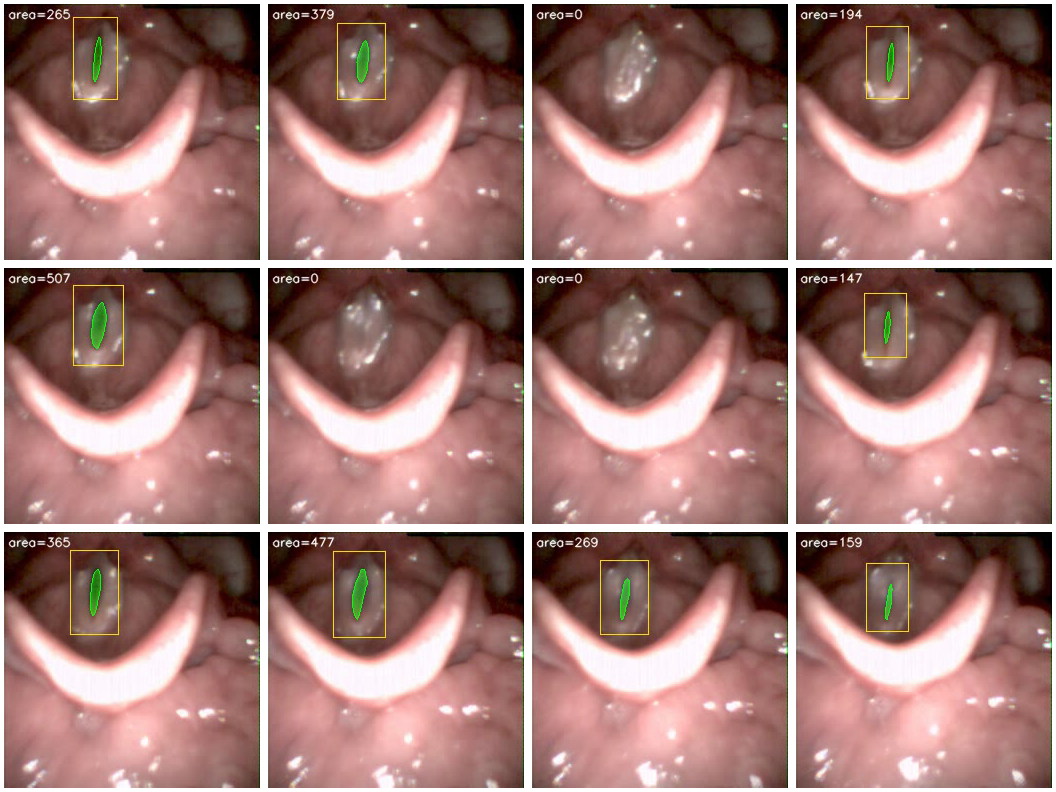
Frame montage: segmentation across a glottal cycle.
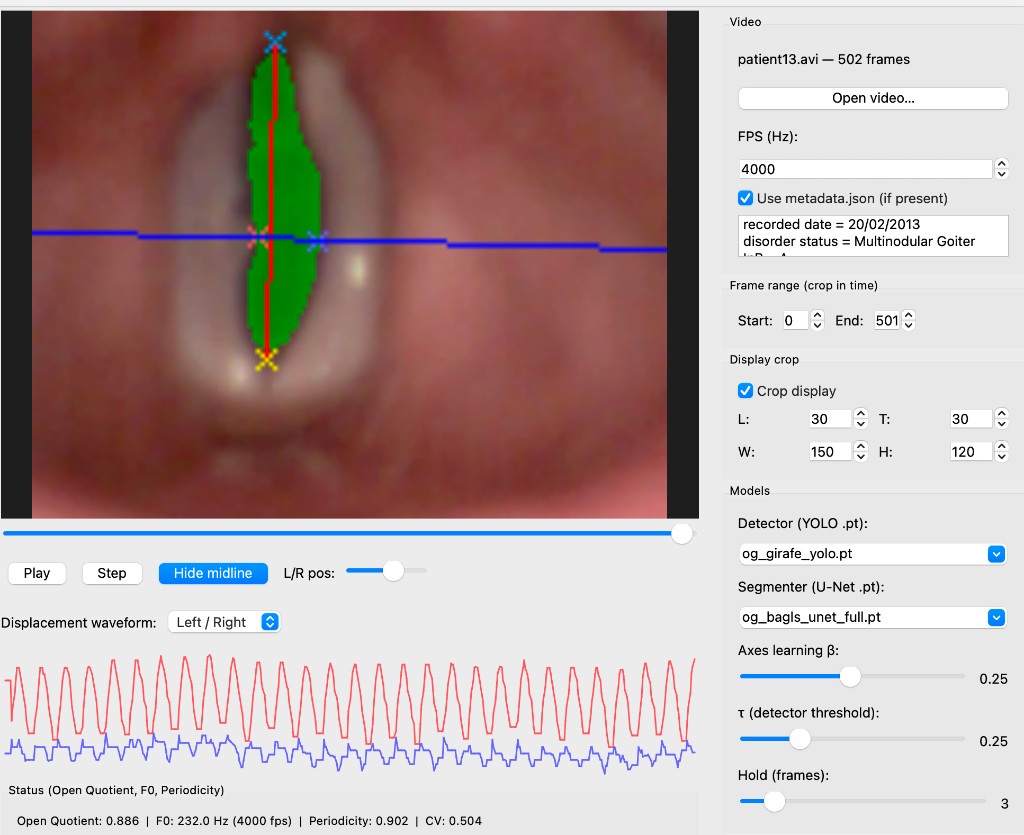
Quantifying L/R asymmetry in a case of vocal fold nodules using the dynamic medial probe and independent displacement waveforms.
Open-source toolkit for automated glottal area segmentation from high-speed videoendoscopy (HSV). Beyond per-frame masks, OpenGlottal extracts glottal area waveforms (GAW) and left/right (L/R) displacement waveforms along a configurable medial axis — enabling quantification of L/R asymmetry and kinematic features (open quotient, F0, periodicity) for clinical assessment.
Building on previous research into nodule-induced vibratory irregularities (Patel, Unnikrishnan, & Donohue, 2016), OpenGlottal provides the first open-source GUI to quantify these L/R asymmetries in real time.
Author: Harikrishnan Unnikrishnan (hari@orchard-robotics.com)
Paper: arXiv:2603.02087 — A Detection-Gated Pipeline for Robust Glottal Area Waveform Extraction and Clinical Pathology Assessment
OpenGlottal combines a YOLOv8 glottis detector, a U-Net pixel-level segmenter, and a temporal vocal fold tracker into a single, reproducible inference and training pipeline — trained and evaluated on the GIRAFE dataset (dataset paper) and BAGLS (Scientific Data, 2020).
Five pipelines are provided (four YOLO-gated and one U-Net-only):
| Pipeline | Flag | Description |
|---|---|---|
| 1 — VFT | vft |
YOLO detects the glottis → crop (size locked to first detection) → motion-based VocalFoldTracker |
| 2 — Guided VFT | guided-vft |
YOLO bbox as ROI mask on the full frame → YOLOGuidedVFT (no cropping) |
| 3 — YOLO+UNet | unet |
YOLO + full-frame U-Net: detection-gated (output only on frames where YOLO fires) |
| 4 — YOLO-Crop+UNet | yolo-crop+unet |
YOLO crop → resize to 256×256 → crop-trained U-Net → project mask back. Weights are provided; use --crop-weights in eval_girafe.py / eval_bagls.py. Once tests validate, these weights will be committed to the repo. |
| 5 — U-Net only | unet-only |
Full-frame U-Net only, no YOLO gate (only --unet-weights required) |
All pipelines produce a per-frame glottal area waveform. The Qt GUI and CLI further support L/R displacement waveform extraction: independent left and right vocal-fold excursions relative to a medial axis, with a configurable sampling position (e.g. anterior–middle junction) to isolate asymmetry. Kinematic features (open quotient, fundamental frequency, periodicity, etc.) are computed from area or L/R waveforms for downstream clinical analysis.
Use a virtual environment in the repo so all commands use the same Python and dependencies.
git clone https://github.com/hari-krishnan/openglottal.git
cd openglottal
python3 -m venv .venv
source .venv/bin/activate # Linux/macOS; on Windows: .venv\Scripts\activate
pip install -e ".[dev]"From a local clone (venv already exists):
cd /path/to/openglottal
source .venv/bin/activate
pip install -e ".[dev]" # only if you need to reinstallAlways use this venv for training, evaluation, and CLI: activate it (source .venv/bin/activate) before running any python/openglottal commands, or use the ./run script (e.g. ./run scripts/train_unet.py ...) which uses .venv/bin/python automatically.
For the Qt GUI, install the optional GUI extra: pip install -e ".[gui]" (adds PyQt5). See Qt GUI below.
(A pip install openglottal option will work once the package is published to PyPI.)
Requirements: Python ≥ 3.9, PyTorch ≥ 2.0, Ultralytics ≥ 8.0, OpenCV ≥ 4.8
Weights: Place pre-trained weights in weights/ (or pass full paths to the scripts). Provided weights:
| Weights file | Description |
|---|---|
weights/og_girafe_unet_full.pt |
GIRAFE-trained U-Net (full-frame, no crop) |
weights/og_girafe_unet_crop.pt |
GIRAFE-trained U-Net (crop mode; use with YOLO-Crop+UNet pipeline) |
weights/og_girafe_yolo.pt |
GIRAFE-trained YOLO glottis detector |
weights/og_bagls_unet_full.pt |
BAGLS-trained U-Net (full-frame, no crop) |
weights/og_bagls_unet_crop.pt |
BAGLS-trained U-Net (crop mode; use with YOLO-Crop+UNet on BAGLS in-distribution) |
weights/og_bagls_yolo.pt |
BAGLS-trained YOLO glottis detector |
To train your own, see Training; trained models are saved under outputs/.
import torch
from openglottal import TemporalDetector, UNet, extract_features_unet
device = torch.device("mps") # or "cuda" / "cpu"
detector = TemporalDetector("weights/og_girafe_yolo.pt")
model = UNet(1, 1, (32, 64, 128, 256)).to(device)
model.load_state_dict(torch.load("weights/og_girafe_unet_full.pt", map_location=device))
model.eval()
features = extract_features_unet("video.avi", detector, model, device)
print(features)
# {'area_mean': 312.4, 'area_std': 98.1, 'open_quotient': 0.61, 'f0': 0.017, ...}# U-Net pipeline (recommended)
openglottal run video.avi \
--yolo-weights weights/og_girafe_yolo.pt \
--unet-weights weights/og_girafe_unet_full.pt \
--pipeline unet \
--output results/
# Motion-based pipeline (no U-Net weights needed)
openglottal run video.avi \
--yolo-weights weights/og_girafe_yolo.pt \
--pipeline guided-vft \
--output results/Desktop app for viewing HSV videos with segmentation overlay, midline/AC–PC axes, L/R displacement waveforms, and kinematic metrics (open quotient, F0, periodicity, etc.). Quantifying L/R asymmetry in cases such as vocal fold nodules is a key use case: the dynamic medial probe and independent left/right waveforms isolate nodule-induced irregularities.
Pathology / L/R asymmetry — For clinicians, the L/R displacement waveform is the main diagnostic view (see screenshot at top).
pip install -e ".[gui]"
openglottal-guiLoad a video, choose detector and U-Net weights from weights/, set frame range, and use Play/Pause or the timeline slider. Optional metadata.json next to the video supplies FPS and other keys. See openglottal/qt_app/README.md for details.
Compare against the GIRAFE paper baselines using GIRAFE-trained weights:
python scripts/eval_girafe.py \
--images-dir GIRAFE/Training/imagesTr \
--labels-dir GIRAFE/Training/labelsTr \
--training-json GIRAFE/Training/training.json \
--unet-weights weights/og_girafe_unet_full.pt \
--yolo-weights weights/og_girafe_yolo.pt \
--device mpsResults are printed alongside the published GIRAFE baselines for direct comparison. Pass --output-json results.json to save raw per-frame scores.
| Method | Det.Recall | Dice | IoU | Dice≥0.5 |
|---|---|---|---|---|
| InP (GIRAFE paper) | n/a | 0.71 | n/a | n/a |
| U-Net (GIRAFE paper) | n/a | 0.64 | n/a | n/a |
| SwinUNetV2 (paper) | n/a | 0.62 | n/a | n/a |
| U-Net only | n/a | 0.81 | 0.70 | 96.2% |
| OTSU (baseline) | 0.95 | 0.22 | 0.13 | 2.5% |
| YOLO+UNet | 0.95 | 0.75 | 0.64 | 88.8% |
| YOLO-Crop+UNet† | 0.95 | 0.70 | 0.57 | 77.5% |
| Motion (baseline) | 0.95 | 0.27 | 0.17 | 9.7% |
- Det.Recall — fraction of frames where YOLO detected a glottis
- Dice — mean Dice coefficient across all test frames (higher is better)
- Dice≥0.5 — fraction of frames meeting the clinical pass threshold
† YOLO-Crop+UNet uses the crop-trained U-Net: weights/og_girafe_unet_crop.pt. Full-frame and crop weight files are not interchangeable.
YOLO+Motion underperforms because GIRAFE test frames are the first 20 frames per patient, providing insufficient temporal context for the motion tracker to converge.
No BAGLS data used in training. Use GIRAFE-trained weights; images are letterboxed to 256×256.
python scripts/eval_bagls.py \
--bagls-dir BAGLS/test \
--unet-weights weights/og_girafe_unet_full.pt \
--crop-weights weights/og_girafe_unet_crop.pt \
--yolo-weights weights/og_girafe_yolo.pt \
--device mps| Method | Det.Recall | Dice | IoU | Dice≥0.5 |
|---|---|---|---|---|
| U-Net only | 1.00 | 0.59 | 0.50 | 67.1% |
| YOLO+UNet | 0.69 | 0.55 | 0.47 | 61.9% |
| YOLO-Crop+UNet | 0.69 | 0.61 | 0.53 | 70.3% |
(Table at default confidence $\tau=0.25$. With --conf 0.02, YOLO-Crop+UNet reaches Dice 0.64 and Dice≥0.5 76.4%; see paper and sweep_bagls_conf.py.)
YOLO-Crop+UNet is the strongest pipeline on the unseen BAGLS data (+2 pp Dice, +3.2 pp Dice≥0.5 over U-Net alone at default
Use the provided BAGLS-trained U-Net (full-frame and crop) and YOLO weights:
python scripts/eval_bagls.py \
--bagls-dir BAGLS/test \
--unet-weights weights/og_bagls_unet_full.pt \
--crop-weights weights/og_bagls_unet_crop.pt \
--yolo-weights weights/og_bagls_yolo.pt \
--device mpsOn the 3 500-frame BAGLS test set this configuration achieves:
| Method | Det.Recall | Dice | IoU | Dice≥0.5 |
|---|---|---|---|---|
| U-Net only | 1.00 | 0.85 | 0.77 | 94.0% |
| YOLO+UNet | 0.87 | 0.85 | 0.78 | 94.6% |
| YOLO-Crop+UNet | 0.87 | 0.74 | 0.64 | 87.1% |
So BAGLS-trained YOLO+UNet sets a strong in-distribution baseline (Dice 0.85), while the GIRAFE-trained YOLO-Crop+UNet remains the best cross-dataset configuration.
openglottal build-dataset \
--images-dir GIRAFE/Training/imagesTr \
--labels-dir GIRAFE/Training/labelsTr \
--training-json GIRAFE/Training/training.json \
--output-dir yolo_dataOr via script:
python scripts/train_yolo.py \
--images-dir GIRAFE/Training/imagesTr \
--labels-dir GIRAFE/Training/labelsTr \
--training-json GIRAFE/Training/training.json \
--epochs 100YOLO saves best weights to outputs/yolo/exp/weights/best.pt (default run name exp). Use that path for train_unet_crop.py below, or copy to weights/og_girafe_yolo.pt for eval/CLI.
python scripts/train_unet.py \
--images-dir GIRAFE/Training/imagesTr \
--labels-dir GIRAFE/Training/labelsTr \
--training-json GIRAFE/Training/training.json \
--output outputs/og_girafe_unet_full.pt \
--epochs 50Uses YOLO to crop each training image to the glottis region, resizes to 256×256, and trains U-Net on these patches. At inference time the YOLO-Crop+UNet pipeline crops the input, runs U-Net, and projects the mask back to full-frame coordinates.
python scripts/train_unet_crop.py \
--images-dir GIRAFE/Training/imagesTr \
--labels-dir GIRAFE/Training/labelsTr \
--training-json GIRAFE/Training/training.json \
--yolo-weights outputs/yolo/exp/weights/best.pt \
--output outputs/og_girafe_unet_crop.pt \
--crop-size 256 \
--epochs 50 \
--device cpuDevice: Use --device cuda on a Linux GPU machine or cloud instance. On Mac, use --device cpu for training (MPS is not used for training); use --device mps for evaluation (eval_bagls.py, eval_girafe.py, analyze_gaw.py) for faster inference.
Both training modes use a 50/50 BCE + Dice loss with cosine annealing, saving the best validation checkpoint automatically.
openglottal/
├── weights/ # pre-trained weights (see table above)
├── outputs/ # trained models (gitignored) when training your own
├── openglottal/
│ ├── models/
│ │ ├── detector.py # TemporalDetector — YOLOv8 + temporal box locking
│ │ ├── tracker.py # VocalFoldTracker, YOLOGuidedVFT
│ │ └── unet.py # UNet, DoubleConv, GlottisDataset (with augmentation)
│ ├── qt_app/ # Qt GUI — overlay, midline, waveform, openglottal-gui
│ │ ├── main.py # Main window, inference worker, playback
│ │ ├── utils.py # Overlay, axes/AC-PC drawing
│ │ └── README.md # GUI usage and features
│ ├── metadata.py # Video I/O, metadata.json, frame loading (used by GUI)
│ ├── features.py # extract_features_{detector,yolo_guided_vft,unet}
│ ├── data.py # mask_to_yolo, build_yolo_dataset
│ ├── utils.py # I/O helpers, dice/IoU metrics, unet_segment_frame
│ └── cli.py # `openglottal` command-line interface
├── scripts/
│ ├── train_yolo.py # standalone YOLO training script
│ ├── train_unet.py # standalone U-Net training script
│ ├── infer.py # batch inference: AVI dir or image sequence → _out.avi
│ ├── eval_girafe.py # per-patient test evaluation vs GIRAFE baselines
│ ├── eval_bagls.py # cross-dataset evaluation on BAGLS (3 500 frames)
│ ├── analyze_gaw.py # GAW feature analysis: Healthy vs Pathological (65 patients)
│ ├── download_datasets.py # download GIRAFE and BAGLS from Zenodo
│ ├── prepare_girafe_splits.py # build GIRAFE training.json from images/labels
│ ├── prepare_bagls_splits.py # build BAGLS train/test splits
│ ├── make_montage.py # build frame montage PNGs for paper figures
│ ├── sweep_bagls_conf.py # YOLO confidence threshold sweep on BAGLS
│ └── train_unet_crop.py # train U-Net on YOLO-cropped patches (higher res)
└── configs/
└── default.yaml # all hyperparameters documented in one place
OpenGlottal is developed and evaluated on GIRAFE and BAGLS. Download them (optional: use the script) then point the training/eval scripts at the extracted directories.
# Download GIRAFE and/or BAGLS to the current directory (or use --output-dir)
python scripts/download_datasets.py --girafe --bagls-
GIRAFE (Zenodo): zenodo.org/records/13773163 — 760 frames (256×256 px), expert-annotated glottal masks. Dataset paper: Data in Brief (2025). After unpacking:
GIRAFE/Training/imagesTr/,GIRAFE/Training/labelsTr/,GIRAFE/Training/training.json; raw videos:GIRAFE/Raw_Data/. If you have a copy in./sdsc/glottal_area, copy it over then run the prepare script:mkdir -p GIRAFE/Training && cp -r sdsc/glottal_area/imagesTr sdsc/glottal_area/labelsTr GIRAFE/Training/ python scripts/prepare_girafe_splits.py --images-dir GIRAFE/Training/imagesTr --labels-dir GIRAFE/Training/labelsTr --output GIRAFE/Training/training.json -
BAGLS (Zenodo): zenodo.org/records/3762320 — benchmark for automatic glottis segmentation. Dataset paper: Gómez et al., Scientific Data (2020). After downloading with
download_datasets.py --bagls, useBAGLS/testas--bagls-dirfor eval andBAGLS/trainingfor training.
| Split (GIRAFE) | Frames |
|---|---|
| Train | ~608 |
| Val | ~76 |
| Test | ~76 |
The following scalar features are extracted from each glottal area waveform (or from the L/R-derived opening signal when using displacement mode):
| Feature | Description |
|---|---|
area_mean |
Mean glottal area (px²) |
area_std |
Standard deviation of area |
area_range |
Max − min area |
open_quotient |
Fraction of cycle with area above 10 % of mean |
f0 |
Dominant frequency from FFT (cycles/frame; multiply by capture fps for Hz) |
periodicity |
Peak autocorrelation at lags 1–50 |
cv |
Coefficient of variation (std / mean) |
Spatial kinematics (L/R asymmetry)
- L/R displacement: Independent tracking of left and right vocal-fold excursions (px) relative to the medial axis. Exported as separate time series (L, R, L−R, area) via the GUI “Save analysis” or the CLI
openglottal displacementcommand. - Dynamic medial probe: Configurable sampling position (0–1 along the anterior–posterior axis) so analysis can be run at a specific anatomical point (e.g. junction of anterior and middle thirds) to isolate nodule-induced irregularities.
Beyond frame-level segmentation, the pipeline produces a Glottal Area Waveform (GAW) — the per-frame glottal area over time — and optionally L/R displacement waveforms (left and right excursions relative to the medial axis). Kinematic features can be extracted from either GAW or L/R-derived signals and used for clinical classification and asymmetry assessment.
python scripts/analyze_gaw.py \
--raw-data-dir GIRAFE/Raw_Data \
--yolo-weights weights/og_girafe_yolo.pt \
--unet-weights weights/og_girafe_unet_full.pt \
--device mps \
--output-dir results/gawThis script processes all 65 GIRAFE patients, extracts kinematic features from each area waveform, and compares Healthy vs Pathological groups using Mann-Whitney U tests.
Because the cohort has a significant sex imbalance (see note below), results are reported stratified by sex rather than pooled.
Female subgroup (12 H / 11 P):
| Feature | Healthy (mean±std) | Pathological (mean±std) | p-value |
|---|---|---|---|
| area_mean | 125.2 ± 43.1 | 247.8 ± 204.6 | 0.230 |
| area_std | 112.9 ± 32.2 | 118.9 ± 96.0 | 0.406 |
| area_range | 336.7 ± 97.6 | 375.5 ± 272.2 | 0.559 |
| open_quotient | 0.760 ± 0.207 | 0.874 ± 0.131 | 0.192 |
| f0 | 241.7 ± 34.8 Hz | 203.5 ± 73.6 Hz | 0.156 |
| periodicity | 0.955 ± 0.008 | 0.946 ± 0.013 | 0.255 |
| cv | 0.95 ± 0.20 | 0.57 ± 0.29 | 0.006* |
Male subgroup (3 H / 14 P):
| Feature | Healthy (mean±std) | Pathological (mean±std) | p-value |
|---|---|---|---|
| area_mean | 192.1 ± 18.3 | 172.7 ± 94.0 | 0.768 |
| area_std | 142.7 ± 35.0 | 92.0 ± 66.9 | 0.197 |
| area_range | 439.7 ± 86.7 | 343.1 ± 212.3 | 0.488 |
| open_quotient | 0.860 ± 0.145 | 0.843 ± 0.186 | 1.000 |
| f0 | 183.3 ± 75.0 Hz | 82.5 ± 79.3 Hz | 0.169 |
| periodicity | 0.962 ± 0.001 | 0.900 ± 0.116 | 0.068 |
| cv | 0.75 ± 0.19 | 0.63 ± 0.40 | 0.509 |
* p < 0.05 (Mann-Whitney U, two-sided)
Sex imbalance note: The Healthy group is 80% female (12F/3M) while the Pathological group is 56% male (14M/11F; Fisher's exact p=0.025). Because f0 is strongly sex-dependent (males ~100 Hz vs females ~224 Hz), pooling would confound f0. After stratifying, only cv (coefficient of variation) reaches significance in the female subgroup (p=0.006). In the male subgroup (n=3 Healthy), cv trends in the same direction (0.75 vs 0.63) but does not reach significance (p=0.509); periodicity approaches significance (p=0.068). The male subgroup is too small for reliable inference.
YOLO acts as a detection gate: when the endoscope moves away from the glottis (scope insertion, patient coughing, instrument in view), YOLO fires no detection and the area is set to zero — preventing spurious waveform spikes that would corrupt downstream feature extraction.
If you use OpenGlottal in your research, please cite:
H. Unnikrishnan. A Detection-Gated Pipeline for Robust Glottal Area Waveform Extraction and Clinical Pathology Assessment. arXiv:2603.02087 [cs.CV], 2 Mar 2026. https://arxiv.org/abs/2603.02087
@misc{unnikrishnan2026openglottal,
title = {A Detection-Gated Pipeline for Robust Glottal Area
Waveform Extraction and Clinical Pathology Assessment},
author = {Unnikrishnan, Harikrishnan},
year = {2026},
eprint = {2603.02087},
archivePrefix = {arXiv},
primaryClass = {cs.CV},
url = {https://arxiv.org/abs/2603.02087}
}Related prior work on glottal kinematics and high-speed imaging:
@article{patel2013invivo,
title = {In vivo measurement of pediatric vocal fold motion using
structured light laser projection},
author = {Patel, Rita R and Donohue, Kevin D and Lau, Daniel and
Unnikrishnan, Harikrishnan},
journal = {The Laryngoscope},
year = {2013}
}
@article{patel2016effects,
title = {Effects of vocal fold nodules on glottal cycle measurements
derived from high-speed digital imaging},
author = {Patel, Rita R and Unnikrishnan, Harikrishnan and Donohue, Kevin D},
journal = {Journal of Speech, Language, and Hearing Research},
year = {2016}
}OpenGlottal is research and educational software. It is not a medical device and is not intended for diagnosis, treatment, or clinical decision-making. Outputs may be inaccurate; always rely on qualified clinical judgment and validated workflows where patient care is involved.
See CONTRIBUTING.md for setup, tests, and linting.
If you use this work in research, cite the arXiv preprint. Repository metadata for citation managers is in CITATION.cff.
MIT License — see LICENSE.