This repository provides a Retrieval-Augmented Generation (RAG) system designed to answer questions based on the content of a collection of PDF documents. The system processes, stores, and retrieves information from PDFs to generate context-aware responses to user queries.
The project is structured around three main stages: data ingestion, retrieval, and generation.
The ingestion pipeline processes PDF files and prepares them for retrieval.
- Document Processing: PDF files placed in the
textbooksdirectory are processed using the Docling library. This step handles document conversion and employs a hybrid chunking strategy to break the content into smaller text segments. - Embedding Generation: Each text chunk is converted into a vector embedding using OpenAI's text-embedding-3-small model.
- Storage: The processed data is stored in a PostgreSQL database with the PGVector extension for vector db capabilities. The ingestion logic is managed by scripts/ingest_pdf.py. It checks the file hash of each document to avoid re-ingesting content that has not changed.
The retrieval stage finds the most relevant document chunks based on a user's query.
- Query Expansion: The initial user query is expanded into multiple variations using Gemini 2.5 Flash to improve the breadth of the search.
- Hybrid Search: A combination of semantic search (using pgvector for cosine similarity on embeddings) and keyword-based search is used to fetch an initial set of relevant chunks.
- Reranking: The retrieved chunks are reranked using the BAAI/bge-reranker-v2-m3 model. This step scores the chunks against the original query and selects the top 5 most relevant results to use as context.
The final answer is generated by a large language model.
- LLM Inference: The top-ranked chunks and the original query are passed to a language model (tested with Google's Gemini 2.5 Flash) via the OpenRouter.ai API to synthesize a final answer.
The system provides both command-line and web-based interfaces for querying the RAG system:
Run queries directly from the terminal using:
python src/scripts/query_rag.py
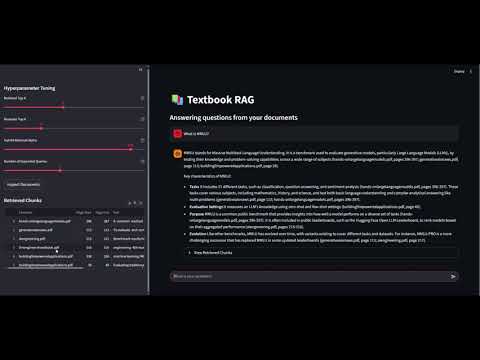
For a more interactive experience, the system includes a Streamlit web application that provides:
- Interactive Chat Interface: A conversational UI where users can ask questions and receive answers with proper formatting and markdown support.
- Real-time Hyperparameter Control: Adjustable sliders in the sidebar allow users to fine-tune retrieval parameters on-the-fly:
- Retrieval Top K: Control how many document chunks to retrieve via semantic and keyword search.
- Reranker Top K: Limit the number of chunks added to context via reranker model sorting and truncating.
- Hybrid Retriever Alpha: Adjust the alpha used to control the weighting for scores from semantic and keyword searches.
- Number of Expanded Queries: Number of additional queries to construct for increase retrieval quality. Total number of retrieved chunks will be the value of retrieval top k multiplied by the number of expanded queries. However, only the reranker top k number of chunks will be added to the context window.
- Source Transparency: The interface displays the relevant document chunks used to generate each answer, showing:
- The source document name and page numbers
- The actual text content that informed the response
- Similarity scores for each chunk
- Session Management: Maintains conversation history within the session for better context.
Launch the Streamlit interface with:
streamlit run src/scripts/streamlit_query_rag.py
The web interface makes it easy to experiment with different parameter settings and understand how the RAG system arrives at its answers by showing the underlying source material.
The system requires a PostgreSQL database with the pgvector extension enabled. The schema consists of two primary tables, which are created by the scripts/init_sql_tables.py script.
- documents table: Stores metadata for each processed PDF.
- file_hash (Primary Key): A SHA256 hash of the document's content.
- doc_name, source_path, file_size, mime, page_count.
- ingested_at: A timestamp that is populated after the document's chunks are successfully ingested.
- chunks table: Stores the individual text chunks and their embeddings.
- Contains the text, heading, page_start, page_end, and embedding for each chunk.
- file_hash (Foreign Key): Links each chunk back to its source document in the documents table.
- Python 3.x (I'm using 3.13)
- PostgreSQL with the pgvector extension
- An OpenAI API key
- An OpenRouter.ai API key
- Streamlit (included in environment.yml)
-
Clone the repository:
git clone \<repository-url\> -
Install dependencies:
conda env create -f environment.yml -
Configure environment variables:
Create a .env file in the project's root directory. Seesrc/settings.pyfor the required variables, which include database connection details and API keys.
-
Initialize the database tables:
python -m src/scripts/init_sql_tables.py -
Ingest documents:
Add your PDF files to the textbooks/ directory and run the ingestion script:
python -m src/scripts/ingest_pdf.py -
Run queries:
Command line interface (not recommended):
python -m src/scripts/query_rag.pyWeb interface (recommended):
python -m streamlit run src/scripts/streamlit_query_rag.pyThe Streamlit interface will open in your browser (typically at http://localhost:8501) where you can interact with the RAG system through an intuitive web interface with real-time parameter adjustment and source document transparency.