With the development of deep learning, a large number of works use brain MRI images for diagnostic prediction. However, many novices do not know much about MRI. Most natural image vision researchers often directly train the original MRI, and often cannot get good training results and explanations, which will waste a lot of time. Therefore, this work will describe how to preprocess the mri, and then directly align it for training. It has been compiled by UKB and ADNI Database.
The following work is to explain the linux platform, windows and mac can also be referred to, just need to pay less attention There are many preprocessing tools, such as Freesurfer, FSL, ANTs, SPM and so on. Here I will explain the combination of FSL and ANTs.
- install FSL from https://fsl.fmrib.ox.ac.uk/fsl/fslwiki/FslInstallation
- It is recommended to use fsinstaller.py for installation
- Offline installation can also be used, but pay attention to using miniconda(or conda) to create a virtual environment named fslpython, and modify the FSL address in bashrc.
- install ANTs from https://github.com/ANTsX/ANTs/wiki/Compiling-ANTs-on-Linux-and-Mac-OS
- follow "Installing developer tools"
- install nipype, nibabel
- Orientation correction
Different shooting machines, doctors, patients and other factors lead to different directions of images. This step rotates all images to the same orientation. (The direction here refers to the head facing)
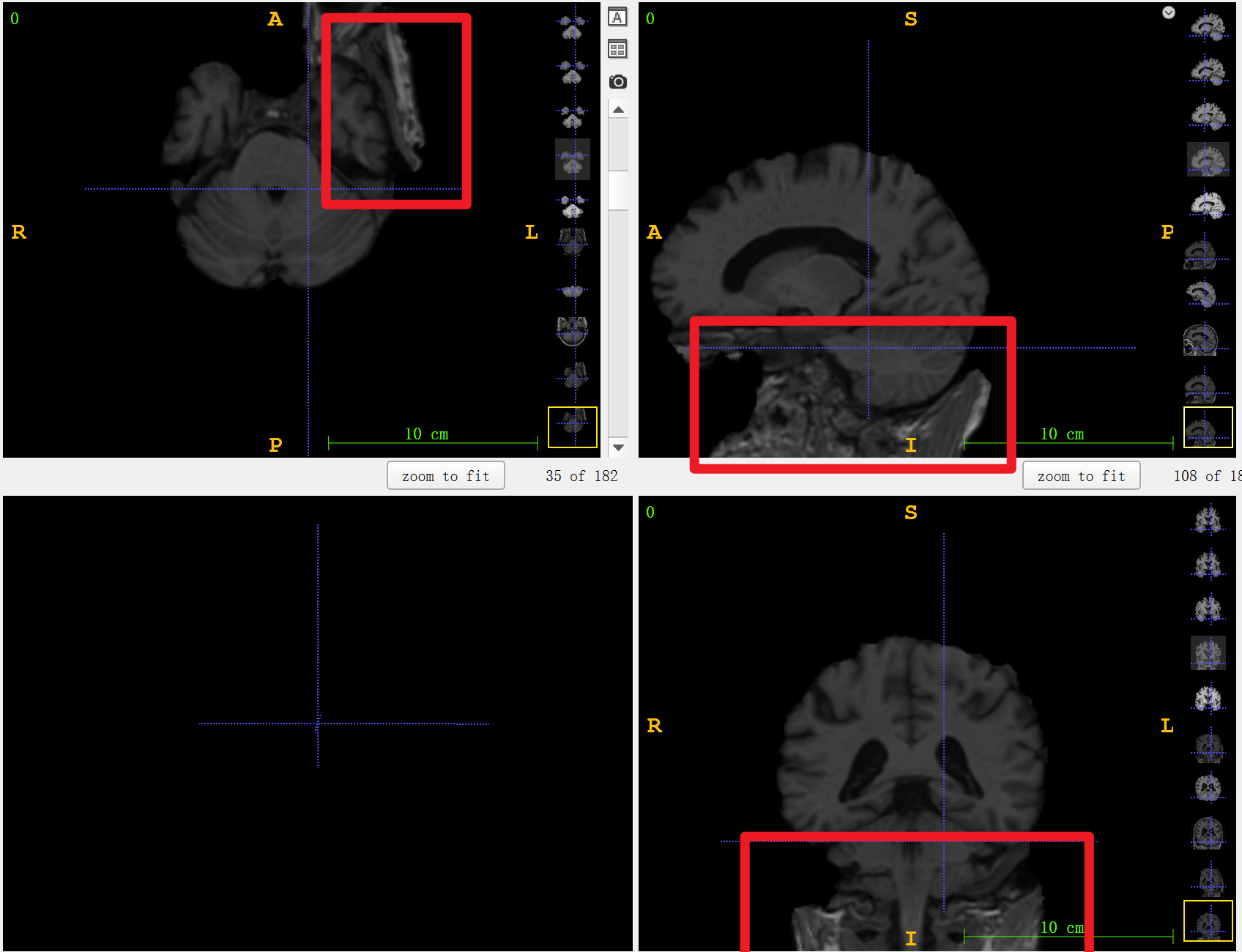
- Crop the image (remove the neck)
Part of the neck area is automatically cut. This step is for better stripping of the skull. This step is not necessary but recommended, since the direct removal of the skull will leave most of the neck area.
- Stripping the skull
Remove useless information about the skull. Laying the basis for white matter and gray matter segmentation
- Registration
Register each image to the standard brain space to facilitate subsequent roi feature extraction, neural network training, etc.
- bias correction
Correction for bias field effects
- Crop the image (remove the neck)** is not necessary but recommended. **TThe picture below will show the schematic diagram of directly stripping the skull without cutting. Please select according to the specific image situation
- The scalping and registration order can be reversed. If registering first, please use the MNI152_T1_1mm.nii.gz template for registration, if you peel off the scalp first and then register, please use MNI152_T1_1mm_brain.nii.gz. However, it is more recommended to peel off the scalp first and then perform registration.
This work uses a multi-threaded batch processing mechanism, you can choose to perform all processes on each image or perform each process on all images in turn. Either one will take a long time, it is recommended to use nohup to observe the running status.
python all_in_one/Process_all.py
python one_by_one/orient.py
python one_by_one/remove_neck.py
python one_by_one/skull.py
python one_by_one/registration.py
python one_by_one/bias_correction.py
https://github.com/junyuchen245/TransMorph_Transformer_for_Medical_Image_Registration/blob/main/PreprocessingMRI.md https://github.com/quqixun/BrainPrep