-
Notifications
You must be signed in to change notification settings - Fork 6
Foreground specification
Using the --foreground option with --fg_format 1 (default setting), you can specify focal lineages, whose branch combinations are identified in the CSUBST output files. You can also limit the analysis to foreground branches only (with --exhaustive_until 1, see here). The input file of --foreground should be a tsv containing 2 columns without a header:
1 Homo_sapiens_.*
2 Mus_musculus_.*
3 Danio_rerio_.*
The first column specifies the unique identifier of lineages.
The second column is the regex-compatible leaf label specification. For example, Homo_sapiens_.* allows CSUBST to select all leaves with the prefix Homo_sapiens_ and their ancestors as foreground branches.
Other options can be combined to accurately specify complex foreground combinations as below.
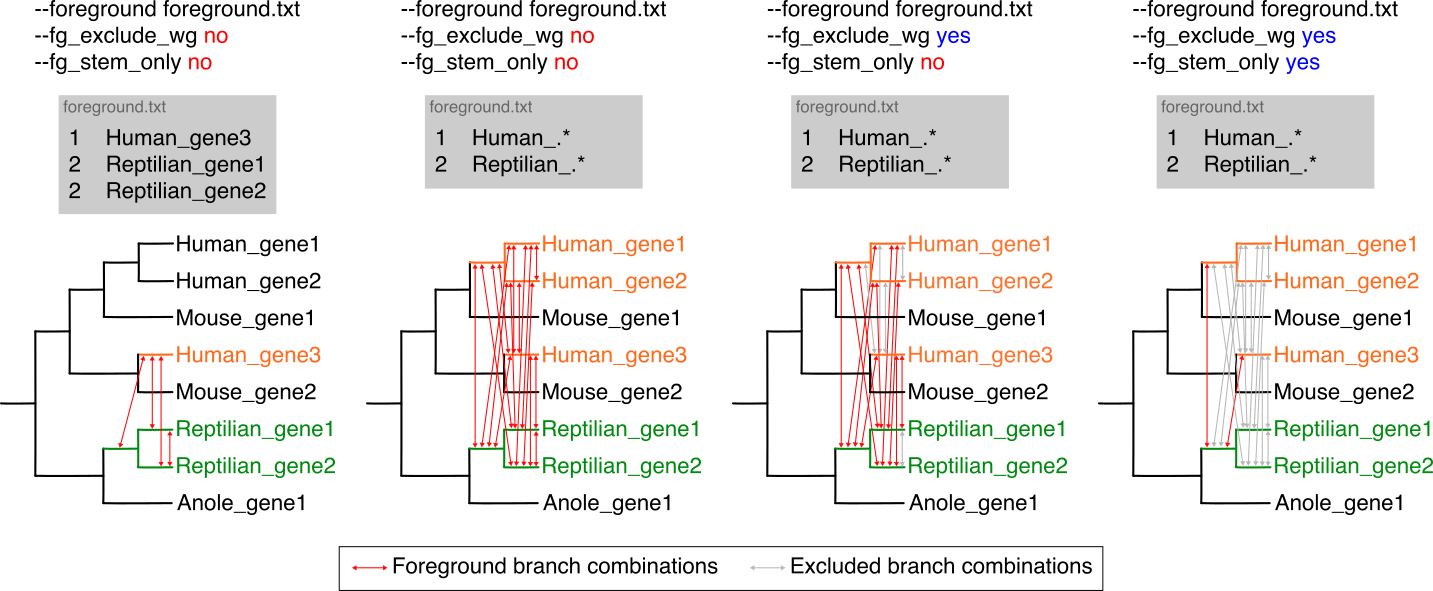
To analyze multiple traits in a single CSUBST run, use --fg_format 2. This format allows --foreground to accept a TSV file with a header line indicating foreground lineages for multiple traits. In this context, lineage ID 0 denotes a background lineage. Example files for --fg_format 1 (named foreground.txt) and --fg_format 2 (named foreground.tsv) can be found with csubst dataset --name PEPC.
The table below shows an example of specifying lineages for different traits:
| name | C4 | C4_monocot | C4_dicot | crop |
|---|---|---|---|---|
| Alternanthera_ficoidea_.* | 0 | 0 | 0 | 0 |
| Alternanthera_pungens_.* | 1 | 0 | 1 | 0 |
| Alternanthera_sessilis_.* | 0 | 0 | 0 | 0 |
| Amaranthus_hypochondriacus_.* | 1 | 0 | 1 | 1 |
| Amborella_trichopoda_.* | 0 | 0 | 0 | 0 |