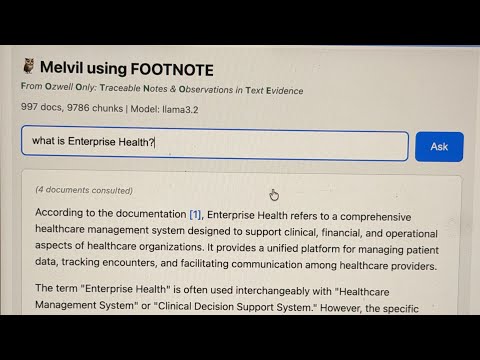
FOOTNOTE — From Ozwell Only: Traceable Notes & Observations in Text Evidence
FOOTNOTE is a portable index format within Artipods designed for RAG (Retrieval-Augmented Generation) with proper citations. It compiles markdown documentation into a self-contained SQLite database that combines:
- sqlite-vec for semantic vector search (find conceptually similar content)
- FTS5 for full-text BM25 search (find exact keywords and phrases)
- Literal search for special characters and code patterns (HL7 messages like
ADT^A04)
The result is a single index.sqlite file that can be deployed anywhere—servers, edge functions, or even in-browser via WebAssembly.
Melvil is an agentic RAG assistant (implemented in ask.ts) that uses FOOTNOTE indexes to answer questions with citations. Named after Melvil Dewey, the inventor of the Dewey Decimal System, Melvil helps users find information in large documentation sets.
Melvil features:
- Tool-based search: The LLM decides which search strategy to use (hybrid, FTS, literal)
- Multi-turn reasoning: Can search multiple times to gather comprehensive answers
- Citation tracking: Every answer includes
[1],[2]references to source documents - Debug & reporting: Saves conversation logs and allows users to flag bad answers for review
This tool (docidx) is a markdown compiler that produces FOOTNOTE-enabled Artipods for server-side RAG consumption.
See https://alexgarcia.xyz/sqlite-vec/wasm.html for in browser example
cd tools/docidx
npm installSmart build (auto-detects everything):
npm run docidx -- buildForce clean rebuild:
npm run docidx -- build --cleanUse specific Ollama model:
npm run docidx -- build --embedding-model ollama:nomic-embed-textUse OpenAI (requires OPENAI_API_KEY):
npm run docidx -- build --embedding-model text-embedding-3-small--root <path>- Hugo project root (auto-detected by finding content/ directory)--content <path>- Content directory relative to root (default: content)--out <path>- Output artipod directory (default: ./artipod relative to Hugo root)--clean- Remove existing index and rebuild from scratch--incremental- Only process changed files--include-drafts- Include draft documents--filter <name=value>- Content filter (e.g.,--filter brand=eh)--embedding-model <model>- Embedding model (auto-detected, see below)--embedding-dim <n>- Embedding dimensions (auto-detected based on model)--pull- Automatically pull Ollama model if not installed--max-tokens <n>- Max tokens per chunk (default: 500)--overlap <n>- Token overlap between chunks (default: 80)--copy-content- Copy markdown files to artipod for grep-based search
Auto-detection behavior:
- Hugo root: Walks up from current directory looking for
content/ - Build mode: Incremental if index exists, clean if not
- Embedder: Checks in order:
- Ollama (if running locally with embedding model like
nomic-embed-text) - OpenAI (if
OPENAI_API_KEYis set) - Error with helpful instructions
- Ollama (if running locally with embedding model like
Supported Ollama embedding models:
nomic-embed-text(768 dim) - recommended, fastbge-m3(1024 dim) - multilingual, high qualitymxbai-embed-large(1024 dim)bge-large(1024 dim)snowflake-arctic-embed(1024 dim)all-minilm(384 dim) - smallest/fastest
Installing Ollama models:
Option 1 - Manually pull first:
ollama pull nomic-embed-textOption 2 - Auto-pull with --pull flag:
./docidx.sh build --embedding-model ollama:bge-m3 --pullnpm run docidx -- query --hybrid "patient registration" --k 5Test different search modes interactively:
./docidx.sh test --mode fts "FHIR API"
./docidx.sh test --mode hybrid "how do I schedule"
./docidx.sh test --mode literal "ADT^A04"
./docidx.sh test --mode grep "copyright" # requires --copy-contentAsk questions with an agentic RAG assistant:
./docidx.sh ask "How do I schedule an appointment?"
./docidx.sh ask -v "What is FHIR?" # verbose: show tool calls
./docidx.sh ask -c "terms of use" # show full chunk contentThe agent has access to these tools:
search_hybrid- Vector + keyword search (best for natural language)search_fts- Full-text BM25 search (best for technical terms)search_literal- Exact substring match (finds special chars like^,|)search_grep- Unix grep on raw files (if--copy-contentenabled)read_document- Read full document contentfind_related- Find documents that link to a given document
The ask agent includes a built-in HTTP server that demonstrates how external applications can consume an artipod. This serves as a proof of concept for building documentation assistants, chatbots, or search interfaces.
./docidx.sh ask --serve --port 3000 # Start server
./docidx.sh ask --serve --port 3000 --verbose # With request loggingEndpoints:
| Endpoint | Method | Description |
|---|---|---|
/ |
GET | Web UI for interactive Q&A |
/health |
GET | Health check + stats (doc count, chunk count, model) |
/ask?q=... |
GET | Ask question, returns JSON response |
/ask |
POST | Ask with JSON body {"question": "..."} |
/ask/stream?q=... |
GET | Streaming SSE response (real-time tokens) |
JSON Response Format:
{
"answer": "The patient registration process involves...",
"references": [
{
"index": 1,
"doc_id": "doc_abc123",
"url": "/functions/patient-registration/",
"title": "Patient Registration",
"headings": ["Manual Entry", "Required Fields"],
"method": "hybrid",
"score": 0.0241
}
]
}Streaming SSE Events:
data: {"type": "thinking", "message": "Searching... (iteration 1)"}
data: {"type": "tool_call", "tool": "search_fts", "query": "patient registration"}
data: {"type": "token", "content": "To register"}
data: {"type": "token", "content": " a patient..."}
data: {"type": "done", "references": [...]}
data: [DONE]
Building Your Own Client:
The artipod is designed to be consumed by any application that can:
- Read SQLite (for direct search access)
- Call an embedding API (Ollama/OpenAI) for vector queries
- Optionally, wrap an LLM for agentic search
See the SqliteStore class for direct database access patterns, or use the HTTP API as a simpler integration point.
The ask command automatically injects a FINAL: answer-marker requirement into all agent prompts. This ensures reliable detection of when the LLM has finished responding, especially in tool-calling workflows where the model may emit multiple intermediate outputs.
With this injection, the agent must output exactly one of two things:
- a tool call expressed as valid JSON only, or
- a completed user-facing answer prefixed with
FINAL:and containing no planning, narration, or tool references.
Why FINAL:? This convention avoids ambiguity in streaming and multi-turn tool workflows and is more robust than XML tags or heuristic detection. The FINAL: prefix is widely used in agent frameworks and has been shown in practice to be the most consistently followed boundary marker across OpenAI, Anthropic, Gemini, and Ollama-hosted models. If you provide a custom prompt via --prompt-file, the tool appends the FINAL: requirement automatically—you don't need to include it yourself.
References / rationale:
https://chatgpt.com/share/69805745-bfb8-8004-a6ba-2d69750cdc6a
- OpenAI & Anthropic agent patterns separating tool calls from final answers
- LangChain / LangGraph best practices for tool-calling agents
- ReAct-style prompting (Yao et al., 2022) emphasizing explicit action vs. answer boundaries
- Production experience across mixed-vendor LLM stacks showing higher compliance with simple textual sentinels than XML-style tags
Benchmarked on ~1000 markdown documents (~10K chunks) on macOS:
| Search Mode | Time | Description |
|---|---|---|
| FTS | ~6ms | SQLite FTS5 with BM25 ranking |
| Hybrid | ~50ms | Vector + FTS combined (includes embedding) |
| Literal | ~120ms | Exact substring match via SQL LIKE |
| Grep | ~220ms | Unix grep on raw files |
FTS (SQLite) - 6ms:
1. [7.71] Terms of Use > General Prohibitions
/resources/.../terms-of-use/
"...infringe any copyright, trademark rights..."
2. [6.86] Terms of API Use > User Content
/resources/.../terms-of-api-use/
"...may constitute copyright or HIPAA infringement..."
Grep - 219ms:
1. [5 matches] terms-of-api-use
L43: Remember, it's not your content...
L145: ...infringe any copyright, trademark...
2. [5 matches] terms-of-use
L43: Remember, it's not your content...
| Feature | SQLite FTS/Hybrid | Grep |
|---|---|---|
| Speed | ~36x faster | Slower (reads all files) |
| Ranking | BM25 relevance scores | Match count only |
| Results | Chunk-level with context | Line-level |
| Special chars | Tokenized (use literal mode) |
Native support |
| Setup | Requires index build | No setup needed |
To enable grep-based search (for comparison or fallback):
./docidx.sh build --copy-contentThis copies markdown files to artipod/content/ (~3MB for 1000 docs).
OPENAI_API_KEY- Required only if using OpenAI embeddings (not needed for Ollama)LOG_LEVEL- Logging verbosity (debug, info, warn, error)
See ARTIPOD-README.md for the full specification.
artipod/
├── index.sqlite # SQLite database (vectors + FTS)
├── manifest.json # Build metadata and configuration
├── .embed-{model}/ # Embedding cache (e.g., .embed-nomic-embed-text)
│ └── {hash[0:2]}/ # Subdirectories by 2-char hash prefix
│ └── {hash[2:]}.bin # Binary Float32 embedding vectors
└── content/ # (optional) Raw markdown for grep search
The .embed-{model} folder caches embeddings by content hash to avoid re-vectorizing unchanged chunks across builds. This provides significant speedups for incremental builds where only a few documents change.
- Location:
{artipod}/.embed-{model}(e.g.,.embed-nomic-embed-text) - Format: Binary Float32Array files (~3KB per 768-dim embedding)
- Organization: Files are stored in subdirectories by 2-char hash prefix (e.g.,
ab/cdef1234.bin) to avoid slow directory listings when cache contains tens of thousands of entries - Benefits:
- Rebuilds with unchanged content are nearly instant (only metadata updates)
- Switching between clean/incremental builds reuses cached embeddings
- Cache persists even if the SQLite database is deleted
The cache is model-specific, so switching embedding models creates a new cache folder.
| Table | Purpose |
|---|---|
chunks |
Main chunk metadata (doc_id, path, url, title, section, tags, headings, content, content_hash) |
vec_chunks |
sqlite-vec virtual table for vector KNN search |
chunks_fts |
FTS5 virtual table for BM25 full-text search |
file_tracking |
Track processed files for incremental builds |
metadata |
Key-value store (currently stores dimension) |
Contains:
schema_version- Artipod schema versionbuild_time_utc- Build timestampsource_git_commit- Git commit hash (if available)chunking- Chunking parameters (max_tokens, overlap)embedding- Embedding model configurationfts_fields- Fields indexed for full-text searchdoc_count- Number of documents indexedchunk_count- Number of chunks indexed
# Run with tsx for development
npm run docidx -- build --root ../.. --content content --out ./artipod --clean --embedding-model mock
# Build TypeScript
npm run build
# Run tests
npm test