter
+
+ assert adapter is not None
+ assert adapter.update_count > 0
+ assert len(adapter.alpha_history) > 0
+
+ for alpha_val in adapter.alpha_history:
+ assert 0.001 <= alpha_val <= 0.999
+
+ assert np.var(adapter.alpha_history) != 0
diff --git a/tests/selection/estimators/test_ensembling.py b/tests/selection/estimators/test_ensembling.py
new file mode 100644
index 0000000..199c4c2
--- /dev/null
+++ b/tests/selection/estimators/test_ensembling.py
@@ -0,0 +1,387 @@
+import pytest
+import numpy as np
+from sklearn.metrics import mean_pinball_loss
+from sklearn.model_selection import train_test_split
+from sklearn.preprocessing import StandardScaler
+from sklearn.linear_model import LinearRegression
+from sklearn.ensemble import RandomForestRegressor
+
+from confopt.selection.estimators.ensembling import (
+ PointEnsembleEstimator,
+ QuantileEnsembleEstimator,
+ QuantileLassoMeta,
+)
+from confopt.selection.estimators.quantile_estimation import (
+ QuantileGBM,
+ QuantileKNN,
+ QuantileLasso,
+)
+
+
+def test_quantile_lasso_meta_fit_predict():
+ """Test that QuantileLassoMeta correctly fits and predicts."""
+ np.random.seed(42)
+ n_samples, n_features = 100, 3
+ X = np.random.randn(n_samples, n_features)
+ y = X @ np.array([0.5, 0.3, 0.2]) + 0.1 * np.random.randn(n_samples)
+
+ quantile_lasso = QuantileLassoMeta(alpha=0.01, quantile=0.5)
+ quantile_lasso.fit(X, y)
+
+ # Check that coefficients sum to 1 (normalized)
+ assert np.isclose(np.sum(quantile_lasso.coef_), 1.0)
+ assert np.all(quantile_lasso.coef_ >= 0) # positive constraint
+
+ # Check prediction works
+ predictions = quantile_lasso.predict(X)
+ assert predictions.shape == (n_samples,)
+
+
+def test_quantile_lasso_meta_different_quantiles():
+ """Test that QuantileLassoMeta gives different weights for different quantiles."""
+ np.random.seed(42)
+ n_samples, n_features = 200, 3
+ X = np.random.randn(n_samples, n_features)
+ y = X @ np.array([0.5, 0.3, 0.2]) + 0.2 * np.random.randn(n_samples)
+
+ quantile_25 = QuantileLassoMeta(alpha=0.01, quantile=0.25)
+ quantile_75 = QuantileLassoMeta(alpha=0.01, quantile=0.75)
+
+ quantile_25.fit(X, y)
+ quantile_75.fit(X, y)
+
+ # Weights might be different for different quantiles
+ assert quantile_25.coef_ is not None
+ assert quantile_75.coef_ is not None
+ assert np.isclose(np.sum(quantile_25.coef_), 1.0)
+ assert np.isclose(np.sum(quantile_75.coef_), 1.0)
+
+
+def test_quantile_lasso_meta_better_than_uniform():
+ """Test that QuantileLassoMeta performs better than uniform weights for quantile loss."""
+ from sklearn.metrics import mean_pinball_loss
+
+ np.random.seed(42)
+ n_samples, n_features = 150, 3
+
+ # Create data where first feature is best for the quantile
+ X = np.random.randn(n_samples, n_features)
+ y = 2 * X[:, 0] + 0.1 * X[:, 1] + 0.05 * X[:, 2] + 0.1 * np.random.randn(n_samples)
+
+ quantile = 0.25
+
+ # Quantile Lasso
+ quantile_lasso = QuantileLassoMeta(alpha=0.01, quantile=quantile)
+ quantile_lasso.fit(X, y)
+ pred_quantile_lasso = quantile_lasso.predict(X)
+
+ # Uniform weights
+ uniform_weights = np.ones(n_features) / n_features
+ pred_uniform = X @ uniform_weights
+
+ # Compare pinball losses
+ loss_quantile_lasso = mean_pinball_loss(y, pred_quantile_lasso, alpha=quantile)
+ loss_uniform = mean_pinball_loss(y, pred_uniform, alpha=quantile)
+
+ # QuantileLasso should perform at least as well as uniform weights
+ assert loss_quantile_lasso <= loss_uniform * 1.05 # Allow small tolerance
+
+
+def create_diverse_quantile_estimators(random_state=42):
+ return [
+ QuantileGBM(
+ learning_rate=0.1,
+ n_estimators=50,
+ min_samples_split=10,
+ min_samples_leaf=5,
+ max_depth=3,
+ random_state=random_state,
+ ),
+ QuantileKNN(n_neighbors=15),
+ QuantileLasso(
+ max_iter=1000,
+ p_tol=1e-6,
+ random_state=random_state,
+ ),
+ ]
+
+
+def create_diverse_point_estimators(random_state=42):
+ return [
+ LinearRegression(),
+ RandomForestRegressor(
+ n_estimators=50,
+ max_depth=3,
+ random_state=random_state,
+ ),
+ ]
+
+
+def evaluate_quantile_performance(y_true, y_pred, quantiles):
+ total_loss = 0.0
+ for i, q in enumerate(quantiles):
+ loss = mean_pinball_loss(y_true, y_pred[:, i], alpha=q)
+ total_loss += loss
+ return total_loss / len(quantiles)
+
+
+def evaluate_point_performance(y_true, y_pred):
+ return np.mean((y_true - y_pred) ** 2)
+
+
+def test_point_ensemble_get_stacking_training_data(toy_dataset, estimator1, estimator2):
+ X, y = toy_dataset
+
+ model = PointEnsembleEstimator(
+ estimators=[estimator1, estimator2], cv=2, random_state=42, alpha=0.01
+ )
+
+ val_indices, val_targets, val_predictions = model._get_stacking_training_data(X, y)
+
+ assert len(np.unique(val_indices)) == len(X)
+ assert val_predictions.shape == (len(X), 2)
+ assert np.array_equal(val_targets, y[val_indices])
+
+
+@pytest.mark.parametrize("weighting_strategy", ["uniform", "linear_stack"])
+def test_point_ensemble_compute_weights(
+ toy_dataset, estimator1, competing_estimator, weighting_strategy
+):
+ X, y = toy_dataset
+
+ model = PointEnsembleEstimator(
+ estimators=[estimator1, competing_estimator],
+ cv=2,
+ weighting_strategy=weighting_strategy,
+ random_state=42,
+ alpha=0.01,
+ )
+
+ weights = model._compute_point_weights(X, y)
+
+ assert len(weights) == 2
+ assert np.isclose(np.sum(weights), 1.0)
+ assert np.all(weights >= 0)
+
+ if weighting_strategy == "uniform":
+ assert np.allclose(weights, np.array([0.5, 0.5]))
+
+
+def test_point_ensemble_predict_with_uniform_weights(
+ toy_dataset, estimator1, estimator2
+):
+ X, _ = toy_dataset
+
+ model = PointEnsembleEstimator(
+ estimators=[estimator1, estimator2],
+ weighting_strategy="uniform",
+ alpha=0.01,
+ )
+ model.weights = np.array([0.5, 0.5])
+
+ predictions = model.predict(X)
+ expected = np.array([3, 5, 7, 9])
+
+ assert predictions[0] == 3
+ assert predictions[-1] == 9
+ assert np.array_equal(predictions, expected)
+
+
+def test_quantile_ensemble_get_stacking_training_data(
+ toy_dataset, quantiles, quantile_estimator1, quantile_estimator2
+):
+ X, y = toy_dataset
+
+ model = QuantileEnsembleEstimator(
+ estimators=[quantile_estimator1, quantile_estimator2],
+ cv=2,
+ random_state=42,
+ alpha=0.01,
+ )
+
+ (
+ val_indices,
+ val_targets,
+ val_predictions,
+ ) = model._get_stacking_training_data(X, y, quantiles)
+
+ assert len(val_indices) == len(val_targets) == len(X)
+ assert val_predictions.shape[0] == len(X)
+ assert val_predictions.shape[1] == 2 * len(quantiles)
+
+
+@pytest.mark.parametrize("weighting_strategy", ["uniform", "linear_stack"])
+def test_quantile_ensemble_compute_quantile_weights(
+ toy_dataset,
+ quantiles,
+ quantile_estimator1,
+ quantile_estimator2,
+ weighting_strategy,
+):
+ X, y = toy_dataset
+
+ model = QuantileEnsembleEstimator(
+ estimators=[quantile_estimator1, quantile_estimator2],
+ cv=2,
+ weighting_strategy=weighting_strategy,
+ random_state=42,
+ alpha=0.01,
+ )
+
+ weights = model._compute_quantile_weights(X, y, quantiles)
+
+ if weighting_strategy == "uniform":
+ assert weights.shape == (len(quantiles), 2)
+ for w in weights:
+ assert np.isclose(np.sum(w), 1.0)
+ assert np.all(w >= 0)
+ elif weighting_strategy == "linear_stack":
+ assert weights.shape == (len(quantiles), 2)
+ for w in weights:
+ assert np.isclose(np.sum(w), 1.0)
+ assert np.all(w >= 0)
+
+
+def test_quantile_ensemble_predict_quantiles(
+ toy_dataset, quantiles, quantile_estimator1, quantile_estimator2
+):
+ X, _ = toy_dataset
+ n_samples = len(X)
+
+ model = QuantileEnsembleEstimator(
+ estimators=[quantile_estimator1, quantile_estimator2],
+ weighting_strategy="uniform",
+ alpha=0.01,
+ )
+ model.quantiles = quantiles
+ model.quantile_weights = np.array([[0.5, 0.5], [0.5, 0.5], [0.5, 0.5]])
+
+ predictions = model.predict(X)
+ expected = np.tile([3.0, 4.0, 5.0], (n_samples, 1))
+ assert np.array_equal(predictions, expected)
+
+ quantile_estimator1.predict.assert_called_with(X)
+ quantile_estimator2.predict.assert_called_with(X)
+
+
+@pytest.mark.slow
+@pytest.mark.parametrize(
+ "data_fixture_name",
+ [
+ "heteroscedastic_data",
+ "diabetes_data",
+ ],
+)
+@pytest.mark.parametrize("weighting_strategy", ["linear_stack"])
+def test_ensemble_outperforms_components_multiple_repetitions(
+ request,
+ data_fixture_name,
+ weighting_strategy,
+ ensemble_test_quantiles,
+):
+ X, y = request.getfixturevalue(data_fixture_name)
+ X_train, X_test, y_train, y_test = train_test_split(
+ X, y, test_size=0.7, random_state=42
+ )
+
+ scaler = StandardScaler()
+ X_train = scaler.fit_transform(X_train)
+ X_test = scaler.transform(X_test)
+
+ n_repetitions = 10
+ success_threshold = 0.51
+
+ pinball_wins = 0
+
+ for rep in range(n_repetitions):
+ estimators = create_diverse_quantile_estimators(random_state=42 + rep)
+
+ individual_losses = []
+ for estimator in estimators:
+ estimator.fit(X_train, y_train, quantiles=ensemble_test_quantiles)
+ y_pred_individual = estimator.predict(X_test)
+ loss = evaluate_quantile_performance(
+ y_test, y_pred_individual, ensemble_test_quantiles
+ )
+ individual_losses.append(loss)
+
+ best_individual_loss = min(individual_losses)
+
+ ensemble = QuantileEnsembleEstimator(
+ estimators=create_diverse_quantile_estimators(random_state=42 + rep),
+ cv=5,
+ weighting_strategy=weighting_strategy,
+ random_state=42 + rep,
+ alpha=0.01, # Reduced alpha for better performance with quantile Lasso
+ )
+
+ ensemble.fit(X_train, y_train, quantiles=ensemble_test_quantiles)
+ y_pred_ensemble = ensemble.predict(X_test)
+ ensemble_loss = evaluate_quantile_performance(
+ y_test, y_pred_ensemble, ensemble_test_quantiles
+ )
+
+ if ensemble_loss <= best_individual_loss:
+ pinball_wins += 1
+
+ pinball_success_rate = pinball_wins / n_repetitions
+ assert pinball_success_rate > success_threshold
+
+
+@pytest.mark.slow
+@pytest.mark.parametrize(
+ "data_fixture_name",
+ [
+ "heteroscedastic_data",
+ "diabetes_data",
+ ],
+)
+@pytest.mark.parametrize("weighting_strategy", ["linear_stack"])
+def test_point_ensemble_outperforms_components_multiple_repetitions(
+ request,
+ data_fixture_name,
+ weighting_strategy,
+):
+ X, y = request.getfixturevalue(data_fixture_name)
+ X_train, X_test, y_train, y_test = train_test_split(
+ X, y, test_size=0.7, random_state=42
+ )
+
+ scaler = StandardScaler()
+ X_train = scaler.fit_transform(X_train)
+ X_test = scaler.transform(X_test)
+
+ n_repetitions = 10
+ success_threshold = 0.51
+
+ mse_wins = 0
+
+ for rep in range(n_repetitions):
+ estimators = create_diverse_point_estimators(random_state=42 + rep)
+
+ individual_losses = []
+ for estimator in estimators:
+ estimator.fit(X_train, y_train)
+ y_pred_individual = estimator.predict(X_test)
+ loss = evaluate_point_performance(y_test, y_pred_individual)
+ individual_losses.append(loss)
+
+ best_individual_loss = min(individual_losses)
+
+ ensemble = PointEnsembleEstimator(
+ estimators=create_diverse_point_estimators(random_state=42 + rep),
+ cv=5,
+ weighting_strategy=weighting_strategy,
+ random_state=42 + rep,
+ alpha=0.01, # Reduced alpha for better performance
+ )
+
+ ensemble.fit(X_train, y_train)
+ y_pred_ensemble = ensemble.predict(X_test)
+ ensemble_loss = evaluate_point_performance(y_test, y_pred_ensemble)
+
+ if ensemble_loss <= best_individual_loss:
+ mse_wins += 1
+
+ mse_success_rate = mse_wins / n_repetitions
+ assert mse_success_rate > success_threshold
diff --git a/tests/selection/estimators/test_quantile_estimation.py b/tests/selection/estimators/test_quantile_estimation.py
new file mode 100644
index 0000000..5f5187f
--- /dev/null
+++ b/tests/selection/estimators/test_quantile_estimation.py
@@ -0,0 +1,187 @@
+import pytest
+import numpy as np
+from typing import List, Dict, Any
+from sklearn.model_selection import train_test_split
+from sklearn.preprocessing import StandardScaler
+from unittest.mock import Mock
+from confopt.selection.estimators.quantile_estimation import (
+ QuantileLasso,
+ QuantileGBM,
+ QuantileForest,
+ QuantileKNN,
+ QuantileGP,
+ QuantileLeaf,
+ QuantRegWrapper,
+)
+
+
+def assess_quantile_quality(
+ y_true: np.ndarray, predictions: np.ndarray, quantiles: List[float]
+) -> Dict[str, Any]:
+ n_samples, n_quantiles = predictions.shape
+
+ hard_violations = 0
+ soft_violations = 0
+ violation_magnitudes = []
+ tolerance = 1e-6
+
+ for i in range(n_samples):
+ pred_row = predictions[i, :]
+ for j in range(n_quantiles - 1):
+ diff = pred_row[j + 1] - pred_row[j]
+ if diff < -tolerance:
+ hard_violations += 1
+ violation_magnitudes.append(abs(diff))
+ elif abs(diff) <= tolerance:
+ soft_violations += 1
+
+ total_comparisons = n_samples * (n_quantiles - 1)
+
+ coverage_errors = []
+ for i, q in enumerate(quantiles):
+ empirical_coverage = np.mean(y_true <= predictions[:, i])
+ coverage_error = abs(empirical_coverage - q)
+ coverage_errors.append(coverage_error)
+
+ return {
+ "hard_violations": hard_violations,
+ "soft_violations": soft_violations,
+ "hard_rate": hard_violations / total_comparisons,
+ "soft_rate": soft_violations / total_comparisons,
+ "mean_violation_magnitude": np.mean(violation_magnitudes)
+ if violation_magnitudes
+ else 0.0,
+ "coverage_errors": coverage_errors,
+ "mean_coverage_error": np.mean(coverage_errors),
+ "total_comparisons": total_comparisons,
+ }
+
+
+QUALITY_THRESHOLDS = {
+ "single_fit": {
+ "max_hard_violation_rate": 0.0,
+ "max_soft_violation_rate": 0.10,
+ "max_coverage_error": 0.20,
+ },
+ "multi_fit": {
+ "max_hard_violation_rate": 0.08,
+ "max_soft_violation_rate": 0.18,
+ "max_coverage_error": 0.20,
+ },
+}
+
+
+@pytest.mark.parametrize(
+ "data_fixture_name",
+ [
+ "linear_regression_data",
+ "heteroscedastic_data",
+ "diabetes_data",
+ ],
+)
+@pytest.mark.parametrize(
+ "estimator_class,estimator_params,estimator_type",
+ [
+ (
+ QuantileGP,
+ {"kernel": "matern", "random_state": 42},
+ "single_fit",
+ ),
+ (
+ QuantileForest,
+ {"n_estimators": 30, "max_depth": 6, "random_state": 42},
+ "single_fit",
+ ),
+ (
+ QuantileLeaf,
+ {"n_estimators": 30, "max_depth": 6, "random_state": 42},
+ "single_fit",
+ ),
+ (QuantileKNN, {"n_neighbors": 8}, "single_fit"),
+ (
+ QuantileGBM,
+ {
+ "learning_rate": 0.1,
+ "n_estimators": 30,
+ "min_samples_split": 8,
+ "min_samples_leaf": 4,
+ "max_depth": 4,
+ "random_state": 42,
+ },
+ "multi_fit",
+ ),
+ (
+ QuantileLasso,
+ {"max_iter": 1000, "p_tol": 1e-6, "random_state": 42},
+ "multi_fit",
+ ),
+ ],
+)
+def test_quantile_estimator_comprehensive_quality(
+ request,
+ data_fixture_name,
+ estimator_class,
+ estimator_params,
+ estimator_type,
+ comprehensive_test_quantiles,
+):
+ X, y = request.getfixturevalue(data_fixture_name)
+ X_train, X_test, y_train, y_test = train_test_split(
+ X, y, test_size=0.3, random_state=42
+ )
+
+ # Standardize features to avoid penalizing scale-sensitive estimators
+ scaler = StandardScaler()
+ X_train = scaler.fit_transform(X_train)
+ X_test = scaler.transform(X_test)
+
+ quantiles = comprehensive_test_quantiles
+ estimator = estimator_class(**estimator_params)
+
+ try:
+ estimator.fit(X_train, y_train, quantiles)
+ predictions = estimator.predict(X_test)
+ except Exception as e:
+ pytest.fail(
+ "Estimator {} failed on {}: {}".format(
+ estimator_class.__name__, data_fixture_name, str(e)
+ )
+ )
+
+ assert predictions.shape == (len(X_test), len(quantiles))
+ assert not np.any(np.isnan(predictions))
+ assert not np.any(np.isinf(predictions))
+
+ base_thresholds = QUALITY_THRESHOLDS[estimator_type].copy()
+
+ quality_stats = assess_quantile_quality(y_test, predictions, quantiles)
+
+ assert quality_stats["hard_rate"] <= base_thresholds["max_hard_violation_rate"]
+ assert quality_stats["soft_rate"] <= base_thresholds["max_soft_violation_rate"]
+ assert quality_stats["mean_coverage_error"] <= base_thresholds["max_coverage_error"]
+
+
+def test_quantreg_wrapper_with_intercept():
+ mock_results = Mock()
+ mock_results.params = np.array([1.0, 2.0, 3.0])
+
+ wrapper = QuantRegWrapper(mock_results, has_intercept=True)
+ X_test = np.array([[1, 2], [3, 4]])
+
+ predictions = wrapper.predict(X_test)
+ expected = np.array([1 + 1 * 2 + 2 * 3, 1 + 3 * 2 + 4 * 3])
+
+ np.testing.assert_array_equal(predictions, expected)
+
+
+def test_quantreg_wrapper_without_intercept():
+ mock_results = Mock()
+ mock_results.params = np.array([2.0, 3.0])
+
+ wrapper = QuantRegWrapper(mock_results, has_intercept=False)
+ X_test = np.array([[1, 2], [3, 4]])
+
+ predictions = wrapper.predict(X_test)
+ expected = np.array([1 * 2 + 2 * 3, 3 * 2 + 4 * 3])
+
+ np.testing.assert_array_equal(predictions, expected)
diff --git a/tests/selection/sampling/__init__.py b/tests/selection/sampling/__init__.py
new file mode 100644
index 0000000..e69de29
diff --git a/tests/selection/sampling/test_bound_samplers.py b/tests/selection/sampling/test_bound_samplers.py
new file mode 100644
index 0000000..957ed2b
--- /dev/null
+++ b/tests/selection/sampling/test_bound_samplers.py
@@ -0,0 +1,413 @@
+"""
+Tests for bound-based acquisition strategies in conformal prediction optimization.
+
+This module tests the bound-based acquisition samplers that use prediction interval
+bounds for optimization decisions. Tests focus on methodological correctness of
+bound extraction, exploration-exploitation balance, adaptive interval width
+adjustment, and mathematical properties of the acquisition functions.
+
+Test coverage includes:
+- PessimisticLowerBoundSampler: Conservative bound-based acquisition
+- LowerBoundSampler: LCB-style exploration with decay schedules
+- Adaptive interval width mechanisms and coverage feedback
+- Mathematical properties and edge cases
+"""
+
+import pytest
+import numpy as np
+from unittest.mock import patch
+from confopt.selection.sampling.bound_samplers import (
+ PessimisticLowerBoundSampler,
+ LowerBoundSampler,
+)
+
+
+class TestPessimisticLowerBoundSampler:
+ """Test conservative acquisition strategy using pessimistic lower bounds."""
+
+ @pytest.mark.parametrize("interval_width", [0.7, 0.8, 0.9, 0.95])
+ def test_initialization_interval_width(self, interval_width):
+ """Test initialization with different interval widths."""
+ sampler = PessimisticLowerBoundSampler(interval_width=interval_width)
+
+ assert sampler.interval_width == interval_width
+ assert sampler.alpha == 1 - interval_width
+ assert 0 < sampler.alpha < 1
+
+ @pytest.mark.parametrize("adapter", [None, "DtACI", "ACI"])
+ def test_initialization_adapter_types(self, adapter):
+ """Test initialization with different adapter configurations."""
+ sampler = PessimisticLowerBoundSampler(interval_width=0.8, adapter=adapter)
+
+ if adapter is None:
+ assert sampler.adapter is None
+ else:
+ assert sampler.adapter is not None
+
+ def test_fetch_alphas_single_value(self):
+ """Test alpha retrieval returns single value list."""
+ sampler = PessimisticLowerBoundSampler(interval_width=0.85)
+ alphas = sampler.fetch_alphas()
+
+ assert isinstance(alphas, list)
+ assert len(alphas) == 1
+ assert abs(alphas[0] - 0.15) < 1e-10
+
+ def test_fetch_alphas_consistency(self):
+ """Test alpha values remain consistent with interval width."""
+ interval_widths = [0.7, 0.8, 0.9]
+ for width in interval_widths:
+ sampler = PessimisticLowerBoundSampler(interval_width=width)
+ alphas = sampler.fetch_alphas()
+ assert alphas[0] == 1 - width
+
+ @patch("confopt.selection.sampling.bound_samplers.update_single_interval_width")
+ def test_update_interval_width_with_adapter(self, mock_update):
+ """Test interval width update with adapter present."""
+ mock_update.return_value = 0.12
+ sampler = PessimisticLowerBoundSampler(interval_width=0.8, adapter="ACI")
+ original_alpha = sampler.alpha
+
+ sampler.update_interval_width(beta=0.85)
+
+ mock_update.assert_called_once_with(sampler.adapter, original_alpha, 0.85)
+ assert sampler.alpha == 0.12
+
+ @patch("confopt.selection.sampling.bound_samplers.update_single_interval_width")
+ def test_update_interval_width_without_adapter(self, mock_update):
+ """Test interval width update without adapter."""
+ mock_update.return_value = 0.2
+ sampler = PessimisticLowerBoundSampler(interval_width=0.8, adapter=None)
+ original_alpha = sampler.alpha
+
+ sampler.update_interval_width(beta=0.85)
+
+ mock_update.assert_called_once_with(None, original_alpha, 0.85)
+ assert sampler.alpha == 0.2
+
+ @pytest.mark.parametrize("beta", [0.5, 0.75, 0.85, 0.95])
+ def test_update_interval_width_coverage_range(self, beta):
+ """Test update with different coverage rates."""
+ sampler = PessimisticLowerBoundSampler(interval_width=0.8, adapter="ACI")
+ sampler.alpha
+
+ sampler.update_interval_width(beta=beta)
+
+ # Alpha should be adjusted based on coverage
+ assert isinstance(sampler.alpha, float)
+ assert 0 < sampler.alpha < 1
+
+ def test_interval_width_bounds(self):
+ """Test interval width parameter bounds."""
+ # Valid ranges
+ for width in [0.5, 0.8, 0.99]:
+ sampler = PessimisticLowerBoundSampler(interval_width=width)
+ assert 0 < sampler.alpha < 1
+
+ # Edge case: very high confidence
+ sampler = PessimisticLowerBoundSampler(interval_width=0.999)
+ assert abs(sampler.alpha - 0.001) < 1e-10
+
+ def test_alpha_interval_width_relationship(self):
+ """Test mathematical relationship between alpha and interval width."""
+ widths = np.linspace(0.5, 0.95, 10)
+ for width in widths:
+ sampler = PessimisticLowerBoundSampler(interval_width=width)
+ assert abs(sampler.alpha + sampler.interval_width - 1.0) < 1e-10
+
+
+class TestLowerBoundSampler:
+ """Test LCB acquisition strategy with adaptive exploration."""
+
+ @pytest.mark.parametrize("interval_width", [0.7, 0.8, 0.9])
+ @pytest.mark.parametrize("adapter", [None, "DtACI", "ACI"])
+ def test_initialization_inheritance(self, interval_width, adapter):
+ """Test proper inheritance from PessimisticLowerBoundSampler."""
+ sampler = LowerBoundSampler(interval_width=interval_width, adapter=adapter)
+
+ assert sampler.interval_width == interval_width
+ assert sampler.alpha == 1 - interval_width
+ if adapter is None:
+ assert sampler.adapter is None
+ else:
+ assert sampler.adapter is not None
+
+ @pytest.mark.parametrize(
+ "beta_decay", [None, "inverse_square_root_decay", "logarithmic_decay"]
+ )
+ def test_initialization_decay_strategies(self, beta_decay):
+ """Test initialization with different decay strategies."""
+ sampler = LowerBoundSampler(beta_decay=beta_decay)
+
+ assert sampler.beta_decay == beta_decay
+ assert sampler.t == 1
+ assert sampler.beta == 1
+
+ @pytest.mark.parametrize("c", [0.1, 1.0, 5.0, 10.0])
+ def test_initialization_exploration_constant(self, c):
+ """Test initialization with different exploration constants."""
+ sampler = LowerBoundSampler(c=c)
+
+ assert sampler.c == c
+
+ @pytest.mark.parametrize("beta_max", [1.0, 5.0, 10.0, 20.0])
+ def test_initialization_beta_max(self, beta_max):
+ """Test initialization with different maximum beta values."""
+ sampler = LowerBoundSampler(beta_max=beta_max)
+
+ assert sampler.beta_max == beta_max
+
+ def test_time_step_initialization(self):
+ """Test initial time step and exploration parameter."""
+ sampler = LowerBoundSampler()
+
+ assert sampler.t == 1
+ assert sampler.beta == 1
+ assert sampler.mu_max == float("-inf")
+
+ def test_update_exploration_step_time_increment(self):
+ """Test time step increment in exploration update."""
+ sampler = LowerBoundSampler()
+ initial_t = sampler.t
+
+ sampler.update_exploration_step()
+
+ assert sampler.t == initial_t + 1
+
+ @pytest.mark.parametrize(
+ "decay_type", ["inverse_square_root_decay", "logarithmic_decay"]
+ )
+ def test_update_exploration_decay_formulas(self, decay_type):
+ """Test exploration decay formula implementations."""
+ c = 2.0
+ sampler = LowerBoundSampler(beta_decay=decay_type, c=c)
+
+ # Run multiple steps to test decay
+ betas = []
+ for _ in range(10):
+ sampler.update_exploration_step()
+ betas.append(sampler.beta)
+
+ # Beta should generally decrease (with possible fluctuations due to log term)
+ assert betas[-1] < betas[0]
+ assert all(beta >= 0 for beta in betas)
+
+ def test_update_exploration_inverse_square_root_decay(self):
+ """Test inverse square root decay implementation."""
+ c = 4.0
+ sampler = LowerBoundSampler(beta_decay="inverse_square_root_decay", c=c)
+
+ sampler.update_exploration_step() # t=2
+ expected_beta = np.sqrt(c / 2)
+ assert abs(sampler.beta - expected_beta) < 1e-10
+
+ sampler.update_exploration_step() # t=3
+ expected_beta = np.sqrt(c / 3)
+ assert abs(sampler.beta - expected_beta) < 1e-10
+
+ def test_update_exploration_logarithmic_decay(self):
+ """Test logarithmic decay implementation."""
+ c = 2.0
+ sampler = LowerBoundSampler(beta_decay="logarithmic_decay", c=c)
+
+ sampler.update_exploration_step() # t=2
+ expected_beta = np.sqrt((c * np.log(2)) / 2)
+ assert abs(sampler.beta - expected_beta) < 1e-10
+
+ sampler.update_exploration_step() # t=3
+ expected_beta = np.sqrt((c * np.log(3)) / 3)
+ assert abs(sampler.beta - expected_beta) < 1e-10
+
+ def test_update_exploration_no_decay(self):
+ """Test behavior when no decay is specified."""
+ sampler = LowerBoundSampler(beta_decay=None)
+ initial_beta = sampler.beta
+
+ for _ in range(5):
+ sampler.update_exploration_step()
+ assert sampler.beta == initial_beta
+
+ def test_update_exploration_invalid_decay(self):
+ """Test error handling for invalid decay strategies."""
+ sampler = LowerBoundSampler()
+ sampler.beta_decay = "invalid_decay"
+
+ with pytest.raises(ValueError, match="beta_decay must be"):
+ sampler.update_exploration_step()
+
+ def test_calculate_ucb_predictions_basic(self, test_predictions_and_widths):
+ """Test basic LCB calculation functionality."""
+ point_estimates, interval_widths = test_predictions_and_widths
+ sampler = LowerBoundSampler()
+
+ lcb_values = sampler.calculate_ucb_predictions(point_estimates, interval_widths)
+
+ assert lcb_values.shape == point_estimates.shape
+ assert isinstance(lcb_values, np.ndarray)
+
+ def test_calculate_ucb_predictions_formula(self, test_predictions_and_widths):
+ """Test LCB formula implementation."""
+ point_estimates, interval_widths = test_predictions_and_widths
+ beta = 2.0
+ sampler = LowerBoundSampler()
+ sampler.beta = beta
+
+ lcb_values = sampler.calculate_ucb_predictions(point_estimates, interval_widths)
+ expected_values = point_estimates - beta * interval_widths
+
+ np.testing.assert_array_almost_equal(lcb_values, expected_values)
+
+ def test_calculate_ucb_predictions_beta_effect(self, test_predictions_and_widths):
+ """Test effect of different beta values on LCB calculations."""
+ point_estimates, interval_widths = test_predictions_and_widths
+
+ beta_low = LowerBoundSampler()
+ beta_low.beta = 0.5
+
+ beta_high = LowerBoundSampler()
+ beta_high.beta = 3.0
+
+ lcb_low = beta_low.calculate_ucb_predictions(point_estimates, interval_widths)
+ lcb_high = beta_high.calculate_ucb_predictions(point_estimates, interval_widths)
+
+ # Higher beta should lead to lower (more conservative) LCB values
+ assert np.all(lcb_high < lcb_low)
+
+ def test_calculate_ucb_predictions_edge_cases(self):
+ """Test LCB calculation with edge case inputs."""
+ sampler = LowerBoundSampler()
+
+ # Zero interval widths
+ point_estimates = np.array([1, 2, 3])
+ interval_widths = np.zeros(3)
+ lcb_values = sampler.calculate_ucb_predictions(point_estimates, interval_widths)
+ np.testing.assert_array_equal(lcb_values, point_estimates)
+
+ # Single point
+ single_point = np.array([5.0])
+ single_width = np.array([1.0])
+ lcb_single = sampler.calculate_ucb_predictions(single_point, single_width)
+ assert lcb_single.shape == (1,)
+
+ def test_calculate_ucb_predictions_negative_inputs(self):
+ """Test LCB calculation with negative inputs."""
+ sampler = LowerBoundSampler()
+ sampler.beta = 1.5
+
+ point_estimates = np.array([-2, -1, 0, 1, 2])
+ interval_widths = np.array([0.5, 1.0, 1.5, 1.0, 0.5])
+
+ lcb_values = sampler.calculate_ucb_predictions(point_estimates, interval_widths)
+ expected = point_estimates - 1.5 * interval_widths
+
+ np.testing.assert_array_almost_equal(lcb_values, expected)
+
+ @pytest.mark.parametrize("t_steps", [1, 5, 10, 50])
+ def test_exploration_decay_convergence(self, t_steps):
+ """Test exploration parameter convergence over multiple steps."""
+ sampler = LowerBoundSampler(beta_decay="logarithmic_decay", c=1.0)
+
+ for _ in range(t_steps):
+ sampler.update_exploration_step()
+
+ # Beta should decrease as t increases
+ assert sampler.beta < 1.0
+ assert sampler.beta > 0
+ assert sampler.t == t_steps + 1
+
+ def test_exploration_decay_asymptotic_behavior(self):
+ """Test asymptotic behavior of exploration decay."""
+ sampler = LowerBoundSampler(beta_decay="inverse_square_root_decay", c=1.0)
+
+ # Run many steps
+ for _ in range(1000):
+ sampler.update_exploration_step()
+
+ # Beta should be very small but positive
+ assert 0 < sampler.beta < 0.1
+
+ def test_inheritance_method_access(self):
+ """Test access to inherited methods from parent class."""
+ sampler = LowerBoundSampler(interval_width=0.85, adapter="ACI")
+
+ # Should have access to parent methods
+ alphas = sampler.fetch_alphas()
+ assert len(alphas) == 1
+ assert abs(alphas[0] - 0.15) < 1e-10
+
+ # Should be able to update interval width
+ sampler.update_interval_width(beta=0.8)
+ assert isinstance(sampler.alpha, float)
+
+ def test_mathematical_properties_lcb_ordering(self, test_predictions_and_widths):
+ """Test mathematical ordering properties of LCB values."""
+ point_estimates, interval_widths = test_predictions_and_widths
+ sampler = LowerBoundSampler()
+ sampler.beta = 1.0
+
+ lcb_values = sampler.calculate_ucb_predictions(point_estimates, interval_widths)
+
+ # LCB should be lower than point estimates when interval_widths > 0
+ mask = interval_widths > 0
+ assert np.all(lcb_values[mask] <= point_estimates[mask])
+
+ def test_exploration_constant_impact(self, test_predictions_and_widths):
+ """Test impact of exploration constant on acquisition behavior."""
+ point_estimates, interval_widths = test_predictions_and_widths
+
+ sampler_conservative = LowerBoundSampler(c=0.1)
+ sampler_conservative.update_exploration_step()
+
+ sampler_aggressive = LowerBoundSampler(c=10.0)
+ sampler_aggressive.update_exploration_step()
+
+ lcb_conservative = sampler_conservative.calculate_ucb_predictions(
+ point_estimates, interval_widths
+ )
+ lcb_aggressive = sampler_aggressive.calculate_ucb_predictions(
+ point_estimates, interval_widths
+ )
+
+ # Aggressive exploration should lead to lower LCB values
+ assert np.mean(lcb_aggressive) < np.mean(lcb_conservative)
+
+ def test_beta_max_constraint(self):
+ """Test that beta values respect maximum constraint."""
+ beta_max = 5.0
+ sampler = LowerBoundSampler(
+ beta_max=beta_max, c=100.0
+ ) # Large c to potentially exceed beta_max
+
+ # Even with large c, beta should not exceed beta_max in early iterations
+ assert sampler.beta <= beta_max
+
+ @pytest.mark.parametrize("array_size", [1, 10, 100, 1000])
+ def test_calculate_ucb_predictions_scalability(self, array_size):
+ """Test LCB calculation scalability with different array sizes."""
+ sampler = LowerBoundSampler()
+
+ point_estimates = np.random.uniform(-5, 5, array_size)
+ interval_widths = np.random.uniform(0.1, 2.0, array_size)
+
+ lcb_values = sampler.calculate_ucb_predictions(point_estimates, interval_widths)
+
+ assert lcb_values.shape == (array_size,)
+ assert len(lcb_values) == array_size
+
+ def test_state_consistency_after_updates(self):
+ """Test state consistency after multiple operations."""
+ sampler = LowerBoundSampler(interval_width=0.8, adapter="ACI", c=2.0)
+ original_interval_width = sampler.interval_width
+
+ # Perform multiple operations
+ sampler.update_exploration_step()
+ sampler.update_interval_width(beta=0.85)
+ sampler.update_exploration_step()
+
+ # State should remain consistent
+ assert isinstance(sampler.alpha, float)
+ assert 0 < sampler.alpha < 1
+ assert sampler.t >= 1
+ assert sampler.beta >= 0
+ # interval_width remains unchanged even when alpha is updated
+ assert sampler.interval_width == original_interval_width
diff --git a/tests/selection/sampling/test_expected_improvement_samplers.py b/tests/selection/sampling/test_expected_improvement_samplers.py
new file mode 100644
index 0000000..e390de3
--- /dev/null
+++ b/tests/selection/sampling/test_expected_improvement_samplers.py
@@ -0,0 +1,102 @@
+"""
+Tests for Expected Improvement acquisition strategies in conformal prediction optimization.
+
+This module tests the Expected Improvement sampler that estimates expected improvement
+through Monte Carlo sampling from prediction intervals. Tests focus on mathematical
+correctness of EI estimation, exploration-exploitation balance, adaptive interval
+width adjustment, and acquisition function properties.
+
+Test coverage includes:
+- ExpectedImprovementSampler: Monte Carlo EI estimation with conformal intervals
+- Best value tracking and improvement computation accuracy
+- Adaptive interval width mechanisms and coverage feedback
+- Mathematical properties of EI acquisition function
+- Edge cases and boundary conditions
+"""
+
+import pytest
+import numpy as np
+from confopt.selection.sampling.expected_improvement_samplers import (
+ ExpectedImprovementSampler,
+)
+
+
+class TestExpectedImprovementSampler:
+ """Test Expected Improvement acquisition strategy using conformal prediction intervals."""
+
+ @pytest.mark.parametrize("n_quantiles", [4, 6, 8])
+ def test_initialization_even_quantiles(self, n_quantiles):
+ """Test initialization with valid even quantile numbers."""
+ sampler = ExpectedImprovementSampler(n_quantiles=n_quantiles)
+
+ assert sampler.n_quantiles == n_quantiles
+ assert len(sampler.alphas) == n_quantiles // 2
+
+ @pytest.mark.parametrize("n_quantiles", [3, 5, 7])
+ def test_initialization_odd_quantiles_raises_error(self, n_quantiles):
+ """Test that odd quantile numbers raise validation errors."""
+ with pytest.raises(ValueError):
+ ExpectedImprovementSampler(n_quantiles=n_quantiles)
+
+ @pytest.mark.parametrize("adapter", [None, "DtACI", "ACI"])
+ def test_initialization_adapter_types(self, adapter):
+ """Test initialization with different adapter configurations."""
+ sampler = ExpectedImprovementSampler(n_quantiles=4, adapter=adapter)
+
+ if adapter is None:
+ assert sampler.adapters is None
+ else:
+ assert sampler.adapters is not None
+ assert len(sampler.adapters) == len(sampler.alphas)
+
+ def test_update_best_value(self):
+ """Test best value updates with improving values."""
+ sampler = ExpectedImprovementSampler(current_best_value=10.0)
+
+ # Better value should update
+ sampler.update_best_value(5.0)
+ assert sampler.current_best_value == 5.0
+
+ # Should not update if new value is worse
+ sampler.update_best_value(10.0)
+ assert sampler.current_best_value == 5.0
+
+ def test_fetch_alphas_returns_correct_format(self):
+ """Test alpha retrieval returns proper list format."""
+ sampler = ExpectedImprovementSampler(n_quantiles=6)
+ alphas = sampler.fetch_alphas()
+
+ assert isinstance(alphas, list)
+ assert len(alphas) == 3 # n_quantiles // 2
+ assert all(0 < alpha < 1 for alpha in alphas)
+
+ def test_calculate_expected_improvement_negative_values(
+ self, simple_conformal_bounds
+ ):
+ """Test EI values are negative for minimization compatibility."""
+ sampler = ExpectedImprovementSampler(
+ n_quantiles=4, num_ei_samples=20, current_best_value=0.1
+ )
+
+ ei_values = sampler.calculate_expected_improvement(simple_conformal_bounds)
+
+ # All EI values should be non-positive (negated for minimization)
+ assert np.all(ei_values <= 0)
+ n_observations = len(simple_conformal_bounds[0].lower_bounds)
+ assert ei_values.shape == (n_observations,)
+
+ def test_calculate_expected_improvement_deterministic_sampling(
+ self, simple_conformal_bounds
+ ):
+ """Test EI calculation consistency with fixed random seed."""
+ sampler = ExpectedImprovementSampler(n_quantiles=4, num_ei_samples=50)
+
+ # Calculate EI with fixed seed
+ np.random.seed(42)
+ ei_values1 = sampler.calculate_expected_improvement(simple_conformal_bounds)
+
+ np.random.seed(42)
+ ei_values2 = sampler.calculate_expected_improvement(simple_conformal_bounds)
+
+ # Results should be identical with same seed
+ np.testing.assert_array_almost_equal(ei_values1, ei_values2)
diff --git a/tests/selection/sampling/test_sampling_utils.py b/tests/selection/sampling/test_sampling_utils.py
new file mode 100644
index 0000000..82c2aac
--- /dev/null
+++ b/tests/selection/sampling/test_sampling_utils.py
@@ -0,0 +1,142 @@
+import pytest
+import numpy as np
+from confopt.selection.sampling.utils import (
+ initialize_quantile_alphas,
+ initialize_multi_adapters,
+ initialize_single_adapter,
+ update_multi_interval_widths,
+ update_single_interval_width,
+ validate_even_quantiles,
+ flatten_conformal_bounds,
+)
+
+
+@pytest.mark.parametrize("n_quantiles", [2, 4, 6, 8, 10])
+def test_initialize_quantile_alphas_even_counts(n_quantiles):
+ """Test quantile alpha initialization with valid even counts."""
+ alphas = initialize_quantile_alphas(n_quantiles)
+
+ # Should return half the input quantiles
+ assert len(alphas) == n_quantiles // 2
+
+ # All alphas should be in valid range
+ assert all(0 < alpha < 1 for alpha in alphas)
+
+ # Spot check:
+ if n_quantiles == 4:
+ expected_alphas = [0.4, 0.8]
+ np.testing.assert_allclose(alphas, expected_alphas, rtol=1e-10)
+
+
+@pytest.mark.parametrize("n_quantiles", [1, 3, 5, 7])
+def test_initialize_quantile_alphas_odd_counts_raises(n_quantiles):
+ """Test that odd quantile counts raise appropriate errors."""
+ with pytest.raises(ValueError):
+ initialize_quantile_alphas(n_quantiles)
+
+
+def test_update_multi_interval_widths_with_adapters(coverage_feedback):
+ """Test multi-interval width updates with adaptation."""
+ alphas = [0.2, 0.1, 0.05]
+ adapters = initialize_multi_adapters(alphas, "DtACI")
+
+ # Store initial alphas
+ initial_alphas = alphas.copy()
+
+ # Update with coverage feedback
+ updated_alphas = update_multi_interval_widths(adapters, alphas, coverage_feedback)
+
+ # Should return list of same length
+ assert len(updated_alphas) == len(initial_alphas)
+
+ # Alphas should be updated (likely different from initial)
+ assert isinstance(updated_alphas, list)
+ assert all(isinstance(alpha, float) for alpha in updated_alphas)
+
+ # All alphas should remain in valid range
+ assert all(0 < alpha < 1 for alpha in updated_alphas)
+
+
+def test_update_multi_interval_widths_without_adapters():
+ """Test multi-interval width updates without adaptation."""
+ alphas = [0.2, 0.1, 0.05]
+ betas = [0.8, 0.9, 0.95]
+
+ updated_alphas = update_multi_interval_widths(None, alphas, betas)
+
+ # Should return original alphas unchanged
+ assert updated_alphas == alphas
+
+
+def test_update_single_interval_width():
+ """Test single interval width update with adaptation."""
+ alpha = 0.1
+ adapter = initialize_single_adapter(alpha, "DtACI")
+ beta = 0.85
+
+ updated_alpha = update_single_interval_width(adapter, alpha, beta)
+
+ # Should return a float in valid range
+ assert isinstance(updated_alpha, float)
+ assert 0 < updated_alpha < 1
+ assert updated_alpha != alpha # Should be updated
+
+
+def test_validate_even_quantiles_valid():
+ """Test validation passes for even quantiles."""
+ # Should not raise any exception
+ validate_even_quantiles(4, "test_sampler")
+ validate_even_quantiles(6, "another_sampler")
+
+
+@pytest.mark.parametrize("n_quantiles", [1, 3, 5, 7])
+def test_validate_even_quantiles_invalid(n_quantiles):
+ """Test validation raises for odd quantiles."""
+ with pytest.raises(
+ ValueError, match="Number of test_sampler quantiles must be even"
+ ):
+ validate_even_quantiles(n_quantiles, "test_sampler")
+
+
+def test_flatten_conformal_bounds_structure(multi_interval_bounds):
+ """Test conformal bounds flattening produces correct structure."""
+ flattened = flatten_conformal_bounds(multi_interval_bounds)
+
+ n_obs = len(multi_interval_bounds[0].lower_bounds)
+ n_intervals = len(multi_interval_bounds)
+ expected_shape = (n_obs, n_intervals * 2)
+
+ # Should have correct shape
+ assert flattened.shape == expected_shape
+
+ # Should be numpy array
+ assert isinstance(flattened, np.ndarray)
+
+
+def test_flatten_conformal_bounds_interleaving(small_dataset):
+ """Test that bounds are correctly interleaved in flattened representation."""
+ flattened = flatten_conformal_bounds(small_dataset)
+
+ # Check that columns alternate between lower and upper bounds
+ for i, bounds in enumerate(small_dataset):
+ lower_col = i * 2
+ upper_col = i * 2 + 1
+
+ np.testing.assert_array_equal(flattened[:, lower_col], bounds.lower_bounds)
+ np.testing.assert_array_equal(flattened[:, upper_col], bounds.upper_bounds)
+
+
+def test_flatten_conformal_bounds_preserves_intervals(nested_intervals):
+ """Test that flattening preserves interval relationships."""
+ flattened = flatten_conformal_bounds(nested_intervals)
+
+ # Check that nested relationships are preserved
+ for obs_idx in range(flattened.shape[0]):
+ # Extract bounds for this observation
+ wide_lower, wide_upper = flattened[obs_idx, 0], flattened[obs_idx, 1]
+ med_lower, med_upper = flattened[obs_idx, 2], flattened[obs_idx, 3]
+ narrow_lower, narrow_upper = flattened[obs_idx, 4], flattened[obs_idx, 5]
+
+ # Verify nesting: narrow ⊆ medium ⊆ wide
+ assert wide_lower <= med_lower <= narrow_lower
+ assert narrow_upper <= med_upper <= wide_upper
diff --git a/tests/selection/sampling/test_thompson_samplers.py b/tests/selection/sampling/test_thompson_samplers.py
new file mode 100644
index 0000000..1f7f2e2
--- /dev/null
+++ b/tests/selection/sampling/test_thompson_samplers.py
@@ -0,0 +1,204 @@
+import pytest
+import numpy as np
+from confopt.selection.sampling.thompson_samplers import ThompsonSampler
+from confopt.wrapping import ConformalBounds
+
+
+@pytest.mark.parametrize("n_quantiles", [2, 4, 6, 8])
+def test_thompson_sampler_initialization_valid_quantiles(n_quantiles):
+ """Test Thompson sampler initialization with valid even quantile counts."""
+ sampler = ThompsonSampler(n_quantiles=n_quantiles)
+
+ assert len(sampler.alphas) == n_quantiles // 2
+ assert sampler.n_quantiles == n_quantiles
+ assert not sampler.enable_optimistic_sampling
+ assert sampler.adapters is None # Default no adapter
+
+
+@pytest.mark.parametrize("adapter", ["DtACI", "ACI", None])
+def test_thompson_sampler_initialization_with_adapters(adapter):
+ """Test Thompson sampler initialization with different adapter strategies."""
+ sampler = ThompsonSampler(n_quantiles=4, adapter=adapter)
+
+ if adapter is None:
+ assert sampler.adapters is None
+ else:
+ assert len(sampler.adapters) == 2 # n_quantiles // 2
+ assert all(hasattr(a, "update") for a in sampler.adapters)
+
+
+def test_update_interval_width_with_adapters(coverage_feedback):
+ """Test interval width updating with adaptation enabled."""
+ sampler = ThompsonSampler(n_quantiles=6, adapter="DtACI")
+ initial_alphas = sampler.alphas.copy()
+
+ sampler.update_interval_width(coverage_feedback)
+
+ assert len(sampler.alphas) == len(initial_alphas)
+ # Alphas should have changed based on coverage feedback
+ assert not np.array_equal(sampler.alphas, initial_alphas)
+
+
+def test_update_interval_width_without_adapters():
+ """Test interval width updating when no adapters are configured."""
+ sampler = ThompsonSampler(n_quantiles=4, adapter=None)
+ initial_alphas = sampler.alphas.copy()
+ betas = [0.85, 0.92]
+
+ # Should return original alphas unchanged when no adapters
+ sampler.update_interval_width(betas)
+ assert np.array_equal(sampler.alphas, initial_alphas)
+
+
+def test_calculate_thompson_predictions_shape(simple_conformal_bounds):
+ """Test Thompson predictions return correct shape."""
+ sampler = ThompsonSampler(n_quantiles=4)
+ predictions = sampler.calculate_thompson_predictions(simple_conformal_bounds)
+
+ n_observations = len(simple_conformal_bounds[0].lower_bounds)
+ assert predictions.shape == (n_observations,)
+
+
+def test_calculate_thompson_predictions_values_within_bounds(simple_conformal_bounds):
+ """Test that Thompson predictions fall within conformal bounds."""
+ sampler = ThompsonSampler(n_quantiles=4)
+ predictions = sampler.calculate_thompson_predictions(simple_conformal_bounds)
+
+ # Get overall bounds across all intervals
+ all_lower = np.minimum(
+ simple_conformal_bounds[0].lower_bounds, simple_conformal_bounds[1].lower_bounds
+ )
+ all_upper = np.maximum(
+ simple_conformal_bounds[0].upper_bounds, simple_conformal_bounds[1].upper_bounds
+ )
+
+ # All predictions should be within the overall bounds
+ assert np.all(predictions >= all_lower)
+ assert np.all(predictions <= all_upper)
+
+
+@pytest.mark.parametrize("n_quantiles", [2, 4, 6])
+def test_calculate_thompson_predictions_stochasticity(
+ simple_conformal_bounds, n_quantiles
+):
+ """Test that Thompson predictions show appropriate stochastic behavior."""
+ sampler = ThompsonSampler(n_quantiles=n_quantiles)
+
+ # Generate multiple samples
+ samples = []
+ for _ in range(50):
+ predictions = sampler.calculate_thompson_predictions(simple_conformal_bounds)
+ samples.append(predictions)
+
+ samples_array = np.array(samples)
+
+ # Check that predictions vary across runs (stochastic behavior)
+ variance_per_observation = np.var(samples_array, axis=0)
+ assert np.all(variance_per_observation > 0) # Should have non-zero variance
+
+
+def test_calculate_thompson_predictions_optimistic_sampling_enabled(
+ simple_conformal_bounds,
+):
+ """Test Thompson predictions with optimistic sampling enabled."""
+ sampler = ThompsonSampler(n_quantiles=4, enable_optimistic_sampling=True)
+ point_estimates = np.array([0.2, 0.4, 0.6]) # Conservative point estimates
+
+ predictions = sampler.calculate_thompson_predictions(
+ simple_conformal_bounds, point_predictions=point_estimates
+ )
+
+ # Predictions should be capped at point estimates
+ assert np.all(predictions <= point_estimates)
+
+
+def test_calculate_thompson_predictions_mathematical_properties(
+ simple_conformal_bounds,
+):
+ """Test mathematical properties of Thompson sampling distribution.
+
+ Thompson sampling uniformly samples from the flattened bounds matrix,
+ which contains all lower and upper bounds from all intervals.
+ For simple_conformal_bounds, each observation should sample uniformly
+ from the set of bounds: [lower1, upper1, lower2, upper2].
+ """
+ sampler = ThompsonSampler(n_quantiles=4) # Creates 2 intervals
+
+ # Extract expected values for each observation from the bounds
+ expected_values_per_obs = []
+ for obs_idx in range(len(simple_conformal_bounds[0].lower_bounds)):
+ values = [
+ simple_conformal_bounds[0].lower_bounds[obs_idx], # interval 1 lower
+ simple_conformal_bounds[0].upper_bounds[obs_idx], # interval 1 upper
+ simple_conformal_bounds[1].lower_bounds[obs_idx], # interval 2 lower
+ simple_conformal_bounds[1].upper_bounds[obs_idx], # interval 2 upper
+ ]
+ expected_values_per_obs.append(values)
+
+ # Generate many samples for statistical analysis
+ n_samples = 10000
+ samples = []
+ for _ in range(n_samples):
+ predictions = sampler.calculate_thompson_predictions(simple_conformal_bounds)
+ samples.append(predictions)
+
+ samples_array = np.array(samples)
+
+ # For each observation, rigorously test uniform sampling from expected values
+ for obs_idx in range(len(simple_conformal_bounds[0].lower_bounds)):
+ obs_samples = samples_array[:, obs_idx]
+ expected_values = expected_values_per_obs[obs_idx]
+
+ # Test 1: All samples should be from the expected discrete set
+ unique_samples = np.unique(obs_samples)
+ np.testing.assert_array_almost_equal(
+ np.sort(unique_samples),
+ np.sort(expected_values),
+ decimal=10,
+ err_msg=f"Observation {obs_idx} samples not from expected bounds set",
+ )
+
+ # Test 2: Each value should appear with approximately equal frequency (uniform)
+ expected_freq = n_samples / len(expected_values)
+ tolerance = 0.05 * n_samples # 5% tolerance for randomness
+
+ for value in expected_values:
+ actual_freq = np.sum(np.isclose(obs_samples, value))
+ assert abs(actual_freq - expected_freq) < tolerance, (
+ f"Observation {obs_idx}, value {value}: expected ~{expected_freq:.0f} "
+ f"occurrences, got {actual_freq}"
+ )
+
+ # Test 3: Sample mean should equal theoretical mean of uniform distribution
+ theoretical_mean = np.mean(expected_values)
+ sample_mean = np.mean(obs_samples)
+
+ # With large sample size, sample mean should be very close to theoretical
+ mean_tolerance = 0.01 * abs(theoretical_mean) # 1% tolerance
+ assert abs(sample_mean - theoretical_mean) < mean_tolerance, (
+ f"Observation {obs_idx}: theoretical mean {theoretical_mean:.6f}, "
+ f"sample mean {sample_mean:.6f}"
+ )
+
+
+def test_thompson_sampler_deterministic_with_seed():
+ """Test that Thompson sampler produces deterministic results with fixed seed."""
+ sampler = ThompsonSampler(n_quantiles=4)
+
+ # Create fixed bounds
+ bounds = [
+ ConformalBounds(
+ lower_bounds=np.array([0.1, 0.2]), upper_bounds=np.array([0.5, 0.6])
+ )
+ ]
+
+ # Set seed and get predictions
+ np.random.seed(42)
+ predictions1 = sampler.calculate_thompson_predictions(bounds)
+
+ # Reset seed and get predictions again
+ np.random.seed(42)
+ predictions2 = sampler.calculate_thompson_predictions(bounds)
+
+ # Should be identical with same seed
+ np.testing.assert_array_equal(predictions1, predictions2)
diff --git a/tests/selection/test_acquisition.py b/tests/selection/test_acquisition.py
new file mode 100644
index 0000000..2f31b92
--- /dev/null
+++ b/tests/selection/test_acquisition.py
@@ -0,0 +1,206 @@
+import pytest
+import numpy as np
+from confopt.selection.acquisition import (
+ QuantileConformalSearcher,
+)
+from confopt.selection.sampling.bound_samplers import (
+ PessimisticLowerBoundSampler,
+ LowerBoundSampler,
+)
+from confopt.selection.sampling.thompson_samplers import ThompsonSampler
+from confopt.selection.sampling.expected_improvement_samplers import (
+ ExpectedImprovementSampler,
+)
+
+from conftest import (
+ QUANTILE_ESTIMATOR_ARCHITECTURES,
+)
+
+
+@pytest.mark.parametrize(
+ "sampler_class,sampler_kwargs",
+ [
+ (PessimisticLowerBoundSampler, {"interval_width": 0.8}),
+ (LowerBoundSampler, {"interval_width": 0.8}),
+ (ThompsonSampler, {"n_quantiles": 4}),
+ (ExpectedImprovementSampler, {"n_quantiles": 4}),
+ ],
+)
+@pytest.mark.parametrize("quantile_arch", QUANTILE_ESTIMATOR_ARCHITECTURES[:1])
+def test_quantile_conformal_searcher(
+ sampler_class, sampler_kwargs, quantile_arch, big_toy_dataset
+):
+ X, y = big_toy_dataset
+ X_train, y_train = X[:7], y[:7]
+ X_val, y_val = X[7:], y[7:]
+
+ sampler = sampler_class(**sampler_kwargs)
+ searcher = QuantileConformalSearcher(
+ quantile_estimator_architecture=quantile_arch,
+ sampler=sampler,
+ n_pre_conformal_trials=5,
+ )
+
+ # Combine train and val data for new interface
+ X_combined = np.vstack((X_train, X_val))
+ y_combined = np.concatenate((y_train, y_val))
+ searcher.fit(
+ X=X_combined,
+ y=y_combined,
+ tuning_iterations=0,
+ random_state=42,
+ )
+
+ predictions = searcher.predict(X_val)
+ assert len(predictions) == len(X_val)
+
+ X_update = X_val[0].reshape(1, -1)
+ y_update = y_val[0]
+ initial_X_train_len = len(searcher.X_train)
+ initial_y_train_len = len(searcher.y_train)
+
+ searcher.update(X_update, y_update)
+
+ # Data doesn't change, only updates samplers and other states:
+ assert len(searcher.X_train) == initial_X_train_len
+ assert len(searcher.y_train) == initial_y_train_len
+
+
+def test_quantile_searcher_prediction_methods(big_toy_dataset):
+ X, y = big_toy_dataset
+ X_train, y_train = X[:7], y[:7]
+ X_val, y_val = X[7:], y[7:]
+ X_test = X_val
+
+ lb_sampler = LowerBoundSampler(interval_width=0.8, beta_decay=None)
+ lb_searcher = QuantileConformalSearcher(
+ quantile_estimator_architecture="ql",
+ sampler=lb_sampler,
+ n_pre_conformal_trials=5,
+ )
+ # Combine train and val data for new interface
+ X_combined = np.vstack((X_train, X_val))
+ y_combined = np.concatenate((y_train, y_val))
+ lb_searcher.fit(
+ X=X_combined,
+ y=y_combined,
+ tuning_iterations=0,
+ random_state=42,
+ )
+ lb_predictions = lb_searcher.predict(X_test)
+ assert len(lb_predictions) == len(X_test)
+
+ thompson_sampler = ThompsonSampler(n_quantiles=4)
+ thompson_searcher = QuantileConformalSearcher(
+ quantile_estimator_architecture="ql",
+ sampler=thompson_sampler,
+ n_pre_conformal_trials=5,
+ )
+ thompson_searcher.fit(
+ X=X_combined,
+ y=y_combined,
+ tuning_iterations=0,
+ random_state=42,
+ )
+ thompson_predictions = thompson_searcher.predict(X_test)
+ assert len(thompson_predictions) == len(X_test)
+
+ ei_sampler = ExpectedImprovementSampler(n_quantiles=4, current_best_value=0.5)
+ ei_searcher = QuantileConformalSearcher(
+ quantile_estimator_architecture="ql",
+ sampler=ei_sampler,
+ n_pre_conformal_trials=5,
+ )
+ ei_searcher.fit(
+ X=X_combined,
+ y=y_combined,
+ tuning_iterations=0,
+ random_state=42,
+ )
+ ei_predictions = ei_searcher.predict(X_test)
+ assert len(ei_predictions) == len(X_test)
+
+ plb_sampler = PessimisticLowerBoundSampler(interval_width=0.8)
+ plb_searcher = QuantileConformalSearcher(
+ quantile_estimator_architecture="ql",
+ sampler=plb_sampler,
+ n_pre_conformal_trials=5,
+ )
+ plb_searcher.fit(
+ X=X_combined,
+ y=y_combined,
+ tuning_iterations=0,
+ random_state=42,
+ )
+ plb_predictions = plb_searcher.predict(X_test)
+ assert len(plb_predictions) == len(X_test)
+
+
+@pytest.mark.parametrize("current_best_value", [0.0, 0.5, 1.0, 10.0])
+def test_expected_improvement_best_value_update(current_best_value, big_toy_dataset):
+ """Test that Expected Improvement properly tracks and updates best values."""
+ X, y = big_toy_dataset
+ X_train, y_train = X[:10], y[:10]
+ X_val, y_val = X[10:20], y[10:20]
+
+ sampler = ExpectedImprovementSampler(
+ n_quantiles=4, current_best_value=current_best_value
+ )
+ searcher = QuantileConformalSearcher(
+ quantile_estimator_architecture="ql",
+ sampler=sampler,
+ n_pre_conformal_trials=5,
+ )
+
+ # Combine train and val data for new interface
+ X_combined = np.vstack((X_train, X_val))
+ y_combined = np.concatenate((y_train, y_val))
+ searcher.fit(X=X_combined, y=y_combined, tuning_iterations=0, random_state=42)
+
+ # Test that sampler has correct initial best value
+ assert sampler.current_best_value == current_best_value
+
+ # Test update with better value (remember: we minimize, so lower is better)
+ new_value = current_best_value - 1.0
+ searcher.update(X_val[0], new_value)
+ assert sampler.current_best_value == new_value
+
+ # Test update with worse value (should not change)
+ worse_value = current_best_value + 1.0
+ searcher.update(X_val[1], worse_value)
+ assert sampler.current_best_value == new_value # Should remain the better value
+
+
+def test_adaptive_alpha_updating(big_toy_dataset):
+ """Test that adaptive alpha updating works correctly for compatible samplers."""
+ X, y = big_toy_dataset
+ X_train, y_train = X[:15], y[:15]
+ X_val, y_val = X[15:30], y[15:30]
+
+ # Test with adaptive sampler
+ sampler = LowerBoundSampler(interval_width=0.8, adapter="DtACI")
+ searcher = QuantileConformalSearcher(
+ quantile_estimator_architecture="ql",
+ sampler=sampler,
+ n_pre_conformal_trials=5,
+ )
+
+ # Combine train and val data for new interface
+ X_combined = np.vstack((X_train, X_val))
+ y_combined = np.concatenate((y_train, y_val))
+ searcher.fit(X=X_combined, y=y_combined, tuning_iterations=0, random_state=42)
+
+ # Store initial alpha values
+ initial_alphas = searcher.sampler.fetch_alphas().copy()
+
+ # Perform several updates
+ for i in range(3):
+ test_point = X_val[i]
+ test_value = y_val[i]
+ searcher.update(test_point, test_value)
+
+ # Check that alphas change:
+ final_alphas = searcher.sampler.fetch_alphas()
+ assert len(final_alphas) == len(initial_alphas)
+ assert all(0 < alpha < 1 for alpha in final_alphas)
+ assert not np.array_equal(initial_alphas, final_alphas)
diff --git a/tests/selection/test_adaptation.py b/tests/selection/test_adaptation.py
new file mode 100644
index 0000000..0b520eb
--- /dev/null
+++ b/tests/selection/test_adaptation.py
@@ -0,0 +1,607 @@
+import numpy as np
+import pytest
+from sklearn.linear_model import LinearRegression
+from confopt.selection.adaptation import DtACI, pinball_loss
+
+
+class SimpleACI:
+ """Simplified ACI implementation from Gibbs & Candès (2021) paper.
+
+ This implements the basic adaptive conformal inference algorithm with the simple update:
+ α_{t+1} = α_t + γ(α - err_t)
+
+ where err_t is a binary indicator: 1 for breach/error, 0 for coverage/no error.
+ This follows the exact formula from equation (2) in the paper.
+ This is used only for testing equivalence with DTACI when using a single gamma value.
+ """
+
+ def __init__(self, alpha: float = 0.1, gamma: float = 0.01):
+ """Initialize Simple ACI.
+
+ Args:
+ alpha: Target miscoverage level (α ∈ (0,1))
+ gamma: Learning rate for alpha updates
+ """
+ if not 0 < alpha < 1:
+ raise ValueError("alpha must be in (0, 1)")
+ if gamma <= 0:
+ raise ValueError("gamma must be positive")
+
+ self.alpha = alpha
+ self.gamma = gamma
+ self.alpha_t = alpha
+ self.alpha_history = []
+
+ def update(self, err_t: int) -> float:
+ """Update alpha based on binary error indicator.
+
+ Args:
+ err_t: Binary error indicator (1 = breach/error, 0 = coverage/no error)
+
+ Returns:
+ Updated miscoverage level α_t+1
+ """
+ if err_t not in [0, 1]:
+ raise ValueError(f"err_t must be 0 or 1, got {err_t}")
+
+ # Simple ACI update from paper: α_{t+1} = α_t + γ(α - err_t)
+ self.alpha_t = self.alpha_t + self.gamma * (self.alpha - err_t)
+ self.alpha_t = np.clip(self.alpha_t, 0.001, 0.999)
+
+ self.alpha_history.append(self.alpha_t)
+ return self.alpha_t
+
+
+class StaticCI:
+ def __init__(self, alpha: float = 0.1):
+ if not 0 < alpha < 1:
+ raise ValueError("alpha must be in (0, 1)")
+ self.alpha = alpha
+ self.alpha_t = alpha
+ self.alpha_history = []
+
+ def update(self, beta: float) -> float:
+ if not 0 <= beta <= 1:
+ raise ValueError(f"beta must be in [0, 1], got {beta}")
+ self.alpha_history.append(self.alpha_t)
+ return self.alpha_t
+
+
+def run_conformal_performance_test(method, X, y, target_alpha, gamma_values=None):
+ """Helper function to run conformal prediction performance tests and return metrics.
+
+ Args:
+ method: Either 'dtaci' or 'static' to specify which method to test
+ X, y: Data for testing
+ target_alpha: Target miscoverage level
+ gamma_values: Learning rates for DtACI (ignored for static method)
+
+ Returns:
+ Dictionary with performance metrics
+ """
+ if method == "dtaci":
+ if gamma_values is None:
+ gamma_values = [0.01, 0.05, 0.1]
+ predictor = DtACI(alpha=target_alpha, gamma_values=gamma_values)
+ elif method == "static":
+ predictor = StaticCI(alpha=target_alpha)
+ else:
+ raise ValueError("method must be 'dtaci' or 'static'")
+
+ breaches = []
+ alpha_evolution = []
+ initial_window = 30
+
+ for i in range(initial_window, len(X)):
+ X_past = X[:i]
+ y_past = y[:i]
+ X_test = X[i].reshape(1, -1)
+ y_test = y[i]
+
+ n_cal = max(int(len(X_past) * 0.3), 10)
+ X_train, X_cal = X_past[:-n_cal], X_past[-n_cal:]
+ y_train, y_cal = y_past[:-n_cal], y_past[-n_cal:]
+
+ model = LinearRegression()
+ model.fit(X_train, y_train)
+ y_cal_pred = model.predict(X_cal)
+ cal_residuals = np.abs(y_cal - y_cal_pred)
+ y_test_pred = model.predict(X_test)[0]
+
+ test_residual = abs(y_test - y_test_pred)
+ beta = np.mean(cal_residuals >= test_residual)
+
+ current_alpha = predictor.update(beta=beta)
+ alpha_evolution.append(current_alpha)
+
+ # Check breach
+ quantile = np.quantile(cal_residuals, 1 - current_alpha, method="linear")
+ lower = y_test_pred - quantile
+ upper = y_test_pred + quantile
+ breach = int(not (lower <= y_test <= upper))
+ breaches.append(breach)
+
+ coverage = 1 - np.mean(breaches)
+ target_coverage = 1 - target_alpha
+ coverage_error = abs(coverage - target_coverage)
+
+ return {
+ "coverage_error": coverage_error,
+ "alpha_variance": np.var(alpha_evolution),
+ "alpha_range": max(alpha_evolution) - min(alpha_evolution),
+ "alpha_evolution": alpha_evolution,
+ }
+
+
+@pytest.mark.parametrize("gamma", [0.01, 0.05, 0.1])
+@pytest.mark.parametrize("target_alpha", [0.1, 0.2])
+def test_dtaci_simple_aci_equivalence(gamma, target_alpha):
+ """Test that DTACI with single gamma produces identical results to SimpleACI.
+
+ Uses empirical quantile definition that matches conformal theory to ensure
+ exact mathematical equivalence between beta-based (DtACI) and interval-based
+ (SimpleACI) error signals. The algorithms should produce identical alpha histories."""
+ np.random.seed(42)
+
+ # Initialize both algorithms with same parameters
+ dtaci = DtACI(alpha=target_alpha, gamma_values=[gamma], use_weighted_average=True)
+ simple_aci = SimpleACI(alpha=target_alpha, gamma=gamma)
+
+ # Generate synthetic data for testing
+ n_samples = 100
+ X = np.random.randn(n_samples, 2)
+ y = X[:, 0] + 0.5 * X[:, 1] + 0.1 * np.random.randn(n_samples)
+
+ dtaci_alphas = []
+ simple_aci_alphas = []
+
+ # Simulate online conformal prediction
+ for i in range(30, n_samples):
+ # Split data
+ X_past = X[:i]
+ y_past = y[:i]
+ X_test = X[i].reshape(1, -1)
+ y_test = y[i]
+
+ # Use simple train/calibration split
+ n_cal = 20
+ X_train, X_cal = X_past[:-n_cal], X_past[-n_cal:]
+ y_train, y_cal = y_past[:-n_cal], y_past[-n_cal:]
+
+ model = LinearRegression()
+ model.fit(X_train, y_train)
+
+ y_cal_pred = model.predict(X_cal)
+ cal_residuals = np.abs(y_cal - y_cal_pred)
+ y_test_pred = model.predict(X_test)[0]
+ test_residual = abs(y_test - y_test_pred)
+
+ current_alpha = dtaci.alpha_t
+
+ # Compute interval coverage using empirical quantile that matches conformal theory
+ # This ensures exact equivalence with beta calculation
+ sorted_residuals = np.sort(cal_residuals)
+ n_cal = len(cal_residuals)
+ target_count = (1 - current_alpha) * n_cal
+ k = int(np.floor(target_count))
+ if k == 0:
+ quantile = sorted_residuals[0]
+ elif k >= n_cal:
+ quantile = sorted_residuals[-1]
+ else:
+ quantile = sorted_residuals[k]
+ lower_bound = y_test_pred - quantile
+ upper_bound = y_test_pred + quantile
+ covered = int(lower_bound <= y_test <= upper_bound)
+ err_t = int(not covered) # 1 if not covered (breach), 0 if covered
+
+ beta = np.mean(cal_residuals >= test_residual)
+ dtaci_alpha = dtaci.update(beta=beta)
+ simple_aci_alpha = simple_aci.update(err_t=err_t)
+
+ dtaci_alphas.append(dtaci_alpha)
+ simple_aci_alphas.append(simple_aci_alpha)
+
+ # Alpha updates should be identical
+ assert np.allclose(dtaci_alphas, simple_aci_alphas, atol=1e-12)
+
+ # Alpha histories should be identical
+ assert np.allclose(dtaci.alpha_history, simple_aci.alpha_history, atol=1e-12)
+
+
+def test_simple_aci_basic_functionality():
+ """Test basic functionality of SimpleACI class."""
+ aci = SimpleACI(alpha=0.1, gamma=0.01)
+
+ # Test initialization
+ assert aci.alpha == 0.1
+ assert aci.gamma == 0.01
+ assert aci.alpha_t == 0.1
+ assert len(aci.alpha_history) == 0
+
+ # Test update with breach (err_t = 1)
+ alpha_new = aci.update(err_t=1) # breach, err_t = 1
+ expected_alpha = 0.1 + 0.01 * (0.1 - 1) # 0.1 + 0.01 * (-0.9) = 0.091
+ assert abs(alpha_new - expected_alpha) < 1e-12
+ assert len(aci.alpha_history) == 1
+
+ # Test update with coverage (err_t = 0)
+ alpha_new = aci.update(err_t=0) # coverage, err_t = 0
+ expected_alpha = expected_alpha + 0.01 * (0.1 - 0) # 0.091 + 0.01 * 0.1 = 0.092
+ assert abs(alpha_new - expected_alpha) < 1e-12
+ assert len(aci.alpha_history) == 2
+
+
+def test_simple_aci_parameter_validation():
+ """Test parameter validation for SimpleACI."""
+ # Test invalid alpha
+ with pytest.raises(ValueError, match="alpha must be in"):
+ SimpleACI(alpha=0.0)
+
+ with pytest.raises(ValueError, match="alpha must be in"):
+ SimpleACI(alpha=1.0)
+
+ # Test invalid gamma
+ with pytest.raises(ValueError, match="gamma must be positive"):
+ SimpleACI(alpha=0.1, gamma=0.0)
+
+ with pytest.raises(ValueError, match="gamma must be positive"):
+ SimpleACI(alpha=0.1, gamma=-0.01)
+
+ # Test invalid err_t in update
+ aci = SimpleACI(alpha=0.1, gamma=0.01)
+ with pytest.raises(ValueError, match="err_t must be 0 or 1"):
+ aci.update(err_t=-1)
+
+ with pytest.raises(ValueError, match="err_t must be 0 or 1"):
+ aci.update(err_t=2)
+
+
+@pytest.mark.parametrize(
+ "beta,theta,alpha,expected",
+ [
+ (0.8, 0.9, 0.1, 0.09),
+ (0.95, 0.9, 0.1, 0.005),
+ (0.9, 0.9, 0.1, 0.0),
+ (0.5, 0.8, 0.2, 0.24),
+ (0.7, 0.6, 0.3, 0.03),
+ ],
+)
+def test_pinball_loss_mathematical_correctness(beta, theta, alpha, expected):
+ """Test pinball loss calculation matches theoretical formula from paper."""
+ result = pinball_loss(beta=beta, theta=theta, alpha=alpha)
+ assert abs(result - expected) < 1e-10
+
+
+def test_pinball_loss_asymmetric_penalty():
+ """Test pinball loss correctly implements asymmetric penalty structure."""
+ alpha = 0.1
+ theta = 0.9
+
+ # Under-coverage case (beta < theta)
+ under_coverage_loss = pinball_loss(beta=0.8, theta=theta, alpha=alpha)
+ # Over-coverage case (beta > theta)
+ over_coverage_loss = pinball_loss(beta=1.0, theta=theta, alpha=alpha)
+
+ # Under-coverage: ℓ(0.8, 0.9) = 0.1*(0.8-0.9) - min{0, 0.8-0.9} = -0.01 - (-0.1) = 0.09
+ # Over-coverage: ℓ(1.0, 0.9) = 0.1*(1.0-0.9) - min{0, 1.0-0.9} = 0.01 - 0 = 0.01
+ assert abs(under_coverage_loss - 0.09) < 1e-10
+ assert abs(over_coverage_loss - 0.01) < 1e-10
+ # Under-coverage should be penalized more than over-coverage
+ assert under_coverage_loss > over_coverage_loss
+
+
+def test_pinball_loss_properties():
+ """Test general mathematical properties of pinball loss function."""
+ alpha = 0.1
+
+ # Test non-negativity and zero at equality
+ for beta in [0.0, 0.3, 0.5, 0.7, 1.0]:
+ for theta in [0.1, 0.4, 0.6, 0.9]:
+ loss = pinball_loss(beta, theta, alpha)
+ assert loss >= 0
+
+ if abs(beta - theta) < 1e-10:
+ assert abs(loss) < 1e-10
+
+
+@pytest.mark.parametrize("alpha", [0.05, 0.1, 0.2, 0.5])
+def test_dtaci_initialization_parameters(alpha):
+ """Test DtACI initializes with correct theoretical parameters."""
+ dtaci = DtACI(alpha=alpha)
+
+ # Check theoretical parameter formulas
+ expected_eta = (
+ np.sqrt(3 / dtaci.interval)
+ * np.sqrt(np.log(dtaci.interval * dtaci.k) + 2)
+ / ((1 - alpha) ** 2 * alpha**2)
+ )
+ expected_sigma = 1 / (2 * dtaci.interval)
+
+ assert abs(dtaci.eta - expected_eta) < 1e-10
+ assert abs(dtaci.sigma - expected_sigma) < 1e-12
+ assert np.allclose(dtaci.alpha_t_candidates, alpha)
+ assert np.allclose(dtaci.weights, 1.0 / dtaci.k)
+ assert abs(np.sum(dtaci.weights) - 1.0) < 1e-10
+
+
+def test_dtaci_invalid_parameters():
+ """Test DtACI raises appropriate errors for invalid parameters."""
+ with pytest.raises(ValueError, match="alpha must be in"):
+ DtACI(alpha=0.0)
+
+ with pytest.raises(ValueError, match="alpha must be in"):
+ DtACI(alpha=1.0)
+
+ with pytest.raises(ValueError, match="gamma values must be positive"):
+ DtACI(alpha=0.1, gamma_values=[0.1, 0.0, 0.2])
+
+
+@pytest.mark.parametrize("beta", [0.0, 0.25, 0.5, 0.75, 1.0])
+def test_dtaci_update_weight_normalization(beta, dtaci_instance):
+ """Test that expert weights remain valid and probabilities can be computed."""
+ for _ in range(10):
+ dtaci_instance.update(beta=beta)
+ # Weights should be non-negative but not necessarily normalized
+ assert np.all(dtaci_instance.weights >= 0)
+ # Should be able to compute valid probabilities
+ weight_sum = np.sum(dtaci_instance.weights)
+ assert (
+ weight_sum > 0
+ ), "Weight sum should be positive for probability computation"
+ probabilities = dtaci_instance.weights / weight_sum
+ assert abs(np.sum(probabilities) - 1.0) < 1e-10
+ assert np.all(probabilities >= 0)
+ # Alpha values should remain in valid range
+ assert np.all(dtaci_instance.alpha_t_candidates > 0)
+ assert np.all(dtaci_instance.alpha_t_candidates < 1)
+
+
+def test_dtaci_theoretical_weight_updates():
+ """Test that weight updates follow theoretical exponential weighting scheme."""
+ dtaci = DtACI(alpha=0.1, gamma_values=[0.01, 0.05])
+
+ initial_weights = dtaci.weights.copy()
+ initial_alphas = dtaci.alpha_t_candidates.copy()
+
+ beta = 0.85
+ dtaci.update(beta=beta)
+
+ # Manually compute expected weight update following the paper's approach
+ losses = np.array(
+ [
+ pinball_loss(beta=beta, theta=alpha_val, alpha=dtaci.alpha)
+ for alpha_val in initial_alphas
+ ]
+ )
+
+ updated_weights = initial_weights * np.exp(-dtaci.eta * losses)
+ sum_of_updated_weights = np.sum(updated_weights)
+ expected_regularized = (1 - dtaci.sigma) * updated_weights + (
+ (dtaci.sigma * sum_of_updated_weights) / dtaci.k
+ )
+
+ assert np.allclose(dtaci.weights, expected_regularized, atol=1e-12)
+
+
+def test_dtaci_expert_alpha_updates():
+ """Test expert alpha values are updated correctly according to theoretical formula."""
+ dtaci = DtACI(alpha=0.1, gamma_values=[0.01, 0.05])
+
+ initial_alphas = dtaci.alpha_t_candidates.copy()
+ beta = 0.85
+ dtaci.update(beta=beta)
+
+ # Verify alpha updates follow: α_t+1^i = α_t^i + γ_i * (α - err_t^i)
+ for i, (initial_alpha, gamma) in enumerate(zip(initial_alphas, dtaci.gamma_values)):
+ err_indicator = float(beta < initial_alpha)
+ expected_alpha = initial_alpha + gamma * (dtaci.alpha - err_indicator)
+ expected_alpha = np.clip(expected_alpha, 0.001, 0.999)
+
+ assert abs(dtaci.alpha_t_candidates[i] - expected_alpha) < 1e-12
+
+
+def test_dtaci_both_selection_methods():
+ """Test that both random sampling and weighted average methods work correctly."""
+ np.random.seed(42)
+ target_alpha = 0.1
+
+ for use_weighted_average in [True, False]:
+ dtaci = DtACI(
+ alpha=target_alpha,
+ gamma_values=[0.01, 0.05],
+ use_weighted_average=use_weighted_average,
+ )
+
+ # Test with series of beta values
+ betas = [0.85, 0.92, 0.88, 0.95, 0.80]
+ alphas = [dtaci.update(beta=beta) for beta in betas]
+
+ # Both methods should produce valid alphas
+ assert all(0.001 <= alpha <= 0.999 for alpha in alphas)
+ # Should show adaptation behavior
+ assert len(set(np.round(alphas, 6))) > 1
+
+
+def test_dtaci_convergence_under_stationary_conditions():
+ """Test DtACI behavior under stationary conditions."""
+ dtaci = DtACI(alpha=0.1, gamma_values=[0.01, 0.02, 0.05])
+
+ # Test under conditions where target coverage is achieved
+ # Use a beta value that should lead to equilibrium near the target alpha
+ target_beta = 0.1 # This should lead to equilibrium around alpha = 0.1
+ alpha_history = []
+
+ for _ in range(500):
+ alpha_t = dtaci.update(beta=target_beta)
+ alpha_history.append(alpha_t)
+
+ # Under stationary conditions, alpha should be relatively stable
+ recent_alphas = alpha_history[-100:]
+ alpha_variance = np.var(recent_alphas)
+ alpha_mean = np.mean(recent_alphas)
+
+ assert alpha_variance < 0.01
+ # With beta = alpha, the algorithm should converge to a value close to alpha
+ assert abs(alpha_mean - dtaci.alpha) < 0.1
+
+
+def test_dtaci_directional_behavior():
+ """Test that DtACI adjusts alpha in the correct direction based on beta values.
+
+ This test verifies the fundamental adaptive behavior:
+ - When beta > alpha (coverage achieved): alpha should increase slightly toward target
+ - When beta < alpha (breach occurred): alpha should decrease more significantly away from target
+
+ This follows the ACI update rule: α_{t+1} = α_t + γ(α - err_t)
+ where err_t = 1 if breach (β < α), 0 if coverage (β ≥ α).
+ """
+ for beta in [0.8, 0.05]:
+ dtaci = DtACI(alpha=0.1, gamma_values=[0.01])
+ initial_alpha = dtaci.alpha_t
+ updated_alpha = dtaci.update(beta=beta)
+ if beta > initial_alpha:
+ assert updated_alpha > initial_alpha
+ else:
+ assert updated_alpha < initial_alpha
+
+
+def test_dtaci_algorithm_behavior():
+ """Test comprehensive DtACI algorithm behavior and theoretical correctness."""
+ dtaci = DtACI(alpha=0.1, gamma_values=[0.01, 0.05])
+
+ # Test algorithm components work as specified
+ betas = [0.85, 0.92, 0.88, 0.95, 0.80]
+
+ for beta in betas:
+ prev_weights = dtaci.weights.copy()
+ prev_alphas = dtaci.alpha_t_candidates.copy()
+
+ alpha_t = dtaci.update(beta=beta)
+
+ # Verify weights remain valid (non-negative and positive sum)
+ assert np.all(dtaci.weights >= 0)
+ assert np.sum(dtaci.weights) > 0
+
+ # Verify weights change when losses differ
+ losses = [
+ pinball_loss(beta, alpha_val, dtaci.alpha) for alpha_val in prev_alphas
+ ]
+ if not np.allclose(losses, losses[0]):
+ assert not np.allclose(dtaci.weights, prev_weights, atol=1e-10)
+
+ # Verify alpha values are in valid range
+ assert np.all(dtaci.alpha_t_candidates >= 0.001)
+ assert np.all(dtaci.alpha_t_candidates <= 0.999)
+ assert 0.001 <= alpha_t <= 0.999
+
+ # Test algorithm adaptation over time
+ alphas_sequence = [dtaci.update(beta=beta) for beta in betas]
+ unique_alphas = len(set(np.round(alphas_sequence, 6)))
+ assert unique_alphas > 1
+
+
+@pytest.mark.parametrize("target_alpha", [0.1, 0.2, 0.5])
+def test_dtaci_moderate_shift_performance(moderate_shift_data, target_alpha):
+ """Test DtACI performance under moderate distribution shift."""
+ X, y = moderate_shift_data
+ results = run_conformal_performance_test("dtaci", X, y, target_alpha)
+
+ tolerance = 0.05
+
+ assert results["coverage_error"] < tolerance
+ # Should show adaptation behavior
+ assert results["alpha_variance"] > 0.00001
+ assert results["alpha_range"] > 0.0001
+
+
+@pytest.mark.parametrize("target_alpha", [0.1, 0.2, 0.5])
+def test_dtaci_high_shift_performance(high_shift_data, target_alpha):
+ """Test DtACI performance under high distribution shift."""
+ X, y = high_shift_data
+ results = run_conformal_performance_test("dtaci", X, y, target_alpha)
+
+ tolerance = 0.05
+
+ assert results["coverage_error"] < tolerance
+ # Should show significant adaptation behavior under high shift
+ assert results["alpha_variance"] > 0.00001
+ assert results["alpha_range"] > 0.005
+
+
+def generate_shifted_data(
+ n_points=300, shift_points=None, noise_levels=None, random_seed=42
+):
+ """Generate synthetic data with distribution shifts for testing adaptive methods.
+
+ Args:
+ n_points: Total number of data points
+ shift_points: Points where distribution shifts occur
+ noise_levels: Noise levels for each segment
+ random_seed: Random seed for reproducibility
+
+ Returns:
+ X, y: Feature matrix and target vector
+ """
+ if shift_points is None:
+ shift_points = [80, 160, 240]
+ if noise_levels is None:
+ noise_levels = [0.1, 0.6, 0.2, 0.8]
+
+ np.random.seed(random_seed)
+
+ segments = []
+ start_idx = 0
+
+ for i, shift_point in enumerate(shift_points + [n_points]):
+ segment_size = shift_point - start_idx
+ X_segment = np.random.randn(segment_size, 2)
+ y_segment = X_segment.sum(axis=1) + noise_levels[i] * np.random.randn(
+ segment_size
+ )
+ segments.append((X_segment, y_segment))
+ start_idx = shift_point
+
+ X = np.vstack([seg[0] for seg in segments])
+ y = np.hstack([seg[1] for seg in segments])
+
+ return X, y
+
+
+@pytest.mark.parametrize("target_alpha", [0.1, 0.2])
+def test_dtaci_vs_static_conformal_multiple_repetitions(target_alpha):
+ """Test that DtACI outperforms static conformal prediction on highly shifted data.
+
+ Runs multiple random repetitions and verifies that DtACI achieves better
+ performance than static conformal prediction at least 75% of the time.
+ """
+ n_repetitions = 20
+ dtaci_wins = 0
+ dtaci_errors = []
+ static_errors = []
+
+ gamma_values = [0.01, 0.05, 0.1]
+
+ for rep in range(n_repetitions):
+ X, y = generate_shifted_data(random_seed=42 + rep)
+
+ # Run both methods using the consolidated function
+ dtaci_results = run_conformal_performance_test(
+ "dtaci", X, y, target_alpha, gamma_values
+ )
+ static_results = run_conformal_performance_test("static", X, y, target_alpha)
+
+ dtaci_errors.append(dtaci_results["coverage_error"])
+ static_errors.append(static_results["coverage_error"])
+
+ if dtaci_results["coverage_error"] < static_results["coverage_error"]:
+ dtaci_wins += 1
+
+ # DtACI should win at least 75% of the time
+ win_rate = dtaci_wins / n_repetitions
+ assert win_rate >= 0.75
+
+ # Additionally, DtACI should have better average performance
+ avg_dtaci_error = np.mean(dtaci_errors)
+ avg_static_error = np.mean(static_errors)
+ assert avg_dtaci_error <= avg_static_error
diff --git a/tests/selection/test_conformalization.py b/tests/selection/test_conformalization.py
new file mode 100644
index 0000000..3bf60b6
--- /dev/null
+++ b/tests/selection/test_conformalization.py
@@ -0,0 +1,266 @@
+import numpy as np
+import pytest
+from confopt.selection.conformalization import (
+ QuantileConformalEstimator,
+ alpha_to_quantiles,
+)
+from confopt.wrapping import ConformalBounds
+from confopt.utils.preprocessing import train_val_split
+from conftest import (
+ AMENDED_SINGLE_FIT_QUANTILE_ESTIMATOR_ARCHITECTURES,
+ AMENDED_QUANTILE_ESTIMATOR_ARCHITECTURES,
+)
+
+POINT_ESTIMATOR_COVERAGE_TOLERANCE = 0.15
+QUANTILE_ESTIMATOR_COVERAGE_TOLERANCE = 0.15
+MINIMUM_CONFORMAL_WIN_RATE = 0.51
+
+# Optional per-architecture tolerance overrides for rare problematic estimators
+ARCH_TOLERANCE_OVERRIDES: dict[str, float] = {
+ # Example only (keep empty unless specific architectures are identified):
+ # "problem_arch": 0.10,
+}
+
+
+def validate_intervals(
+ intervals: list[ConformalBounds],
+ y_true: np.ndarray,
+ alphas: list[float],
+ tolerance: float,
+) -> tuple[float, bool]:
+ coverages = []
+ errors = []
+ for i, alpha in enumerate(alphas):
+ lower_bound = intervals[i].lower_bounds
+ upper_bound = intervals[i].upper_bounds
+ coverage = np.mean((y_true >= lower_bound) & (y_true <= upper_bound))
+ error = abs(coverage - (1 - alpha)) > tolerance
+
+ coverages.append(coverage)
+ errors.append(error)
+
+ return coverages, errors
+
+
+@pytest.mark.parametrize("alpha", [0.1, 0.2, 0.3])
+def test_alpha_to_quantiles(alpha):
+ lower, upper = alpha_to_quantiles(alpha)
+ assert lower == alpha / 2
+ assert upper == 1 - alpha / 2
+ assert lower <= upper
+
+
+@pytest.mark.slow
+@pytest.mark.parametrize(
+ "data_fixture_name",
+ ["diabetes_data"],
+)
+@pytest.mark.parametrize(
+ "estimator_architecture", AMENDED_QUANTILE_ESTIMATOR_ARCHITECTURES
+)
+@pytest.mark.parametrize("tuning_iterations", [0])
+@pytest.mark.parametrize("alphas", [[0.1], [0.1, 0.3, 0.9]])
+@pytest.mark.parametrize(
+ "calibration_split_strategy", ["train_test_split", "cv", "adaptive"]
+)
+def test_quantile_fit_and_predict_intervals_shape_and_coverage(
+ request,
+ data_fixture_name,
+ estimator_architecture,
+ tuning_iterations,
+ alphas,
+ calibration_split_strategy,
+):
+ X, y = request.getfixturevalue(data_fixture_name)
+ (X_train, y_train, X_test, y_test,) = train_val_split(
+ X, y, train_split=0.8, normalize=False, ordinal=False, random_state=42
+ )
+
+ estimator = QuantileConformalEstimator(
+ quantile_estimator_architecture=estimator_architecture,
+ alphas=alphas,
+ n_pre_conformal_trials=15,
+ n_calibration_folds=3,
+ calibration_split_strategy=calibration_split_strategy,
+ )
+ estimator.fit(
+ X=X_train,
+ y=y_train,
+ tuning_iterations=tuning_iterations,
+ random_state=42,
+ )
+ assert len(estimator.fold_scores_per_alpha) == len(alphas)
+
+ intervals = estimator.predict_intervals(X_test)
+ assert len(intervals) == len(alphas)
+
+ tol = ARCH_TOLERANCE_OVERRIDES.get(
+ estimator_architecture, QUANTILE_ESTIMATOR_COVERAGE_TOLERANCE
+ )
+ _, errors = validate_intervals(intervals, y_test, alphas, tol)
+ assert not any(errors)
+
+
+def test_quantile_calculate_betas_output_properties(
+ dummy_expanding_quantile_gaussian_dataset,
+):
+ estimator = QuantileConformalEstimator(
+ quantile_estimator_architecture=AMENDED_QUANTILE_ESTIMATOR_ARCHITECTURES[0],
+ alphas=[0.1, 0.2, 0.3],
+ n_pre_conformal_trials=15,
+ )
+ X, y = dummy_expanding_quantile_gaussian_dataset
+ X_train, y_train, X_val, y_val = train_val_split(
+ X, y, train_split=0.8, normalize=False, ordinal=False, random_state=42
+ )
+ estimator.fit(X=X_train, y=y_train, random_state=42)
+ test_point = X_val[0]
+ test_value = y_val[0]
+ betas = estimator.calculate_betas(test_point, test_value)
+ assert len(betas) == len(estimator.alphas)
+ assert all(0 <= beta <= 1 for beta in betas)
+
+
+@pytest.mark.parametrize(
+ "n_trials,expected_conformalize",
+ [
+ (5, False),
+ (50, True),
+ ],
+)
+def test_quantile_conformalization_decision_logic(n_trials, expected_conformalize):
+ estimator = QuantileConformalEstimator(
+ quantile_estimator_architecture=AMENDED_SINGLE_FIT_QUANTILE_ESTIMATOR_ARCHITECTURES[
+ 0
+ ],
+ alphas=[0.2],
+ n_pre_conformal_trials=20,
+ )
+ total_size = n_trials
+ X = np.random.rand(total_size, 3)
+ y = np.random.rand(total_size)
+ X_train, y_train, _, _ = train_val_split(
+ X, y, train_split=0.8, normalize=False, ordinal=False, random_state=42
+ )
+ estimator.fit(X=X_train, y=y_train)
+ assert estimator.conformalize_predictions == expected_conformalize
+
+
+@pytest.mark.parametrize(
+ "initial_alphas,new_alphas",
+ [
+ ([0.2], [0.15, 0.25]),
+ ([0.1, 0.2], [0.05, 0.15, 0.3]),
+ ([0.3], [0.1]),
+ ],
+)
+def test_quantile_alpha_update_mechanism(initial_alphas, new_alphas):
+ estimator = QuantileConformalEstimator(
+ quantile_estimator_architecture=AMENDED_QUANTILE_ESTIMATOR_ARCHITECTURES[0],
+ alphas=initial_alphas,
+ )
+ estimator.update_alphas(new_alphas)
+ assert estimator.updated_alphas == new_alphas
+ assert estimator.alphas == initial_alphas
+
+
+@pytest.mark.slow
+@pytest.mark.parametrize(
+ "data_fixture_name",
+ [
+ # "heteroscedastic_data",
+ "diabetes_data",
+ ],
+)
+@pytest.mark.parametrize("estimator_architecture", ["qrf", "qgbm"])
+@pytest.mark.parametrize("alphas", [[0.2, 0.4, 0.6, 0.8]])
+@pytest.mark.parametrize("calibration_split_strategy", ["cv"])
+def test_conformalized_vs_non_conformalized_quantile_estimator_coverage(
+ request,
+ data_fixture_name,
+ estimator_architecture,
+ alphas,
+ calibration_split_strategy,
+):
+ X, y = request.getfixturevalue(data_fixture_name)
+
+ n_repeats = 10
+ np.random.seed(42)
+ random_states = [np.random.randint(0, 10000) for _ in range(n_repeats)]
+ better_or_equal_count = 0
+ for random_state in random_states:
+ (X_train, y_train, X_test, y_test,) = train_val_split(
+ X,
+ y,
+ # A low value, given we care about distributional coverage
+ # on hold out set and we want to simulate a finite training dataset:
+ train_split=0.7,
+ normalize=False,
+ ordinal=False,
+ random_state=random_state,
+ )
+
+ conformalized_estimator = QuantileConformalEstimator(
+ quantile_estimator_architecture=estimator_architecture,
+ alphas=alphas,
+ n_pre_conformal_trials=32,
+ calibration_split_strategy=calibration_split_strategy,
+ n_calibration_folds=5,
+ normalize_features=True,
+ )
+
+ conformalized_estimator.fit(
+ X=X_train,
+ y=y_train,
+ random_state=random_state,
+ )
+
+ non_conformalized_estimator = QuantileConformalEstimator(
+ quantile_estimator_architecture=estimator_architecture,
+ alphas=alphas,
+ n_pre_conformal_trials=10000,
+ calibration_split_strategy=calibration_split_strategy,
+ n_calibration_folds=5,
+ normalize_features=True,
+ )
+
+ non_conformalized_estimator.fit(
+ X=X_train,
+ y=y_train,
+ random_state=random_state,
+ )
+
+ assert conformalized_estimator.conformalize_predictions
+ assert not non_conformalized_estimator.conformalize_predictions
+
+ conformalized_intervals = conformalized_estimator.predict_intervals(X_test)
+ non_conformalized_intervals = non_conformalized_estimator.predict_intervals(
+ X_test
+ )
+ conformalized_coverages, _ = validate_intervals(
+ conformalized_intervals,
+ y_test,
+ alphas,
+ QUANTILE_ESTIMATOR_COVERAGE_TOLERANCE,
+ )
+ non_conformalized_coverages, _ = validate_intervals(
+ non_conformalized_intervals,
+ y_test,
+ alphas,
+ QUANTILE_ESTIMATOR_COVERAGE_TOLERANCE,
+ )
+
+ for i, alpha in enumerate(alphas):
+ target_coverage = 1 - alpha
+ conformalized_coverage = conformalized_coverages[i]
+ non_conformalized_coverage = non_conformalized_coverages[i]
+
+ conformalized_error = abs(conformalized_coverage - target_coverage)
+ non_conformalized_error = abs(non_conformalized_coverage - target_coverage)
+
+ if conformalized_error <= non_conformalized_error:
+ better_or_equal_count += 1
+
+ total_comparisons = n_repeats * len(alphas)
+ percentage_better_or_equal = better_or_equal_count / total_comparisons
+ assert percentage_better_or_equal >= MINIMUM_CONFORMAL_WIN_RATE
diff --git a/tests/selection/test_estimation.py b/tests/selection/test_estimation.py
new file mode 100644
index 0000000..f75b5fa
--- /dev/null
+++ b/tests/selection/test_estimation.py
@@ -0,0 +1,136 @@
+import pytest
+
+from confopt.selection.estimation import (
+ initialize_estimator,
+ average_scores_across_folds,
+)
+
+from confopt.selection.estimator_configuration import ESTIMATOR_REGISTRY
+
+
+@pytest.mark.parametrize("estimator_architecture", list(ESTIMATOR_REGISTRY.keys()))
+def test_initialize_estimator_returns_expected_type(estimator_architecture):
+ """Test that initialize_estimator returns the correct estimator type."""
+ estimator = initialize_estimator(estimator_architecture, random_state=42)
+ expected_class = ESTIMATOR_REGISTRY[estimator_architecture].estimator_class
+ assert isinstance(estimator, expected_class)
+
+
+@pytest.mark.parametrize("random_state", [None, 42, 123])
+def test_initialize_estimator_with_random_state(random_state):
+ """Test that random_state is properly set when supported by estimator."""
+ estimator = initialize_estimator(
+ estimator_architecture="gbm",
+ initialization_params={"random_state": 42} if random_state else {},
+ random_state=random_state,
+ )
+ assert estimator.random_state == random_state
+
+
+@pytest.mark.parametrize("split_type", ["k_fold", "ordinal_split"])
+@pytest.mark.parametrize("n_searches", [1, 3, 10])
+def test_point_tuner_returns_valid_configuration(
+ point_tuner, estimation_test_data, split_type, n_searches
+):
+ """Test that PointTuner returns a valid configuration for different search counts."""
+ X_train, X_val, y_train, y_val = estimation_test_data
+
+ # Use an estimator we know exists
+ estimator_architecture = "gbm"
+ estimator_config = ESTIMATOR_REGISTRY[estimator_architecture]
+
+ best_config = point_tuner.tune(
+ X_train,
+ y_train,
+ estimator_architecture,
+ n_searches=n_searches,
+ train_split=0.8,
+ split_type=split_type,
+ )
+
+ # Configuration should be a dictionary
+ assert isinstance(best_config, dict)
+
+ # All parameter keys should be valid for this estimator
+ valid_params = set(estimator_config.estimator_parameter_space.keys())
+ assert set(best_config.keys()).issubset(valid_params)
+
+
+@pytest.mark.parametrize("split_type", ["k_fold", "ordinal_split"])
+def test_quantile_tuner_returns_valid_configuration(
+ quantile_tuner_with_quantiles, estimation_test_data, split_type
+):
+ """Test that QuantileTuner returns valid configuration for quantile estimators."""
+ tuner, quantiles = quantile_tuner_with_quantiles
+ X_train, X_val, y_train, y_val = estimation_test_data
+
+ # Find a quantile estimator
+ quantile_architectures = [
+ arch
+ for arch, config in ESTIMATOR_REGISTRY.items()
+ if config.is_quantile_estimator()
+ ]
+ estimator_architecture = quantile_architectures[0]
+ estimator_config = ESTIMATOR_REGISTRY[estimator_architecture]
+
+ best_config = tuner.tune(
+ X_train,
+ y_train,
+ estimator_architecture,
+ n_searches=3,
+ train_split=0.8,
+ split_type=split_type,
+ )
+
+ # Configuration should be a dictionary
+ assert isinstance(best_config, dict)
+
+ # All parameter keys should be valid for this estimator
+ valid_params = set(estimator_config.estimator_parameter_space.keys())
+ assert set(best_config.keys()).issubset(valid_params)
+
+
+def test_tuning_with_forced_configurations_prioritizes_them(
+ point_tuner, estimation_test_data
+):
+ """Test that forced configurations are prioritized in tuning process."""
+ X_train, X_val, y_train, y_val = estimation_test_data
+
+ estimator_architecture = "gbm"
+ estimator_config = ESTIMATOR_REGISTRY[estimator_architecture]
+ forced_config = estimator_config.default_params
+
+ best_config = point_tuner.tune(
+ X_train,
+ y_train,
+ estimator_architecture,
+ n_searches=1, # Only one search, should return forced config
+ train_split=0.8,
+ split_type="ordinal_split",
+ forced_param_configurations=[forced_config],
+ )
+
+ assert best_config == forced_config
+
+
+def test_correct_averaging_and_ordering():
+ """Test that order of unique configurations is preserved during averaging."""
+ configs = [
+ {"param": "first"},
+ {"param": "second"},
+ {"param": "first"}, # duplicate
+ {"param": "third"},
+ ]
+ scores = [1.0, 2.0, 3.0, 4.0]
+
+ unique_configs, unique_scores = average_scores_across_folds(configs, scores)
+
+ # First unique should be "first", second should be "second", third should be "third"
+ assert unique_configs[0]["param"] == "first"
+ assert unique_configs[1]["param"] == "second"
+ assert unique_configs[2]["param"] == "third"
+
+ # Check scores are averaged correctly
+ assert unique_scores[0] == 2.0 # (1.0 + 3.0) / 2
+ assert unique_scores[1] == 2.0
+ assert unique_scores[2] == 4.0
diff --git a/tests/test_estimation.py b/tests/test_estimation.py
deleted file mode 100644
index 42a2b1d..0000000
--- a/tests/test_estimation.py
+++ /dev/null
@@ -1,470 +0,0 @@
-from typing import Dict
-
-import numpy as np
-import pytest
-
-from confopt.config import GBM_NAME, RF_NAME, QGBM_NAME, QRF_NAME
-from confopt.estimation import (
- QuantileConformalRegression,
- LocallyWeightedConformalRegression,
- initialize_point_estimator,
- initialize_quantile_estimator,
- cross_validate_configurations,
-)
-
-DEFAULT_SEED = 1234
-DEFAULT_SEARCH_POINT_ESTIMATOR = GBM_NAME
-DEFAULT_SEARCH_QUANTILE_ESTIMATOR = QRF_NAME
-
-
-def get_discretized_quantile_dict(
- X: np.array, y: np.array, quantile_level: float
-) -> Dict:
- """
- Helper function to create dictionary of quantiles per X value.
-
- Parameters
- ----------
- X :
- Explanatory variables.
- y :
- Target variable.
- quantile_level :
- Desired quantile to take.
-
- Returns
- -------
- quantile_dict :
- Dictionary relating X values to their quantile.
- """
- quantile_dict = {}
- for discrete_x_coordinate in np.unique(X):
- conditional_y_at_x = y[X == discrete_x_coordinate]
- quantile_dict[discrete_x_coordinate] = np.quantile(
- conditional_y_at_x, quantile_level
- )
- return quantile_dict
-
-
-def test_initialize_point_estimator():
- initialized_estimator = initialize_point_estimator(
- estimator_architecture=DEFAULT_SEARCH_POINT_ESTIMATOR,
- initialization_params={},
- random_state=DEFAULT_SEED,
- )
-
- assert hasattr(initialized_estimator, "predict")
-
-
-def test_initialize_point_estimator__reproducibility():
- initialized_estimator_first_call = initialize_point_estimator(
- estimator_architecture=DEFAULT_SEARCH_POINT_ESTIMATOR,
- initialization_params={},
- random_state=DEFAULT_SEED,
- )
- initialized_estimator_second_call = initialize_point_estimator(
- estimator_architecture=DEFAULT_SEARCH_POINT_ESTIMATOR,
- initialization_params={},
- random_state=DEFAULT_SEED,
- )
- assert (
- initialized_estimator_first_call.random_state
- == initialized_estimator_second_call.random_state
- )
-
-
-def test_initialize_quantile_estimator():
- dummy_pinball_loss_alpha = [0.25, 0.75]
-
- initialized_estimator = initialize_quantile_estimator(
- estimator_architecture=DEFAULT_SEARCH_QUANTILE_ESTIMATOR,
- initialization_params={},
- pinball_loss_alpha=dummy_pinball_loss_alpha,
- random_state=DEFAULT_SEED,
- )
-
- assert hasattr(initialized_estimator, "predict")
-
-
-def test_initialize_quantile_estimator__reproducibility():
- dummy_pinball_loss_alpha = [0.25, 0.75]
-
- initialized_estimator_first_call = initialize_quantile_estimator(
- estimator_architecture=DEFAULT_SEARCH_QUANTILE_ESTIMATOR,
- initialization_params={},
- pinball_loss_alpha=dummy_pinball_loss_alpha,
- random_state=DEFAULT_SEED,
- )
- initialized_estimator_second_call = initialize_quantile_estimator(
- estimator_architecture=DEFAULT_SEARCH_QUANTILE_ESTIMATOR,
- initialization_params={},
- pinball_loss_alpha=dummy_pinball_loss_alpha,
- random_state=DEFAULT_SEED,
- )
-
- assert (
- initialized_estimator_first_call.random_state
- == initialized_estimator_second_call.random_state
- )
-
-
-def test_cross_validate_configurations__point_estimator(
- dummy_gbm_configurations, dummy_stationary_gaussian_dataset
-):
- X, y = (
- dummy_stationary_gaussian_dataset[:, 0].reshape(-1, 1),
- dummy_stationary_gaussian_dataset[:, 1],
- )
-
- scored_configurations, scores = cross_validate_configurations(
- configurations=dummy_gbm_configurations,
- estimator_architecture=DEFAULT_SEARCH_POINT_ESTIMATOR,
- X=X,
- y=y,
- k_fold_splits=3,
- random_state=DEFAULT_SEED,
- )
-
- assert len(scored_configurations) == len(scores)
- assert len(scored_configurations) == len(dummy_gbm_configurations)
-
- stringified_scored_configurations = []
- for configuration in scored_configurations:
- stringified_scored_configurations.append(
- str(dict(sorted(configuration.items())))
- )
- assert sorted(list(set(stringified_scored_configurations))) == sorted(
- stringified_scored_configurations
- )
-
- for score in scores:
- assert score >= 0
-
-
-def test_cross_validate_configurations__point_estimator__reproducibility(
- dummy_gbm_configurations, dummy_stationary_gaussian_dataset
-):
- X, y = (
- dummy_stationary_gaussian_dataset[:, 0].reshape(-1, 1),
- dummy_stationary_gaussian_dataset[:, 1],
- )
-
- (
- scored_configurations_first_call,
- scores_first_call,
- ) = cross_validate_configurations(
- configurations=dummy_gbm_configurations,
- estimator_architecture=DEFAULT_SEARCH_POINT_ESTIMATOR,
- X=X,
- y=y,
- k_fold_splits=3,
- random_state=DEFAULT_SEED,
- )
- (
- scored_configurations_second_call,
- scores_second_call,
- ) = cross_validate_configurations(
- configurations=dummy_gbm_configurations,
- estimator_architecture=DEFAULT_SEARCH_POINT_ESTIMATOR,
- X=X,
- y=y,
- k_fold_splits=3,
- random_state=DEFAULT_SEED,
- )
-
- assert scored_configurations_first_call == scored_configurations_second_call
- assert scores_first_call == scores_second_call
-
-
-@pytest.mark.parametrize("confidence_level", [0.2, 0.8])
-@pytest.mark.parametrize("tuning_param_combinations", [0, 1, 3])
-@pytest.mark.parametrize("quantile_estimator_architecture", [QGBM_NAME, QRF_NAME])
-def test_quantile_conformal_regression__fit(
- dummy_fixed_quantile_dataset,
- confidence_level,
- tuning_param_combinations,
- quantile_estimator_architecture,
-):
- X, y = (
- dummy_fixed_quantile_dataset[:, 0].reshape(-1, 1),
- dummy_fixed_quantile_dataset[:, 1],
- )
- train_split = 0.8
- X_train, y_train = (
- X[: round(len(X) * train_split), :],
- y[: round(len(y) * train_split)],
- )
- X_val, y_val = X[round(len(X) * train_split) :, :], y[round(len(y) * train_split) :]
-
- qcr = QuantileConformalRegression(
- quantile_estimator_architecture=quantile_estimator_architecture,
- )
- qcr.fit(
- X_train=X_train,
- y_train=y_train,
- X_val=X_val,
- y_val=y_val,
- confidence_level=confidence_level,
- tuning_iterations=tuning_param_combinations,
- random_state=DEFAULT_SEED,
- )
-
- assert qcr.nonconformity_scores is not None
- assert qcr.quantile_estimator is not None
-
-
-@pytest.mark.parametrize("confidence_level", [0.2, 0.8])
-@pytest.mark.parametrize("tuning_param_combinations", [5])
-@pytest.mark.parametrize("quantile_estimator_architecture", [QGBM_NAME, QRF_NAME])
-def test_quantile_conformal_regression__predict(
- dummy_fixed_quantile_dataset,
- confidence_level,
- tuning_param_combinations,
- quantile_estimator_architecture,
-):
- X, y = (
- dummy_fixed_quantile_dataset[:, 0].reshape(-1, 1),
- dummy_fixed_quantile_dataset[:, 1],
- )
- train_split = 0.8
- X_train, y_train = (
- X[: round(len(X) * train_split), :],
- y[: round(len(y) * train_split)],
- )
- X_val, y_val = X[round(len(X) * train_split) :, :], y[round(len(y) * train_split) :]
-
- qcr = QuantileConformalRegression(
- quantile_estimator_architecture=quantile_estimator_architecture,
- )
- qcr.fit(
- X_train=X_train,
- y_train=y_train,
- X_val=X_val,
- y_val=y_val,
- confidence_level=confidence_level,
- tuning_iterations=tuning_param_combinations,
- random_state=DEFAULT_SEED,
- )
- y_low_bounds, y_high_bounds = qcr.predict(X_val, confidence_level=confidence_level)
-
- # Check lower bound is always lower than higher bound:
- for y_low, y_high in zip(y_low_bounds, y_high_bounds):
- assert y_low <= y_high
-
- # Compute observed quantiles per X slice during training
- # (would only work for univariate dummy datasets):
- low_quantile_dict_train = get_discretized_quantile_dict(
- X_train.reshape(
- -1,
- ),
- y_train,
- confidence_level + ((1 - confidence_level) / 2),
- )
- high_quantile_dict_train = get_discretized_quantile_dict(
- X_train.reshape(
- -1,
- ),
- y_train,
- (1 - confidence_level) / 2,
- )
- # Check that predictions return observed quantiles during training
- # Prediction error deviations of more than this amount
- # will count as a breach:
- y_breach_threshold = 1
- # More than this percentage of breaches will fail the test:
- breach_tolerance = 0.3
- low_margin_breaches, high_margin_breaches = 0, 0
- for x_obs, y_low, y_high in zip(
- X_train.reshape(
- -1,
- ),
- y_low_bounds,
- y_high_bounds,
- ):
- if abs(y_low - low_quantile_dict_train[x_obs]) > y_breach_threshold:
- low_margin_breaches += 1
- if abs(y_high - high_quantile_dict_train[x_obs]) > y_breach_threshold:
- high_margin_breaches += 1
- assert low_margin_breaches < len(X_train) * breach_tolerance
- assert high_margin_breaches < len(X_train) * breach_tolerance
-
- # Check conformal interval coverage on validation data
- # (note validation data is actively used by the searcher
- # to calibrate its conformal intervals, so this is not an
- # OOS test, just a sanity check):
- interval_breach_states = []
- for y_obs, y_low, y_high in zip(y_val, y_low_bounds, y_high_bounds):
- is_interval_breach = 0 if y_high > y_obs > y_low else 1
- interval_breach_states.append(is_interval_breach)
-
- interval_breach_rate = sum(interval_breach_states) / len(interval_breach_states)
- breach_margin = 0.01
- assert (
- (confidence_level - breach_margin)
- <= (1 - interval_breach_rate)
- <= (confidence_level + breach_margin)
- )
-
-
-@pytest.mark.parametrize("confidence_level", [0.2, 0.8])
-@pytest.mark.parametrize("tuning_param_combinations", [0, 1, 3])
-@pytest.mark.parametrize("point_estimator_architecture", [GBM_NAME, RF_NAME])
-@pytest.mark.parametrize("demeaning_estimator_architecture", [GBM_NAME])
-@pytest.mark.parametrize("variance_estimator_architecture", [GBM_NAME])
-def test_locally_weighted_conformal_regression__fit(
- dummy_fixed_quantile_dataset,
- confidence_level,
- tuning_param_combinations,
- point_estimator_architecture,
- demeaning_estimator_architecture,
- variance_estimator_architecture,
-):
- X, y = (
- dummy_fixed_quantile_dataset[:, 0].reshape(-1, 1),
- dummy_fixed_quantile_dataset[:, 1],
- )
- train_split = 0.8
- X_train, y_train = (
- X[: round(len(X) * train_split), :],
- y[: round(len(y) * train_split)],
- )
- pe_split = 0.8
- X_pe, y_pe = (
- X_train[: round(len(X_train) * pe_split), :],
- y_train[: round(len(y_train) * pe_split)],
- )
- X_ve, y_ve = (
- X_train[round(len(X_train) * pe_split) :, :],
- y_train[round(len(y_train) * pe_split) :],
- )
- X_val, y_val = X[round(len(X) * train_split) :, :], y[round(len(y) * train_split) :]
-
- lwcr = LocallyWeightedConformalRegression(
- point_estimator_architecture=point_estimator_architecture,
- demeaning_estimator_architecture=demeaning_estimator_architecture,
- variance_estimator_architecture=variance_estimator_architecture,
- )
- lwcr.fit(
- X_pe=X_pe,
- y_pe=y_pe,
- X_ve=X_ve,
- y_ve=y_ve,
- X_val=X_val,
- y_val=y_val,
- tuning_iterations=tuning_param_combinations,
- random_state=DEFAULT_SEED,
- )
-
- assert lwcr.nonconformity_scores is not None
- assert lwcr.pe_estimator is not None
- assert lwcr.ve_estimator is not None
-
-
-@pytest.mark.parametrize("confidence_level", [0.2, 0.8])
-@pytest.mark.parametrize("tuning_param_combinations", [5])
-@pytest.mark.parametrize("point_estimator_architecture", [GBM_NAME, RF_NAME])
-@pytest.mark.parametrize("demeaning_estimator_architecture", [GBM_NAME])
-@pytest.mark.parametrize("variance_estimator_architecture", [GBM_NAME])
-def test_locally_weighted_conformal_regression__predict(
- dummy_fixed_quantile_dataset,
- confidence_level,
- tuning_param_combinations,
- point_estimator_architecture,
- demeaning_estimator_architecture,
- variance_estimator_architecture,
-):
- X, y = (
- dummy_fixed_quantile_dataset[:, 0].reshape(-1, 1),
- dummy_fixed_quantile_dataset[:, 1],
- )
- train_split = 0.8
- X_train, y_train = (
- X[: round(len(X) * train_split), :],
- y[: round(len(y) * train_split)],
- )
- pe_split = 0.8
- X_pe, y_pe = (
- X_train[: round(len(X_train) * pe_split), :],
- y_train[: round(len(y_train) * pe_split)],
- )
- X_ve, y_ve = (
- X_train[round(len(X_train) * pe_split) :, :],
- y_train[round(len(y_train) * pe_split) :],
- )
- X_val, y_val = X[round(len(X) * train_split) :, :], y[round(len(y) * train_split) :]
-
- lwcr = LocallyWeightedConformalRegression(
- point_estimator_architecture=point_estimator_architecture,
- demeaning_estimator_architecture=demeaning_estimator_architecture,
- variance_estimator_architecture=variance_estimator_architecture,
- )
- lwcr.fit(
- X_pe=X_pe,
- y_pe=y_pe,
- X_ve=X_ve,
- y_ve=y_ve,
- X_val=X_val,
- y_val=y_val,
- tuning_iterations=tuning_param_combinations,
- random_state=DEFAULT_SEED,
- )
-
- y_low_bounds, y_high_bounds = lwcr.predict(X_val, confidence_level=confidence_level)
-
- # Check lower bound is always lower than higher bound:
- for y_low, y_high in zip(y_low_bounds, y_high_bounds):
- assert y_low <= y_high
-
- # Compute observed quantiles per X slice during training (only works for univariate dummy datasets):
- low_quantile_dict_train = get_discretized_quantile_dict(
- X_train.reshape(
- -1,
- ),
- y_train,
- confidence_level + ((1 - confidence_level) / 2),
- )
- high_quantile_dict_train = get_discretized_quantile_dict(
- X_train.reshape(
- -1,
- ),
- y_train,
- (1 - confidence_level) / 2,
- )
-
- # Check that predictions return observed quantiles during training
- # Prediction error deviations of more than this amount
- # will count as a breach:
- y_breach_threshold = 1
- # More than this percentage of breaches will fail the test:
- breach_tolerance = 0.3
- low_margin_breaches, high_margin_breaches = 0, 0
- for x_obs, y_low, y_high in zip(
- X_train.reshape(
- -1,
- ),
- y_low_bounds,
- y_high_bounds,
- ):
- if abs(y_low - low_quantile_dict_train[x_obs]) > y_breach_threshold:
- low_margin_breaches += 1
- if abs(y_high - high_quantile_dict_train[x_obs]) > y_breach_threshold:
- high_margin_breaches += 1
- assert low_margin_breaches < len(X_train) * breach_tolerance
- assert high_margin_breaches < len(X_train) * breach_tolerance
-
- # Check conformal interval coverage on validation data
- # (note validation data is actively used by the searcher
- # to calibrate its conformal intervals, so this is not an
- # OOS test, just a sanity check):
- interval_breach_states = []
- for y_obs, y_low, y_high in zip(y_val, y_low_bounds, y_high_bounds):
- is_interval_breach = 0 if y_high > y_obs > y_low else 1
- interval_breach_states.append(is_interval_breach)
-
- interval_breach_rate = sum(interval_breach_states) / len(interval_breach_states)
- breach_margin = 0.01
- assert (
- (confidence_level - breach_margin)
- <= (1 - interval_breach_rate)
- <= (confidence_level + breach_margin)
- )
diff --git a/tests/test_optimization.py b/tests/test_optimization.py
deleted file mode 100644
index 2914213..0000000
--- a/tests/test_optimization.py
+++ /dev/null
@@ -1,53 +0,0 @@
-import time
-
-import pytest
-
-from confopt.optimization import derive_optimal_tuning_count, RuntimeTracker
-
-
-def test_runtime_tracker__return_runtime():
- dummy_tracker = RuntimeTracker()
- sleep_time = 5
- time.sleep(sleep_time)
- time_elapsed = dummy_tracker.return_runtime()
- assert sleep_time - 1 < round(time_elapsed) < sleep_time + 1
-
-
-def test_runtime_tracker__pause_runtime():
- dummy_tracker = RuntimeTracker()
- dummy_tracker.pause_runtime()
- sleep_time = 5
- time.sleep(sleep_time)
- dummy_tracker.resume_runtime()
- time_elapsed = dummy_tracker.return_runtime()
- assert time_elapsed < 1
-
-
-@pytest.mark.parametrize("base_model_runtime", [1, 100])
-@pytest.mark.parametrize("search_model_runtime", [1, 100])
-@pytest.mark.parametrize("search_to_base_runtime_ratio", [0.5, 2])
-@pytest.mark.parametrize("search_retraining_freq", [1, 10])
-def test_derive_optimal_tuning_count(
- base_model_runtime,
- search_model_runtime,
- search_to_base_runtime_ratio,
- search_retraining_freq,
-):
- n_iterations = derive_optimal_tuning_count(
- baseline_model_runtime=base_model_runtime,
- search_model_runtime=search_model_runtime,
- search_to_baseline_runtime_ratio=search_to_base_runtime_ratio,
- search_model_retraining_freq=search_retraining_freq,
- )
- assert n_iterations >= 1
- assert isinstance(n_iterations, int)
-
-
-def test_derive_optimal_tuning_count__no_iterations():
- n_iterations = derive_optimal_tuning_count(
- baseline_model_runtime=1,
- search_model_runtime=1,
- search_to_baseline_runtime_ratio=1,
- search_model_retraining_freq=1,
- )
- assert n_iterations == 1
diff --git a/tests/test_tuning.py b/tests/test_tuning.py
index c7058cd..36b2edf 100644
--- a/tests/test_tuning.py
+++ b/tests/test_tuning.py
@@ -1,430 +1,353 @@
-import random
-from copy import deepcopy
-
-import numpy as np
-import pandas as pd
import pytest
+import numpy as np
+from typing import Dict
+from itertools import product
-from confopt.config import GBM_NAME
-from confopt.optimization import RuntimeTracker
-from confopt.tuning import (
- score_predictions,
- get_best_configuration_idx,
- process_and_split_estimation_data,
- normalize_estimation_data,
- update_adaptive_confidence_level,
-)
-
-DEFAULT_SEED = 1234
-
+from confopt.tuning import ConformalTuner, stop_search
+from confopt.wrapping import CategoricalRange, IntRange
+from confopt.utils.tracking import RuntimeTracker
+from confopt.selection.acquisition import QuantileConformalSearcher, LowerBoundSampler
-@pytest.mark.parametrize("optimization_direction", ["direct", "inverse"])
-def test_get_best_configuration_idx(optimization_direction):
- lower_bound = np.array([5, 4, 3, 2, 1])
- higher_bound = lower_bound + 1
- dummy_performance_bounds = (lower_bound, higher_bound)
- best_idx = get_best_configuration_idx(
- configuration_performance_bounds=dummy_performance_bounds,
- optimization_direction=optimization_direction,
+def test_stop_search_no_remaining_configurations():
+ assert stop_search(
+ n_remaining_configurations=0,
+ current_iter=5,
+ current_runtime=10.0,
+ max_runtime=100.0,
+ max_searches=50,
)
- assert best_idx >= 0
- if optimization_direction == "direct":
- assert best_idx == np.argmax(higher_bound)
- elif optimization_direction == "inverse":
- assert best_idx == np.argmin(lower_bound)
+@pytest.mark.parametrize("max_runtime", [10.0, 15.0, 20.0])
+def test_stop_search_runtime_exceeded(max_runtime):
+ current_runtime = 25.0
+ should_stop = current_runtime >= max_runtime
+ assert (
+ stop_search(
+ n_remaining_configurations=10,
+ current_iter=5,
+ current_runtime=current_runtime,
+ max_runtime=max_runtime,
+ max_searches=50,
+ )
+ == should_stop
+ )
-@pytest.mark.parametrize(
- "scoring_function", ["accuracy_score", "mean_squared_error", "log_loss"]
-)
-def test_score_predictions__perfect_score(scoring_function):
- dummy_y_obs = np.array([1, 0, 1, 0, 1, 1])
- dummy_y_pred = deepcopy(dummy_y_obs)
- score = score_predictions(
- y_obs=dummy_y_obs, y_pred=dummy_y_pred, scoring_function=scoring_function
+@pytest.mark.parametrize("max_searches", [10, 20, 30])
+def test_stop_search_iterations_exceeded(max_searches):
+ current_iter = 25
+ should_stop = current_iter >= max_searches
+ assert (
+ stop_search(
+ n_remaining_configurations=10,
+ current_iter=current_iter,
+ current_runtime=5.0,
+ max_runtime=100.0,
+ max_searches=max_searches,
+ )
+ == should_stop
)
- if scoring_function == "accuracy_score":
- assert score == 1
- elif scoring_function == "mean_squared_error":
- assert score == 0
- elif scoring_function == "log_loss":
- assert 0 < score < 0.001
-
-def test_process_and_split_estimation_data(dummy_configurations):
- train_split = 0.5
- dummy_searched_configurations = pd.DataFrame(dummy_configurations).to_numpy()
- stored_dummy_searched_configurations = deepcopy(dummy_searched_configurations)
- dummy_searched_performances = np.array(
- [random.random() for _ in range(len(dummy_configurations))]
- )
- stored_dummy_searched_performances = deepcopy(dummy_searched_performances)
-
- X_train, y_train, X_val, y_val = process_and_split_estimation_data(
- searched_configurations=dummy_searched_configurations,
- searched_performances=dummy_searched_performances,
- train_split=train_split,
- filter_outliers=False,
- outlier_scope=None,
- random_state=DEFAULT_SEED,
+def test_stop_search_continue_search():
+ assert not stop_search(
+ n_remaining_configurations=10,
+ current_iter=5,
+ current_runtime=10.0,
+ max_runtime=100.0,
+ max_searches=50,
)
- assert len(X_val) == len(y_val)
- assert len(X_train) == len(y_train)
- assert len(X_val) + len(X_train) == len(dummy_searched_configurations)
+def test_check_objective_function_wrong_argument_count(dummy_parameter_grid):
+ def invalid_objective(config1, config2):
+ return 1.0
- assert (
- abs(len(X_train) - round(len(dummy_searched_configurations) * train_split)) <= 1
- )
- assert (
- abs(len(X_val) - round(len(dummy_searched_configurations) * (1 - train_split)))
- <= 1
- )
+ with pytest.raises(
+ ValueError, match="Objective function must take exactly one argument"
+ ):
+ ConformalTuner(
+ objective_function=invalid_objective,
+ search_space=dummy_parameter_grid,
+ minimize=True,
+ )
- # Assert there is no mutability of input:
- assert np.array_equal(
- dummy_searched_configurations, stored_dummy_searched_configurations
- )
- assert np.array_equal(
- dummy_searched_performances, stored_dummy_searched_performances
- )
+def test_check_objective_function_wrong_argument_name(dummy_parameter_grid):
+ def invalid_objective(config):
+ return 1.0
-def test_process_and_split_estimation_data__reproducibility(dummy_configurations):
- train_split = 0.5
- dummy_searched_configurations = pd.DataFrame(dummy_configurations).to_numpy()
- dummy_searched_performances = np.array(
- [random.random() for _ in range(len(dummy_configurations))]
- )
+ with pytest.raises(
+ ValueError,
+ match="The objective function must take exactly one argument named 'configuration'",
+ ):
+ ConformalTuner(
+ objective_function=invalid_objective,
+ search_space=dummy_parameter_grid,
+ minimize=True,
+ )
- (
- X_train_first_call,
- y_train_first_call,
- X_val_first_call,
- y_val_first_call,
- ) = process_and_split_estimation_data(
- searched_configurations=dummy_searched_configurations,
- searched_performances=dummy_searched_performances,
- train_split=train_split,
- filter_outliers=False,
- outlier_scope=None,
- random_state=DEFAULT_SEED,
- )
- (
- X_train_second_call,
- y_train_second_call,
- X_val_second_call,
- y_val_second_call,
- ) = process_and_split_estimation_data(
- searched_configurations=dummy_searched_configurations,
- searched_performances=dummy_searched_performances,
- train_split=train_split,
- filter_outliers=False,
- outlier_scope=None,
- random_state=DEFAULT_SEED,
- )
- assert np.array_equal(X_train_first_call, X_train_second_call)
- assert np.array_equal(y_train_first_call, y_train_second_call)
- assert np.array_equal(X_val_first_call, X_val_second_call)
- assert np.array_equal(y_val_first_call, y_val_second_call)
+def test_evaluate_configuration(tuner):
+ config = {"param_1": 0.5, "param_2": 10, "param_3": "option1"}
+ performance, runtime = tuner._evaluate_configuration(config)
-def test_normalize_estimation_data(dummy_configurations):
- # Proportion of all candidate configurations that
- # have already been searched:
- searched_split = 0.5
- # Split of searched configurations that is used as
- # training data for the search estimator:
- train_split = 0.5
+ assert performance == 2
+ assert runtime >= 0
- dummy_searched_configurations = dummy_configurations[
- : round(len(dummy_configurations) * searched_split)
- ]
- dummy_searchable_configurations = pd.DataFrame(
- dummy_configurations[round(len(dummy_configurations) * searched_split) :]
- ).to_numpy()
- stored_dummy_searchable_configurations = deepcopy(dummy_searchable_configurations)
- dummy_training_searched_configurations = pd.DataFrame(
- dummy_searched_configurations[
- : round(len(dummy_searched_configurations) * train_split)
- ]
- ).to_numpy()
- stored_dummy_training_searched_configurations = deepcopy(
- dummy_training_searched_configurations
- )
- dummy_validation_searched_configurations = pd.DataFrame(
- dummy_searched_configurations[
- round(len(dummy_searched_configurations) * train_split) :
- ]
- ).to_numpy()
- stored_dummy_validation_searched_configurations = deepcopy(
- dummy_validation_searched_configurations
- )
- (
- normalized_training_searched_configurations,
- normalized_validation_searched_configurations,
- normalized_searchable_configurations,
- ) = normalize_estimation_data(
- training_searched_configurations=dummy_training_searched_configurations,
- validation_searched_configurations=dummy_validation_searched_configurations,
- searchable_configurations=dummy_searchable_configurations,
- )
+def test_random_search_with_warm_start(
+ mock_constant_objective_function, dummy_parameter_grid
+):
+ warm_start_configs = [
+ ({"param_1": 0.5, "param_2": 10, "param_3": "option1"}, 0.8),
+ ]
- assert len(normalized_training_searched_configurations) == len(
- dummy_training_searched_configurations
- )
- assert len(normalized_validation_searched_configurations) == len(
- normalized_validation_searched_configurations
- )
- assert len(normalized_searchable_configurations) == len(
- normalized_searchable_configurations
+ tuner = ConformalTuner(
+ objective_function=mock_constant_objective_function,
+ search_space=dummy_parameter_grid,
+ minimize=True,
+ warm_starts=warm_start_configs,
)
- # Assert there is no mutability of inputs:
- assert np.array_equal(
- dummy_training_searched_configurations,
- stored_dummy_training_searched_configurations,
- )
- assert np.array_equal(
- dummy_validation_searched_configurations,
- stored_dummy_validation_searched_configurations,
- )
- assert np.array_equal(
- dummy_searchable_configurations, stored_dummy_searchable_configurations
- )
+ tuner.initialize_tuning_resources()
+ tuner.search_timer = RuntimeTracker()
+ assert len(tuner.study.trials) == 1
+ assert tuner.study.trials[0].acquisition_source == "warm_start"
-@pytest.mark.parametrize("breach", [True, False])
-@pytest.mark.parametrize("true_confidence_level", [0.2, 0.8])
-@pytest.mark.parametrize("learning_rate", [0.01, 0.1])
-def test_update_adaptive_interval(breach, true_confidence_level, learning_rate):
- updated_confidence_level = update_adaptive_confidence_level(
- true_confidence_level=true_confidence_level,
- last_confidence_level=true_confidence_level,
- breach=breach,
- learning_rate=learning_rate,
+ tuner.random_search(
+ max_random_iter=3,
+ verbose=False,
)
- assert 0 < updated_confidence_level < 1
- if breach:
- assert updated_confidence_level >= true_confidence_level
- else:
- assert updated_confidence_level <= true_confidence_level
-
+ assert len(tuner.study.trials) == 4
+ assert tuner.study.trials[0].acquisition_source == "warm_start"
+ assert all(trial.acquisition_source == "rs" for trial in tuner.study.trials[1:])
-def test_get_tuning_configurations(dummy_initialized_conformal_searcher__gbm_mse):
- stored_search_space = dummy_initialized_conformal_searcher__gbm_mse.search_space
- tuning_configurations = (
- dummy_initialized_conformal_searcher__gbm_mse._get_tuning_configurations()
- )
+def test_random_search_with_nan_performance(dummy_parameter_grid):
+ def nan_objective(configuration: Dict) -> float:
+ return np.nan
- for configuration in tuning_configurations:
- for param_name, param_value in configuration.items():
- # Check configuration only has parameter names from parameter grid prompt:
- assert param_name in stored_search_space.keys()
- # Check values in configuration come from range in parameter grid prompt:
- assert param_value in stored_search_space[param_name]
- # Test for mutability:
- assert (
- stored_search_space
- == dummy_initialized_conformal_searcher__gbm_mse.search_space
+ tuner = ConformalTuner(
+ objective_function=nan_objective,
+ search_space=dummy_parameter_grid,
+ minimize=True,
)
+ tuner.initialize_tuning_resources()
+ tuner.search_timer = RuntimeTracker()
-def test_get_tuning_configurations__reproducibility(
- dummy_initialized_conformal_searcher__gbm_mse,
-):
- assert (
- dummy_initialized_conformal_searcher__gbm_mse._get_tuning_configurations()
- == dummy_initialized_conformal_searcher__gbm_mse._get_tuning_configurations()
+ tuner.random_search(
+ max_random_iter=3,
+ verbose=False,
)
+ # Should handle NaN gracefully and not crash
+ assert len(tuner.study.trials) == 0
-def test_evaluate_configuration_performance(
- dummy_initialized_conformal_searcher__gbm_mse, dummy_gbm_configurations
-):
- # Arbitrarily select the first configuration in the list:
- dummy_configuration = dummy_gbm_configurations[0]
- stored_dummy_configuration = deepcopy(dummy_configuration)
- performance = dummy_initialized_conformal_searcher__gbm_mse._evaluate_configuration_performance(
- configuration=dummy_configuration, random_state=DEFAULT_SEED
- )
+@pytest.mark.parametrize("random_state", [42, 123, 999])
+def test_tune_method_reproducibility(dummy_parameter_grid, random_state):
+ """Test that tune method produces identical results with same random seed"""
- assert performance > 0
- # Test for mutability:
- assert stored_dummy_configuration == dummy_configuration
+ def complex_objective(configuration: Dict) -> float:
+ # Complex objective with multiple terms
+ x1 = configuration["param_1"]
+ x2 = configuration["param_2"]
+ x3_val = {"option1": 1, "option2": 2, "option3": 3}[configuration["param_3"]]
+ return x1**2 + np.sin(x2) + x3_val * 0.5
+ def run_tune_session():
+ # Create fresh searcher for each run to avoid state contamination
+ searcher = QuantileConformalSearcher(
+ quantile_estimator_architecture="ql",
+ sampler=LowerBoundSampler(
+ interval_width=0.1,
+ adapter="DtACI",
+ beta_decay="logarithmic_decay",
+ c=1,
+ ),
+ n_pre_conformal_trials=5,
+ )
-def test_evaluate_configuration_performance__reproducibility(
- dummy_initialized_conformal_searcher__gbm_mse, dummy_gbm_configurations
-):
- # Arbitrarily select the first configuration in the list:
- dummy_configuration = dummy_gbm_configurations[0]
+ tuner = ConformalTuner(
+ objective_function=complex_objective,
+ search_space=dummy_parameter_grid,
+ minimize=True,
+ n_candidates=200,
+ )
- assert dummy_initialized_conformal_searcher__gbm_mse._evaluate_configuration_performance(
- configuration=dummy_configuration, random_state=DEFAULT_SEED
- ) == dummy_initialized_conformal_searcher__gbm_mse._evaluate_configuration_performance(
- configuration=dummy_configuration, random_state=DEFAULT_SEED
- )
+ tuner.tune(
+ n_random_searches=10,
+ searcher=searcher,
+ optimizer_framework=None,
+ random_state=random_state,
+ max_searches=25,
+ max_runtime=None,
+ verbose=False,
+ )
+ return tuner.study
-def test_random_search(dummy_initialized_conformal_searcher__gbm_mse):
- n_searches = 5
- max_runtime = 30
- dummy_initialized_conformal_searcher__gbm_mse.search_timer = RuntimeTracker()
-
- (
- searched_configurations,
- searched_performances,
- searched_timestamps,
- runtime_per_search,
- ) = dummy_initialized_conformal_searcher__gbm_mse._random_search(
- n_searches=n_searches,
- max_runtime=max_runtime,
- random_state=DEFAULT_SEED,
- )
+ # Run twice with same seed
+ study1 = run_tune_session()
+ study2 = run_tune_session()
- for performance in searched_performances:
- assert performance > 0
- assert len(searched_configurations) > 0
- assert len(searched_performances) > 0
- assert len(searched_timestamps) > 0
- assert (
- len(searched_configurations)
- == len(searched_performances)
- == len(searched_timestamps)
- )
- assert len(searched_configurations) == n_searches
- assert 0 < runtime_per_search < max_runtime
+ # Verify identical results
+ assert len(study1.trials) == len(study2.trials)
+ for trial1, trial2 in zip(study1.trials, study2.trials):
+ assert trial1.configuration == trial2.configuration
+ assert trial1.performance == trial2.performance
+ # Skip acquisition_source comparison as it contains object addresses
-def test_random_search__reproducibility(
- dummy_initialized_conformal_searcher__gbm_mse,
-):
- n_searches = 5
- max_runtime = 30
- dummy_initialized_conformal_searcher__gbm_mse.search_timer = RuntimeTracker()
-
- (
- searched_configurations_first_call,
- searched_performances_first_call,
- _,
- _,
- ) = dummy_initialized_conformal_searcher__gbm_mse._random_search(
- n_searches=n_searches,
- max_runtime=max_runtime,
- random_state=DEFAULT_SEED,
- )
- (
- searched_configurations_second_call,
- searched_performances_second_call,
- _,
- _,
- ) = dummy_initialized_conformal_searcher__gbm_mse._random_search(
- n_searches=n_searches,
- max_runtime=max_runtime,
- random_state=DEFAULT_SEED,
- )
- assert searched_configurations_first_call == searched_configurations_second_call
- assert searched_performances_first_call == searched_performances_second_call
-
-
-def test_search(dummy_initialized_conformal_searcher__gbm_mse):
- # TODO: Below I hard coded a slice of possible inputs, but consider
- # pytest parametrizing these (though test will be very heavy,
- # so tag as slow and only run when necessary)
- confidence_level = 0.2
- conformal_model_type = GBM_NAME
- conformal_retraining_frequency = 1
- conformal_learning_rate = 0.01
- enable_adaptive_intervals = True
- max_runtime = 120
- min_training_iterations = 20
-
- stored_search_space = dummy_initialized_conformal_searcher__gbm_mse.search_space
- stored_tuning_configurations = (
- dummy_initialized_conformal_searcher__gbm_mse.tuning_configurations
+@pytest.mark.slow
+@pytest.mark.parametrize("dynamic_sampling", [True, False])
+def test_tune_method_comprehensive_integration(
+ comprehensive_minimizing_tuning_setup, dynamic_sampling
+):
+ """Comprehensive integration test for tune method (single run, logic only)"""
+ tuner, searcher, warm_start_configs, _ = comprehensive_minimizing_tuning_setup(
+ dynamic_sampling
)
- dummy_initialized_conformal_searcher__gbm_mse.search(
- conformal_search_estimator=conformal_model_type,
- confidence_level=confidence_level,
- n_random_searches=min_training_iterations,
- runtime_budget=max_runtime,
- conformal_retraining_frequency=conformal_retraining_frequency,
- conformal_learning_rate=conformal_learning_rate,
- enable_adaptive_intervals=enable_adaptive_intervals,
- verbose=0,
+ tuner.tune(
+ n_random_searches=15,
+ searcher=searcher,
+ optimizer_framework=None,
+ random_state=42,
+ max_searches=50,
+ max_runtime=5 * 60,
+ verbose=False,
)
+ study = tuner.study
- assert (
- len(dummy_initialized_conformal_searcher__gbm_mse.searched_configurations) > 0
- )
- assert len(dummy_initialized_conformal_searcher__gbm_mse.searched_performances) > 0
- assert len(
- dummy_initialized_conformal_searcher__gbm_mse.searched_configurations
- ) == len(dummy_initialized_conformal_searcher__gbm_mse.searched_performances)
- # Test for mutability:
- assert (
- stored_search_space
- == dummy_initialized_conformal_searcher__gbm_mse.search_space
- )
- assert (
- stored_tuning_configurations
- == dummy_initialized_conformal_searcher__gbm_mse.tuning_configurations
- )
+ # Test 1: Verify correct number of trials
+ assert len(study.trials) == 50
+ # Test 2: Verify warm starts are present
+ warm_start_trials = [
+ t for t in study.trials if t.acquisition_source == "warm_start"
+ ]
+ assert len(warm_start_trials) == 3
+ warm_start_performances = [t.performance for t in warm_start_trials]
+ expected_performances = [perf for _, perf in warm_start_configs]
+ assert set(warm_start_performances) == set(expected_performances)
+
+ # Test 3: Verify trial sources
+ rs_trials = [t for t in study.trials if t.acquisition_source == "rs"]
+ conformal_trials = [
+ t for t in study.trials if t.acquisition_source not in ["warm_start", "rs"]
+ ]
+ assert len(rs_trials) == 12
+ assert len(conformal_trials) == 35
+
+ # Test 4: Verify configurations are diverse
+ all_configs = [t.configuration for t in study.trials]
+ unique_configs = set(str(config) for config in all_configs)
+ assert len(unique_configs) == len(all_configs)
+
+ # Test 5: Verify study methods work correctly
+ best_config = study.get_best_configuration()
+ best_value = study.get_best_performance()
+ assert best_config in all_configs
+ assert best_value == min(t.performance for t in study.trials)
+
+
+@pytest.mark.slow
+@pytest.mark.parametrize("minimize", [True, False])
+@pytest.mark.parametrize("dynamic_sampling", [True, False])
+def test_conformal_vs_random_performance_averaged(
+ comprehensive_minimizing_tuning_setup,
+ comprehensive_maximizing_tuning_setup,
+ minimize,
+ dynamic_sampling,
+):
+ """Compare conformal vs random search win rate over multiple runs."""
+ n_repeats = 20
+ conformal_wins, total_comparisons = 0, 0
-def test_search__reproducibility(dummy_initialized_conformal_searcher__gbm_mse):
- confidence_level = 0.2
- conformal_model_type = GBM_NAME
- conformal_retraining_frequency = 1
- conformal_learning_rate = 0.01
- enable_adaptive_intervals = True
- max_runtime = 120
- min_training_iterations = 20
-
- searcher_first_call = deepcopy(dummy_initialized_conformal_searcher__gbm_mse)
- searcher_second_call = deepcopy(dummy_initialized_conformal_searcher__gbm_mse)
-
- searcher_first_call.search(
- conformal_search_estimator=conformal_model_type,
- confidence_level=confidence_level,
- n_random_searches=min_training_iterations,
- runtime_budget=max_runtime,
- conformal_retraining_frequency=conformal_retraining_frequency,
- conformal_learning_rate=conformal_learning_rate,
- enable_adaptive_intervals=enable_adaptive_intervals,
- verbose=0,
- random_state=DEFAULT_SEED,
- )
- searcher_second_call.search(
- conformal_search_estimator=conformal_model_type,
- confidence_level=confidence_level,
- n_random_searches=min_training_iterations,
- runtime_budget=max_runtime,
- conformal_retraining_frequency=conformal_retraining_frequency,
- conformal_learning_rate=conformal_learning_rate,
- enable_adaptive_intervals=enable_adaptive_intervals,
- verbose=0,
- random_state=DEFAULT_SEED,
- )
+ if minimize:
+ tuning_setup = comprehensive_minimizing_tuning_setup
+ else:
+ tuning_setup = comprehensive_maximizing_tuning_setup
+
+ for seed in range(n_repeats):
+ # Run conformal tuner (15 random + 35 conformal searches)
+ conformal_tuner, searcher, _, _ = tuning_setup(dynamic_sampling)
+ conformal_tuner.tune(
+ n_random_searches=10,
+ searcher=searcher,
+ optimizer_framework=None,
+ random_state=seed,
+ max_searches=40,
+ max_runtime=5 * 60,
+ verbose=False,
+ )
+ conformal_best = conformal_tuner.get_best_value()
+
+ # Run pure random search tuner (40 random searches, no conformal)
+ random_tuner, searcher, _, _ = tuning_setup(dynamic_sampling)
+ random_tuner.tune(
+ n_random_searches=40,
+ searcher=searcher,
+ optimizer_framework=None,
+ random_state=seed,
+ max_searches=40, # This ensures only 40 random searches, no conformal
+ max_runtime=5 * 60,
+ verbose=False,
+ )
+ random_best = random_tuner.get_best_value()
+
+ if minimize:
+ conformal_wins_round = conformal_best < random_best
+ else:
+ conformal_wins_round = conformal_best > random_best
+
+ if conformal_wins_round:
+ conformal_wins += 1
+ total_comparisons += 1
+
+ assert conformal_wins / total_comparisons >= 0.8
+
+
+@pytest.mark.parametrize("minimize", [True, False])
+def test_best_fetcher_methods(minimize):
+ grid = {
+ "x": CategoricalRange(choices=[0, 1]),
+ "y": IntRange(min_value=0, max_value=2),
+ }
+
+ def objective(configuration):
+ return configuration["x"] + configuration["y"] * 10
+
+ tuner = ConformalTuner(
+ objective_function=objective,
+ search_space=grid,
+ minimize=minimize,
+ n_candidates=100,
+ )
+ tuner.initialize_tuning_resources()
+ tuner.search_timer = RuntimeTracker()
+
+ total_configs = len(list(product([0, 1], [0, 1, 2])))
+ tuner.random_search(max_random_iter=total_configs, verbose=False)
+
+ # Use built-in methods to get best config and value
+ best_config = tuner.get_best_params()
+ best_value = tuner.get_best_value()
+
+ if minimize:
+ expected_config = {"x": 0, "y": 0}
+ else:
+ expected_config = {"x": 1, "y": 2}
+ expected_value = objective(expected_config)
- assert (
- searcher_first_call.searched_configurations
- == searcher_second_call.searched_configurations
- )
- assert (
- searcher_first_call.searched_performances
- == searcher_second_call.searched_performances
- )
+ assert best_config == expected_config
+ assert best_value == expected_value
diff --git a/tests/test_utils.py b/tests/test_utils.py
deleted file mode 100644
index f9fc341..0000000
--- a/tests/test_utils.py
+++ /dev/null
@@ -1,81 +0,0 @@
-from confopt.utils import (
- get_tuning_configurations,
- get_perceptron_layers,
-)
-
-DEFAULT_SEED = 1234
-
-
-def test_get_perceptron_layers():
- dummy_n_layers_grid = [2, 3, 4]
- dummy_layer_size_grid = [16, 32, 64, 128]
-
- layer_list = get_perceptron_layers(
- n_layers_grid=dummy_n_layers_grid,
- layer_size_grid=dummy_layer_size_grid,
- random_seed=DEFAULT_SEED,
- )
-
- for layer in layer_list:
- assert isinstance(layer, tuple)
- assert min(dummy_n_layers_grid) <= len(layer) <= max(dummy_n_layers_grid)
- for layer_size in layer:
- assert (
- min(dummy_layer_size_grid) <= layer_size <= max(dummy_layer_size_grid)
- )
-
-
-def test_get_perceptron_layers__reproducibility():
- dummy_n_layers_grid = [2, 3, 4]
- dummy_layer_size_grid = [16, 32, 64, 128]
-
- layer_list_first_call = get_perceptron_layers(
- n_layers_grid=dummy_n_layers_grid,
- layer_size_grid=dummy_layer_size_grid,
- random_seed=DEFAULT_SEED,
- )
- layer_list_second_call = get_perceptron_layers(
- n_layers_grid=dummy_n_layers_grid,
- layer_size_grid=dummy_layer_size_grid,
- random_seed=DEFAULT_SEED,
- )
- for layer_first_call, layer_second_call in zip(
- layer_list_first_call, layer_list_second_call
- ):
- assert layer_first_call == layer_second_call
-
-
-def test_get_tuning_configurations(dummy_parameter_grid):
- dummy_n_configurations = 10
-
- tuning_configurations = get_tuning_configurations(
- parameter_grid=dummy_parameter_grid,
- n_configurations=dummy_n_configurations,
- random_state=DEFAULT_SEED,
- )
- assert len(tuning_configurations) < dummy_n_configurations
- for configuration in tuning_configurations:
- for k, v in configuration.items():
- # Check configuration only has parameter names from parameter grid prompt:
- assert k in dummy_parameter_grid.keys()
- # Check values in configuration come from range in parameter grid prompt:
- assert v in dummy_parameter_grid[k]
-
-
-def test_get_tuning_configurations__reproducibility(dummy_parameter_grid):
- dummy_n_configurations = 10
-
- tuning_configurations_first_call = get_tuning_configurations(
- parameter_grid=dummy_parameter_grid,
- n_configurations=dummy_n_configurations,
- random_state=DEFAULT_SEED,
- )
- tuning_configurations_second_call = get_tuning_configurations(
- parameter_grid=dummy_parameter_grid,
- n_configurations=dummy_n_configurations,
- random_state=DEFAULT_SEED,
- )
- for configuration_first_call, configuration_second_call in zip(
- tuning_configurations_first_call, tuning_configurations_second_call
- ):
- assert configuration_first_call == configuration_second_call
diff --git a/tests/utils/configurations/test_encoding.py b/tests/utils/configurations/test_encoding.py
new file mode 100644
index 0000000..2664149
--- /dev/null
+++ b/tests/utils/configurations/test_encoding.py
@@ -0,0 +1,66 @@
+from confopt.utils.configurations.encoding import ConfigurationEncoder
+from confopt.wrapping import IntRange, FloatRange, CategoricalRange
+
+
+def test_configuration_encoder():
+ """Test that ConfigurationEncoder properly encodes configurations"""
+ # Create configurations with mixed parameter types
+ configs = [
+ {"numeric1": 1.0, "numeric2": 5, "cat1": "a", "cat2": True},
+ {"numeric1": 2.0, "numeric2": 10, "cat1": "b", "cat2": False},
+ {"numeric1": 3.0, "numeric2": 15, "cat1": "a", "cat2": True},
+ ]
+
+ # Define search space with categorical parameters
+ search_space = {
+ "numeric1": FloatRange(min_value=0.0, max_value=10.0),
+ "numeric2": IntRange(min_value=0, max_value=20),
+ "cat1": CategoricalRange(choices=["a", "b", "c"]),
+ "cat2": CategoricalRange(choices=[True, False]),
+ }
+
+ # Test initialization
+ encoder = ConfigurationEncoder(search_space)
+
+ # Verify categorical mappings are created correctly
+ assert "cat1" in encoder.categorical_mappings
+ assert "cat2" in encoder.categorical_mappings
+
+ # Test transformation
+ df = encoder.transform(configs)
+
+ # Check shape - should have columns for numeric1, numeric2, cat1_a, cat1_b, cat1_c, cat2_False, cat2_True
+ assert df.shape[0] == 3 # 3 rows
+
+ # Verify numeric columns are preserved
+ assert "numeric1" in df.columns
+ assert "numeric2" in df.columns
+
+ # Check one-hot encoding worked correctly for string categorical values
+ cat1_cols = [col for col in df.columns if col.startswith("cat1_")]
+ assert (
+ len(cat1_cols) == 3
+ ) # "a", "b", and "c" (all possible values from search space)
+
+ cat1_a_col = next(col for col in cat1_cols if "a" in col)
+ cat1_b_col = next(col for col in cat1_cols if "b" in col)
+
+ # First row has cat1="a", so a=1, b=0
+ assert df.loc[0, cat1_a_col] == 1
+ assert df.loc[0, cat1_b_col] == 0
+
+ # Second row has cat1="b", so a=0, b=1
+ assert df.loc[1, cat1_a_col] == 0
+ assert df.loc[1, cat1_b_col] == 1
+
+ # Check boolean categorical values
+ cat2_cols = [col for col in df.columns if col.startswith("cat2_")]
+ assert len(cat2_cols) == 2 # True and False
+
+ # Boolean values are encoded with their string representation: 'cat2_True' and 'cat2_False'
+ cat2_true_col = "cat2_True"
+ cat2_false_col = "cat2_False"
+
+ # First row has cat2=True, so True=1, False=0
+ assert df.loc[0, cat2_true_col] == 1
+ assert df.loc[0, cat2_false_col] == 0
diff --git a/tests/utils/configurations/test_sampling_configurations.py b/tests/utils/configurations/test_sampling_configurations.py
new file mode 100644
index 0000000..1904a92
--- /dev/null
+++ b/tests/utils/configurations/test_sampling_configurations.py
@@ -0,0 +1,60 @@
+import pytest
+
+
+from confopt.utils.configurations.sampling import get_tuning_configurations
+
+RANDOM_STATE = 1234
+
+
+@pytest.mark.parametrize("method", ["uniform", "sobol"])
+def test_reproducibility(dummy_parameter_grid, method):
+ configs1 = get_tuning_configurations(
+ parameter_grid=dummy_parameter_grid,
+ n_configurations=10,
+ random_state=RANDOM_STATE,
+ sampling_method=method,
+ )
+ configs2 = get_tuning_configurations(
+ parameter_grid=dummy_parameter_grid,
+ n_configurations=10,
+ random_state=RANDOM_STATE,
+ sampling_method=method,
+ )
+ assert configs1 == configs2
+
+
+@pytest.mark.parametrize("method", ["uniform", "sobol"])
+def test_config_value_ranges(dummy_parameter_grid, method):
+ n = 50
+ configs = get_tuning_configurations(
+ parameter_grid=dummy_parameter_grid,
+ n_configurations=n,
+ random_state=RANDOM_STATE,
+ sampling_method=method,
+ )
+ assert len(configs) == n
+
+ for config in configs:
+ int_val = config["param_2"]
+ assert isinstance(int_val, int)
+ assert 1 <= int_val <= 100
+
+ float_val = config["param_1"]
+ assert isinstance(float_val, float)
+ assert 0.01 <= float_val <= 100
+
+ cat_val = config["param_3"]
+ assert cat_val in dummy_parameter_grid["param_3"].choices
+
+
+@pytest.mark.parametrize("method", ["uniform", "sobol"])
+def test_sampling_uniqueness(dummy_parameter_grid, method):
+ n = 100
+ configs = get_tuning_configurations(
+ parameter_grid=dummy_parameter_grid,
+ n_configurations=n,
+ random_state=123,
+ sampling_method=method,
+ )
+ unique_configs = {frozenset(cfg.items()) for cfg in configs}
+ assert len(unique_configs) == len(configs)
diff --git a/tests/utils/test_optimization.py b/tests/utils/test_optimization.py
new file mode 100644
index 0000000..20c17db
--- /dev/null
+++ b/tests/utils/test_optimization.py
@@ -0,0 +1,137 @@
+import pytest
+from confopt.utils.optimization import FixedSearcherOptimizer, DecayingSearcherOptimizer
+
+
+@pytest.fixture
+def fixed_surrogate_tuner():
+ """Fixture to create a FixedSurrogateTuner instance."""
+ return FixedSearcherOptimizer(n_tuning_episodes=8, tuning_interval=6)
+
+
+def test_fixed_surrogate_tuner_initialization():
+ """Test initialization of FixedSurrogateTuner."""
+ tuner = FixedSearcherOptimizer(tuning_interval=7)
+ assert tuner.fixed_interval == 7
+
+
+def test_fixed_surrogate_tuner_select_arm(fixed_surrogate_tuner):
+ """Test that select_arm returns the fixed values."""
+ arm = fixed_surrogate_tuner.select_arm()
+ assert arm == (8, 6)
+
+
+def test_fixed_surrogate_tuner_update(fixed_surrogate_tuner):
+ """Test that update method doesn't change behavior."""
+ fixed_surrogate_tuner.update(
+ search_iter=10,
+ )
+
+ arm = fixed_surrogate_tuner.select_arm()
+ assert arm == (8, 6)
+
+
+@pytest.fixture
+def decaying_tuner():
+ """Fixture to create a DecayingSearcherOptimizer instance."""
+ return DecayingSearcherOptimizer(
+ n_tuning_episodes=10,
+ initial_tuning_interval=2,
+ decay_rate=0.5,
+ decay_type="linear",
+ max_tuning_interval=20,
+ )
+
+
+def test_decaying_tuner_initialization():
+ """Test that the DecayingSearcherOptimizer initializes correctly."""
+ tuner = DecayingSearcherOptimizer(
+ initial_tuning_interval=3,
+ )
+ assert tuner.initial_tuning_interval == 3
+
+ tuner = DecayingSearcherOptimizer(
+ initial_tuning_interval=4,
+ )
+ assert tuner.initial_tuning_interval == 4
+
+
+def test_decaying_tuner_invalid_decay_type():
+ """Test that invalid decay_type raises ValueError."""
+ with pytest.raises(ValueError, match="decay_type must be one of"):
+ DecayingSearcherOptimizer(decay_type="invalid")
+
+
+def test_decaying_tuner_linear_decay(decaying_tuner):
+ """Test linear decay calculation."""
+ # At iteration 0
+ decaying_tuner.update(search_iter=0)
+ arm = decaying_tuner.select_arm()
+ assert arm[0] == 10 # n_tuning_episodes should remain constant
+ assert arm[1] == 2 # initial_tuning_interval
+
+ # At iteration 2: interval = 2 + 0.5 * 2 = 3, rounded to 3
+ decaying_tuner.update(search_iter=2)
+ arm = decaying_tuner.select_arm()
+ assert arm[0] == 10
+ assert arm[1] == 3
+
+ # At iteration 10: interval = 2 + 0.5 * 10 = 7, rounded to 7
+ decaying_tuner.update(search_iter=10)
+ arm = decaying_tuner.select_arm()
+ assert arm[0] == 10
+ assert arm[1] == 7
+
+
+def test_decaying_tuner_exponential_decay():
+ """Test exponential decay calculation."""
+ tuner = DecayingSearcherOptimizer(
+ n_tuning_episodes=5,
+ initial_tuning_interval=2,
+ decay_rate=0.1,
+ decay_type="exponential",
+ max_tuning_interval=20,
+ )
+
+ # At iteration 0
+ tuner.update(search_iter=0)
+ arm = tuner.select_arm()
+ assert arm[0] == 5
+ assert arm[1] == 2 # initial_tuning_interval
+
+ # At iteration 5: interval = 2 * (1.1)^5 ≈ 3.22, rounded to 3
+ tuner.update(search_iter=5)
+ arm = tuner.select_arm()
+ assert arm[0] == 5
+ assert arm[1] == 3
+
+
+def test_decaying_tuner_logarithmic_decay():
+ """Test logarithmic decay calculation."""
+ tuner = DecayingSearcherOptimizer(
+ n_tuning_episodes=8,
+ initial_tuning_interval=2,
+ decay_rate=2.0,
+ decay_type="logarithmic",
+ max_tuning_interval=20,
+ )
+
+ # At iteration 0
+ tuner.update(search_iter=0)
+ arm = tuner.select_arm()
+ assert arm[0] == 8
+ assert arm[1] == 2 # initial_tuning_interval
+
+ # At iteration 4: interval = 2 + 2.0 * log(5) ≈ 5.22, rounded to 5
+ tuner.update(search_iter=4)
+ arm = tuner.select_arm()
+ assert arm[0] == 8
+ assert arm[1] == 5
+
+
+def test_decaying_tuner_max_interval_cap(decaying_tuner):
+ """Test that tuning interval is capped at max_tuning_interval."""
+ # Set a very high iteration to exceed max_tuning_interval
+ decaying_tuner.update(search_iter=100)
+ arm = decaying_tuner.select_arm()
+ assert arm[0] == 10
+ assert arm[1] == 20 # Should be capped at max_tuning_interval
diff --git a/tests/test_preprocessing.py b/tests/utils/test_preprocessing.py
similarity index 97%
rename from tests/test_preprocessing.py
rename to tests/utils/test_preprocessing.py
index cd4dd65..e780d4f 100644
--- a/tests/test_preprocessing.py
+++ b/tests/utils/test_preprocessing.py
@@ -1,7 +1,7 @@
import numpy as np
import pytest
-from confopt.preprocessing import train_val_split
+from confopt.utils.preprocessing import train_val_split
DEFAULT_SEED = 1234
diff --git a/tests/utils/test_tracking.py b/tests/utils/test_tracking.py
new file mode 100644
index 0000000..0aa4753
--- /dev/null
+++ b/tests/utils/test_tracking.py
@@ -0,0 +1,22 @@
+import time
+
+
+from confopt.utils.tracking import RuntimeTracker
+
+
+def test_runtime_tracker__return_runtime():
+ dummy_tracker = RuntimeTracker()
+ sleep_time = 2
+ time.sleep(sleep_time)
+ time_elapsed = dummy_tracker.return_runtime()
+ assert sleep_time - 1 < round(time_elapsed) < sleep_time + 1
+
+
+def test_runtime_tracker__pause_runtime():
+ dummy_tracker = RuntimeTracker()
+ dummy_tracker.pause_runtime()
+ sleep_time = 2
+ time.sleep(sleep_time)
+ dummy_tracker.resume_runtime()
+ time_elapsed = dummy_tracker.return_runtime()
+ assert time_elapsed < 1
 +
+ +
+ +
+