Dali PCA equivalent (correspondence analysis) on aligned pdbs #2063
Replies: 9 comments
-
|
Yes with a bit of hacking. You just need to provide the PCA with an array of coordinate sets or an object that can give it one. Why would you do that though? What are you trying to do? |
Beta Was this translation helpful? Give feedback.
-
|
Thank you. The output of the DALI server provides a plot of the first two PCs, so I was trying to recreate this (since DaliRecord allows the mappings to be used in making an ensemble). However, it wasn't clear to me if this would produce the same result as DALI, and if making an ensemble is the correct route (if dummy atoms are introduced in mapping?). |
Beta Was this translation helpful? Give feedback.
-
|
Oh, that's interesting. Is that a new feature on DALI? You're right that dummy atoms could change things, although I think the way they are handled should mitigate this. What you could do is trim the ensemble to an occupancy level where you don't have many sites with significant numbers of dummy atoms or even to 1.0 to remove all of them and see what happens. |
Beta Was this translation helpful? Give feedback.
-
|
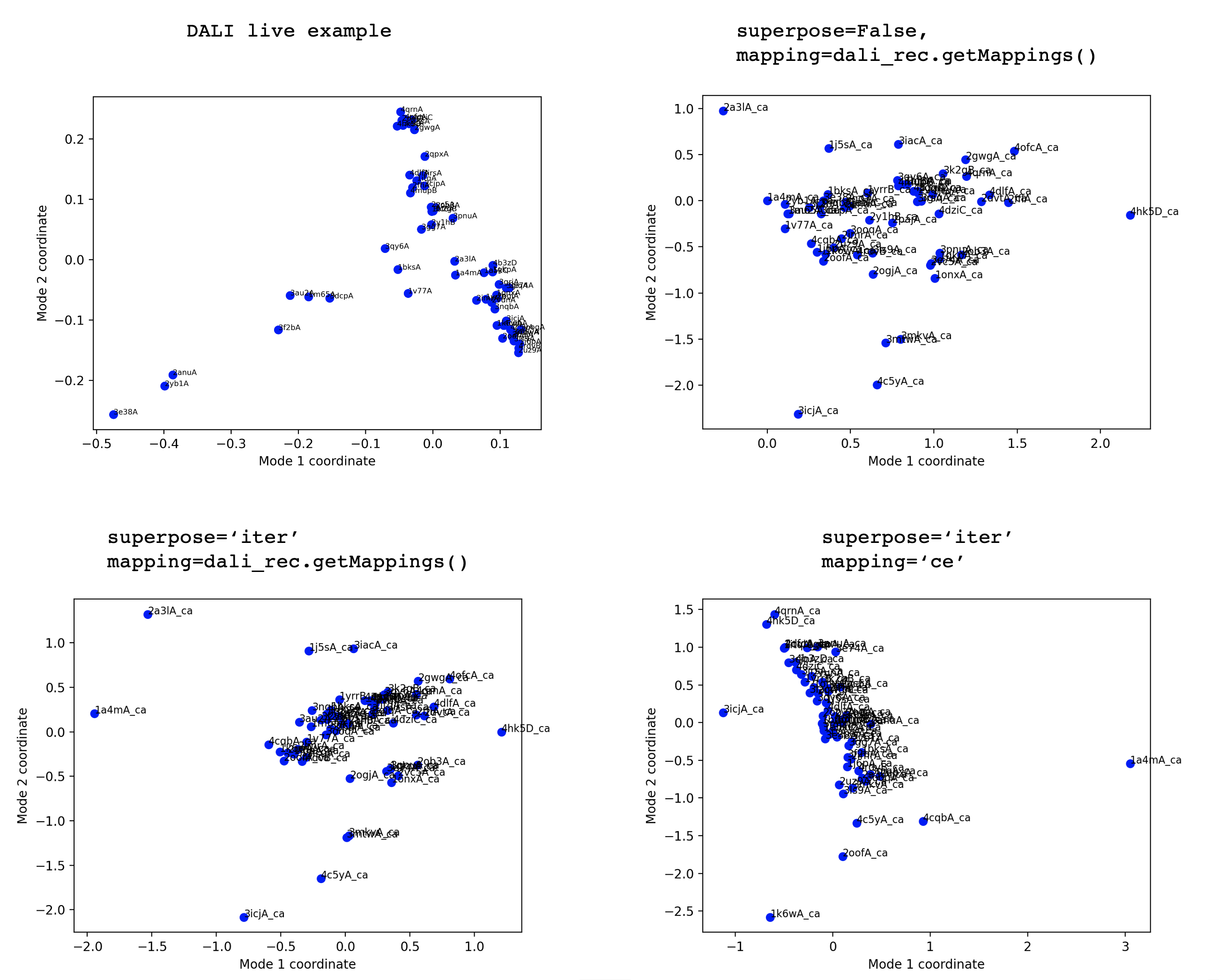
I'm not sure how new it is — there is a live example here: http://ekhidna2.biocenter.helsinki.fi/daliviewer/matrix_EXAMPLE/ Thanks, I will try recreating the PCA plot in their example using trimPDBEnsemble with different occupancy values (e.g. 1 or 0) like you suggest, and see how these compare. Going back to your initial suggestion, how would you go about hacking to provide an array of coordinate sets, from a list of pdbs? I could then compare with this also. |
Beta Was this translation helpful? Give feedback.
-
|
Actually, I'm not sure hacking it would work because you'd still have the problem with different numbers of residues. What you could do is use buildPDBEnsemble with superpose=False |
Beta Was this translation helpful? Give feedback.
-
|
trimPDBEnsemble with occupancy=1 seems to get rid of too many atoms, for this data set at least (59 chains in DALI example above) so have continued with default occupancy for now. So far I haven't been able to recreate the PCA plot trying buildPDBEnsemble superpose=False or 'iter', with mappings from DALI or with mapping='ce' (seqid=0, overlap=0). This is after applying the translation-rotation matrices from DALI to align all structures to 1a4m-A. |
Beta Was this translation helpful? Give feedback.
-
|
Do we know that Dali is using PCA? Could it be some other dimensionality reduction? |
Beta Was this translation helpful? Give feedback.
-
|
It could also be because we're not using Dali transformations. Once I manage to parse and apply them too then this could be worth trying again with superpose=False In any case, I notice that you're missing the option superpose=True for just running one superposition rather than the iterative superposition. I'm not sure if that might help. |
Beta Was this translation helpful? Give feedback.
-
In fact, Dali is using correspondence analysis of the similarity matrix rather than a PCA of the positional covariance. It’s not surprising then that the output is quite different. It may be possible to add such an analysis into ProDy though. |
Beta Was this translation helpful? Give feedback.
Uh oh!
There was an error while loading. Please reload this page.
Uh oh!
There was an error while loading. Please reload this page.
-
Apologies for the naive question, but is it possible to perform PCA as described in the tutorials on a list of aligned (heterogeneous) pdb files rather than on a PDBEnsemble made from them?
Thank you,
Beta Was this translation helpful? Give feedback.
All reactions